Manuscript accepted on :04-06-2025
Published online on: 19-06-2025
Plagiarism Check: Yes
Reviewed by: Dr. Hassan Shora
Second Review by: Dr. Randa Salah Gomaa
Final Approval by: Dr. Patorn Piromchai
Khalid Mohammad. Abu Khadra1* , Mohammad Ahmad Wahsha2
, Mohammad Ahmad Wahsha2 and Tariq Hassan Al-Najjar3
and Tariq Hassan Al-Najjar3
1Department of Biological Sciences, Yarmouk University, Irbid, Jordan
2Marine Science Station, The University of Jordan, Aqaba Branch, Jordan
3Department of Marine Biology, The University of Jordan, Aqaba Branch, Jordan
Corresponding Author E-mail:khalidmohd99@yahoo.com
DOI : https://dx.doi.org/10.13005/bpj/3182
Abstract
Genetic association studies are beginning to play a role in studying and analysing the risk factors of many complicated metabolic disorders, such as Type 2 Diabetes Mellitus (T2DM). This study appraises the association between the SLC30A8 gene polymorphism (rs13266634 C/T) and T2DM in North Jordan as a potential genetic risk factor for T2DM. The rs13266634 polymorphism was analysed using polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP). The CT genotype was the most prevalent in T2DM (84%) and nondiabetic control groups (91.2%). The TT genotype was observed in only 5.5% of the T2DM group and 4.9% of the nondiabetic group. Allelic analysis disclosed that 49% of the studied population have the T allele, and 51% have the C allele. There were no significant differences in the genotype distribution between the T2DM and the control groups, suggesting that the SLC30A8 gene polymorphism (rs13266634 C/T) is not associated with T2DM in North Jordan and indicating that the SLC30A8 rs13266634 polymorphism is not considered a genetic risk factor for T2DM in the North Jordanian population.
Keywords
Gene Polymorphism; Genetic Association; North Jordan; SLC30A8; Type 2 Diabetes Mellitus
Download this article as:| Copy the following to cite this article: Abu-Khadra K. M, Wahsha M. A, Al-Najjar T. H. Analysis of the Association between SLC30A8 Gene Polymorphism (rs13266634 C/T) and Type 2 Diabetes Mellitus in the North of Jordan. Biomed Pharmacol J 2025;18(2). |
| Copy the following to cite this URL: Abu-Khadra K. M, Wahsha M. A, Al-Najjar T. H. Analysis of the Association between SLC30A8 Gene Polymorphism (rs13266634 C/T) and Type 2 Diabetes Mellitus in the North of Jordan. Biomed Pharmacol J 2025;18(2). Available from: https://bit.ly/4eddinJ |
Introduction
Diabetes is a great public health concern, it’s the seventh leading cause of mortality worldwide.1 Diabetes mellitus (DM) is a metabolic disease caused by many risk factors leading to damage in pancreatic β-cells or malfunction in insulin, leading to a collapse in macronutrient metabolism.2,3 The most common form of DM is Type 2 diabetes mellitus (T2DM), a complicated chronic disorder arising from an array of dysfunctions characterised by hyperglycemia and resulting from a combination of insufficient insulin secretion or resistance to insulin action.4,5 The pathology of T2DM is influenced by multiple etiological factors involving complex interactions between environmental and genetic factors.6,7,8 The prevalence of T2DM and its associated complications has rapidly increased in developed and developing countries.9,10 The T2DM incidence is projected to increase significantly over the last thirty years, driven by rising obesity rates.11
Numerous genetic variants have been linked to T2DM according to research studies exploring the relationship between specific genetic variations and susceptibility to T2DM.12,13 However, the mechanisms through which these genetic variations influence pancreatic β-cell function and insulin secretion require further investigation and research.13,14 Several single-nucleotide polymorphisms (SNPS) have been identified as causal factors for T2DM.15,16 SNPS affecting genes related to glucose metabolism, β-cell function, and insulin receptors are particularly significant in this context. One gene that plays a crucial role in T2DM susceptibility is SLC30A8, the gene coding for zinc transporter 8 (Znt8).17,18 Znt 8, a novel member of the zinc transporter family, is predominantly expressed in pancreatic β-cells and is responsible for maintaining zinc homeostasis and facilitating the efflux of zinc from the cytoplasm into intracellular vesicles.19,20 Znt8 is vital for insulin production, storage, and biological availability.21,22 Variants in the SLC30A8 gene, such as those located at rs13266634 and rs16889462 loci, have been shown to affect pancreatic β-cell function and increase susceptibility to T2DM.23,24,25 The C/T variant of rs13266634 (arginine to tryptophan at position 325, R325W) has been associated with T2DM in several ethnic groups.26,27,28 Genome-wide association studies (GWAS) have established a direct link between the SLC30A8 rs13266634 polymorphism and T2DM across multiple populations.29,30,31 The precise association between the SLC30A8 rs13266634 SNPS and their directional impact on T2DM risk remains critical.15,32,33 Our study is designed to analyse the genotype and allelic frequencies of the SLC30A8 (rs13266634 C/T) in northern Jordan. This study explores the relationship between the SLC30A8 (rs13266634 C/T) and T2DM in northern Jordan, as recent genetic studies have confirmed the association of this locus with T2DM.15,33,34 The study findings could clarify and identify the genetic risk factors for T2DM in northern Jordan and contribute to the extensive field of diabetes research by providing a comprehensive overview of SLC30A8 polymorphisms in the rising incidence of T2DM among the Jordanian population.33,33,34
Materials and Methods
Study Design and Subjects
This case-control study was conducted on 406 adults, comprising 186 (45.8%) males and 220 (54.2%) females, from April to June 2018, aged between 35 and 53 years. The study included 201 T2DM patients (101 males and 100 females) and 205 non-diabetic healthy controls (85 males and 120 females). Diabetes was defined according to the American Diabetes Association (ADA) criteria. T2DM individuals were recruited if their diagnosis was confirmed by fasting blood sugar and HbA1c, and they were receiving antidiabetic medications. The T2DM participants were sourced from those referred to the diabetic clinic at Prince Basma Hospital in Irbid, Jordan. The control group consisted of volunteers who met the inclusion criteria: no family history of diabetes and fasting blood sugar levels below 100 mg/dl. Body Mass Index (BMI) was determined by measuring weight and height and dividing the weight (kg) by the height squared (m). All protocols were reviewed and approved by the Institutional Review Board Council at Yarmouk University (Irbid, Jordan) IRB committee (IRB/2025/231). Patients and volunteers from the control group involved in the study were informed about its purpose and significance. Participants’ rights to confidentiality, privacy, and safety were assured. The collected information was used exclusively for research purposes and remained accessible only to the researchers involved in the study. Written informed consent was obtained from all participants after thoroughly addressing their questions and concerns.
Sample Collection and Biochemical Analysis
Peripheral blood was obtained from each participant who was fasting for 12 hours. The collected blood was divided into two portions: 5 ml in a plain tube and 5 ml in an EDTA-containing tube. The blood of the plain tube was left to clot before being centrifuged at 5000 rpm for 8 minutes to obtain serum for fasting blood sugar and biochemical tests. HbA1c was carried out on EDTA-containing blood using an enzymatic immunoassay method
Molecular Analysis (Genotype Analysis)
DNA of each participant was extracted from blood samples using the Wizard Genomic DNA purification kit (Promega, Wisconsin, USA). The quality of the extracted DNA was verified using electrophoresis on a 1.5% agarose gel under UV light. The extracted DNA was kept in the refrigerator at -70°C for molecular biology study. The SLC30A8 polymorphism (rs13266634 C/T) was analysed using polymerase chain reaction (PCR) with well-designed primers based on the SLC30A8 gene available on Genebank data (GenBank Accession Number Z14065.1). Primer designs were developed by Microsynth AG (Schützenstrasse, Balgach, Switzerland). The sequence of primers was as follows: Forward, 5’-GGACAGAAAGAGTTCCCATAGCG-3’; Reverse, 5’ ATAGCAGCATGTTTGAAGGTGGC-3’. The PCR cycling conditions were as follows: initial denaturation for 5 minutes at 95 °c. After that, 35 cycles consist of denaturation at 95°C for 30 seconds, annealing for 30 seconds at 67°C, and extension for 1 minute at 72°C. After completing the 35 cycles, extension for 5 minutes at 72°C is performed to finalize the PCR reaction. The volume of the PCR mixture with a total of 25 µL consists of 1 µL of GoTaq DNA polymerase (Promega, USA), 2 µL of dNTPs, 2 µL of 0.5 M of each primer, 5 µL of 10X buffer, 3 µL of genomic DNA, and 10 µL of nuclease-free water. PCR products were quantified using agarose gel electrophoresis (1.5% w/v) to ensure the generation of the DNA fragment of 429 bp. The PCR products were subjected to restriction fragment length polymorphism (PCR-RFLP) analysis. The PCR products were subjected to MspI restriction enzyme (Fermentas, Germany) at 37°C for 15 minutes. The digested products were then resolved on a 1.5 % agarose gel in Tris-Borate-EDTA buffer. The RFLP patterns were determined by estimating the size of each fragment on the electrophoresis gel to determine the genotype for each participant.
Statistical Analysis
Statistical analysis was performed to evaluate the associations between SLC30A8 gene polymorphism (rs13266634 C/T) and T2DM; a p-value of less than 0.05 was considered statistically significant. The genotype distribution in the diabetic and nondiabetic control groups was obtained using a chi-squared test for Independence. The statistical analyses were done using the SPSS for Windows statistical software package (SPSS Inc., Chicago, IL, USA, 2001.35 The student ‘t’ test was employed to evaluate the differences in means between the two studied groups.
Results
This study includes 406 adults (185 males and 221 females). The T2DM group consisted of 201 patients (101 females and 100 males), while the nondiabetic control group included 205 individuals (85 males and 120 females), as shown in Table 1. The genotypic and allelic frequencies of the SLC30A8 gene (rs13266634 C/T) among the population of North Jordan revealed the following: The CC genotype of rs13266634 (wild type), characterized by two fragments of 234 bp and 195 bp, was observed at 7.1%. Heterozygous CT genotype, representing 87.7% and characterized by the following fragments: 429 bp, 234 bp, and 195bp, However, homozygote TT genotype revealed a 429 bp single RFLP fragment with 5.2% frequency (Figure 1). These results show that the CT genotype is the most prevalent in this population, followed by the TT genotype, with the CC genotype being the least common. The allelic frequencies obtained in our study population were 51% and 49% for the C allele and T allele, respectively. The allelic frequencies suggest a slightly higher prevalence of the T allele compared to the C allele.
Table 1: Clinical and demographic parameters for the T2DM and nondiabetic control group.
|
Variable |
T2DM Group (n = 201) |
Nondiabetic control Group (n = 205) |
|
Age |
49.0 ± 4.6 |
43.5 ± 5.6 |
|
Male |
100 ± 5 |
85 ± 5 |
|
Female |
101 |
120 |
|
BMI |
31.4 ± 5 |
29 ± 3.2 |
|
HbA1c (%) |
7.85 ± 1.8 |
5.2 ± 0.4 |
p > 0.05
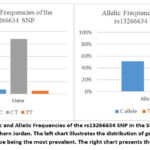 |
Figure 1: Genotypic and Allelic Frequencies of the rs13266634 SNP in the SLC30A8 Gene in the studied group, Northern Jordan. The left chart illustrates the distribution of genotypes (CC, CT, TT), with the CT genotype being the most prevalent. The right chart presents the allelic frequencies.
|
The distribution of SLC30A8 genotypes (CC, CT, TT) was analysed in both T2DM and healthy control groups to assess the association of these genotypes with T2DM in North Jordan (Table 2). The analysis revealed that the CT genotype was the most prevalent in T2DM and nondiabetic control groups, accounting for 84% of the T2DM group and 91.2% of the nondiabetic control group (Table 2). The TT genotype was the least common, observed in only 5.5% of the T2DM studied group and 4.9% of the nondiabetic studied group. The CC genotype was 10.5% for the T2DM study group and 3.9% for the nondiabetic group. Statistically, no significant difference was revealed between the T2DM study group and the nondiabetic control group (p>0.05). Our findings suggest that the SLC30A8 rs13266634 polymorphism may not contribute to the development of T2DM and is not considered a risk factor for T2DM in North Jordan.
Table 2: Genotype Distribution and Allele Frequencies in T2DM and Nondiabetic Control Groups
|
Genotype and Allele Frequencies |
Diabetic Group (n = 201) |
Nondiabetic control Group (n = 205) |
|
CC |
21 (10.5%) |
8 (3.9%) |
|
CT |
169 (84%) |
187 (91.2%) |
|
TT |
11 (5.5%) |
10 (4.9%) |
|
C |
211 (52.5%) |
203 (49.5%) |
|
T |
191 (47.5%) |
207 (50.5%) |
The study also examined the distribution of SLC30A8 genotypes across genders within the T2DM group to determine if gender influences the genetic predisposition of T2DM. The results indicated that the CT genotype is the most prevalent in males and females (Table 3). The SLC30A8 Genotype distribution among males and females reveals that the T2DM genetic risk is not associated with gender. HbA1c levels were compared across different SLC30A8 genotypes. A significant difference in HbA1c level was found when comparing the T2DM group to the Nondiabetic Control Group. The mean HbA1c level was slightly higher in individuals with the CT genotype (7.64%) compared to those with the CC genotype (7.26%). However, the difference was not statistically significant (p = 0.461), indicating that the SLC30A8 genotypes do not substantially affect HbA1c levels.
Table 3: Genotype distribution by gender in the diabetic and nondiabetic control Group.
|
Genotype |
T2DM Group |
Nondiabetic Control Group |
||
|
|
Male (n = 86) |
Females (n = 115) |
Males (n = 85) |
Females (n = 120) |
|
CC |
9 (10.5%) |
12 (10.4%) |
2 (2.3%) |
6 (5%) |
|
CT |
73 (84.9%) |
96 (83.5%) |
79 (93%) |
108 (90%) |
|
TT |
4 (4.6%) |
7 (6.1%) |
4 (4.7%) |
6 (5%) |
p > 0.05
Discussion
Certain genetic variants and their associated polymorphisms have been found to affect the development of T2DM. Such genetic variants disrupt the expression of their corresponding phenotypes, impacting the presentation and severity of T2DM.35,36 T2DM pathogenesis could be linked to genetic variants that affect insulin secretion and bioavailability.35,36 Genetic variants could gain-of-function mutations and abandon the encoded enzyme and protein associated with T2DM.37,38 Genotyping data reported a risk variant associated with T2DM, setting a specific allele that failed to encode functional proteins as a risk allele.39,40 Many alleles of the SLC30A8 gene, such as the rs13266634 C allele, were correlated with increased susceptibility and development of T2DM. However, generated protein models of the wild-type ZnT-8 transporter conclude that the Arg325Trp variant of the rs13266634 polymorphism could tolerate sufficient bioavailable zinc to pancreatic β-cells.35,42
Previous studies in various populations revealed that the C/T polymorphism of rs13266634 directly affects T2DM development.29,30,31,32 The genotype distribution of the SLC30A8 (rs13266634 C/T) in Northern Jordan indicated that the CT genotype was more prevalent among both T2DM and healthy control groups. These findings align with results obtained from other studies in different countries, confirming the prevalence of the CT genotype in T2DM and nondiabetic control groups.29,31,43 In contrast, the CC genotype was more common in T2DM and nondiabetic control groups in Iran.42 The results demonstrate that the observed differences in genotype distribution and allele frequencies were not statistically significant, suggesting no meaningful association between the SLC30A8 rs13266634 C/T polymorphism and the development of T2DM in Northern Jordan. These findings corroborate previous studies conducted across various populations, indicating that the rs13266634 C/T polymorphism of the SLC30A8 gene was not significantly associated with an increased risk of T2DM.40,44,45,46 The lack of significant association between the SLC30A8 rs13266634 genotype and T2DM was contradicted by prior studies that showed a significant association between the SLC30A8 (rs13266634) genotype and T2DM.34,43,48,49 According to genome-wide association studies, polymorphisms of SLC30A8 rs13266634 were reported to be associated with T2DM, supported by significant risk allele-specific odds ratios (ORs) and confidence intervals (95% CI).49,50,51 Meta-analyses have also confirmed the association between T2DM and the SLC30A8 rs13266634 polymorphism.15,29,30 In some instances, a meta-analysis study failed to prove the association between T2DM and SLC30A8 (rs13266634) polymorphism, as found in a case-control study within the same population.47 A significant association between the C/T rs13266634 SNP of SLC30A8 and T2DM was identified in a case-control study conducted in Jordan.34 The distribution of the CC genotype versus the TT genotype among T2DM in our study mirrors their results.34 The contradictory results found in the two case-control studies conducted in Jordan may be attributed to the differences in the population composition of each study.3 Our study was conducted in Northern Jordan, whereas the study that found a significant association between the C/T rs13266634 SNP of SLC30A8 and T2DM was performed in the Amman district in the center of the kingdom, which possesses a highly diverse population undergoing dynamic transformation and vigorous changes. The SLC30A8 (rs13266634 C/T) genotype distribution among the T2DM study group and the nondiabetic control group shows no significant difference between the two studied groups in Northern Jordan. The lack of a significant association of the rs13266634 C/T polymorphism of the SLC30A8 gene with T2DM revealed by our study is a noteworthy finding that warrants investigation of other risk factors associated with T2DM among North Jordanians. Our findings regarding the allele distribution contradict those from other populations, where the high frequency of the C allele commonly represents the majority.39,44,46,49 The high frequency of the risk allele (C) has been reported in the African-American community (91.5 %), as well as in Moroccan (83.5 %) and Tunisian (82.5 %) populations.27,40,44 The C allele frequency of rs13266634 in our study (51%) was lower than that of other populations, such as Eastern Asians. Similar results regarding the C allele frequency reported in our study were also found for the Hunan district population in China.43,47 The C allele frequency of rs13266634 among the Hong Kong and Japanese populations was 55.2% and 57.9 %, respectively.47,52 The T allele frequency revealed in our study (49%) was consistent with the findings of a limited number of populations, yet contradicted the findings from many others. The highest frequency for the T allele was detected in African populations, whereas the T allele frequency is low in the Middle East, with the lowest frequency noted in East Asia. The data concerning T allele frequency and C allele frequency obtained in our study, along with the genotype distribution and the negative association between T2DM and genotype distribution, confirm the emerging new trend of cumulative effects and interactions of different risk alleles in the development of T2DM.
The SLC30A8 (rs13266634 C/T) genotype distribution by gender revealed that both males and females predominantly carried the CT genotype. The distribution of the CC genotype was relatively equal between genders, with no significant gender differences in genotype distribution. This observation is consistent with the findings from other studies, which reported that gender does not significantly influence the distribution of SLC30A8 (rs13266634 C/T).53,54 In addition to genotype frequency and distribution, the study analysed the relationship between SLC30A8 genotypes and HbA1c levels. It was found that while the CT genotype was associated with slightly higher HbA1c levels compared to the CC genotype, this association was not significant. These findings regarding the association between SLC30A8 genotypes and HbA1c levels do not substantially affect the glycemic control in the study population. Similar findings regarding the SLC30A8 genotypes and HbA1c levels have been reported, where no significant association between SLC30A8 polymorphisms and HbA1c levels appears.15,48,49
Considering the value of our findings, it is important to realise that our research was limited by certain constraints, such as the case-control design, which may not reflect the rapid transformation and changes in the population over time, and the potential impact of unmeasured factors. Future research, especially association studies and interventional designs, will be critical in fixing this limitation and becoming more conclusive. However, the eventual nature of our study design safeguards reliability and consistency, providing a snapshot of the current condition of the study population to identify further SNPS that may be associated with T2DM.
Conclusion
Our study findings revealed that the SLC30A8 gene polymorphism (rs13266634 C/T) does not have a significant impact on the development of T2DM in the North Jordanian population. No significant differences were found between the T2DM study group and the nondiabetic control group concerning genotype distribution. Based on that, future research in the North Jordanian population should explore other genetic factors or potential gene-environment interactions that may contribute to the development of T2DM. Additionally, further studies could investigate large and more diverse populations to confirm these findings and potentially uncover other genetic markers associated with T2DM risk.
Acknowledgement
The authors would like to thank Yarmouk University for granting Master degree research work. The Department of Biological Sciences, Faculty of Science, Yarmouk University, is highly appreciated for their kind willingness to conduct scientific research.
Funding Sources
This research was supported by the Deanship of Scientific Research and Graduate Studies at Yarmouk University. Financial support project No. 51/2020.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This trial is registered at the Institutional Review Board Council at Yarmouk University (Irbid, Jordan) IRB committee (YUIRB DSR 2020/63).
Permission to reproduce material from other sources
Not Applicable
Author Contributions
- Khalid Abu Khadra: Conceptualisation, Funding Acquisition, Methodology, Data collection, and Writing – Original Draft.
- Mohammad Wahsha: Data analysis, draft preparation, Review, & Editing.
- Tariq Al-Najjar: Methodology, Statistical analysis, draft preparation, Writing – Review and Editing.
References
- International Diabetes Federation. IDF Diabetes Atlas. Ninth edition 2019. https://www.diabetesatlas.org/ upload/resources/2019/ IDF_Atlas_9th_Edition_2019.pdf. Accessed: 24 Nov 2019.
- Matsuda M., and Shimomura I. Increased oxidative stress in obesity: implications for metabolic syndrome, diabetes, hypertension, dyslipidemia, atherosclerosis, and cancer. Obes Res Clin Pract. 2013; 7(5): e330-41. DOI: 1016/j.orcp.2013.05.004.
CrossRef - Abu Khadra K, Bataineh M, Khalil A, Salah J, Oxidative stress and type 2 diabetes: the development and the pathogenesis, Jordanian cross-sectional study. Eur J Med Res.2024;29, 370. https://doi.org/10.1186/s40001-024-01906-4.
CrossRef - Elbein S, Ma J, Genetic variants associated with type 2 diabetes and obesity. Current Diab Rep. 2006; 6 (5), 399-406. https://doi.org/10.1007/s11892-006-0054-8.
- Mccarthy I, Froguel P, Genetic approaches to the molecular understanding of type 2 Am J Physio Endo Meta. 2002;283 (2) 17-25. DOI: 1152/ajpendo.00099.2002
CrossRef - Kavvoura K, Pappa M, Evangelou E, Ntzani E. The genetic architecture of type 2 diabetes pharmacotherapy: the emerging genomic evidence. Curr Pharm Des. 2014;20(22):3610-9. doi: 10.2174/13816128113196660675.
CrossRef - Lyssenko V, Jonsson A, Almgren P, et al. Clinical risk factors, DNA variants, and the development of type 2 diabetes. N. Engl. J. Med. 2009;360,1360–1361. https://doi.org/10.1056/NEJMc082624.
CrossRef - Ntzani E, Kavvoura K. Genetic risk factors for type 2 diabetes: insights from the emerging genomic evidence. Curr Vasc Pharmacol. 2012;10(2):147-55. doi: 10.2174/157016112799305030.
CrossRef - Wild S, Roglic G, Green A, Sicree R, King H. Global prevalence of diabetes: estimates for the year 2000 and projections for 2030. Diabetes Care. 2004;27 (5):1047-53. doi: 10.2337/diacare.27.5.1047.
CrossRef - Abuyassin B, Laher I. Diabetes epidemic sweeping the Arab world. World J. Diabetes 2016;7(165). https://doi.org/10.4239/wjd.v7.i8.165.
CrossRef - Awad F, Huangfu P, Dargham R, Ajlouni K, Batieha A, Khader Y, Critchley J, Abu‑Raddad J. Characterizing the type 2 diabetes mellitus epidemic in Jordan up to 2050. Scientific Reports 2020; 10:21001.
CrossRef - Williams L, Jacobs B, Moreno-Macías H, et al. Sequence variants in SLC16A11 are a common risk factor for type 2 diabetes in Mexico. Nature. 2014;506(7486):97-101. doi: 10.1038/nature12828.
CrossRef - Walford A, Green T, Neale B, et al. Common genetic variants differentially influence the transition from clinically defined states of fasting glucose metabolism. Diabetologia. 2012;55(2):331-339. doi:10.1007/s00125-011-2353-8.
CrossRef - Spranger, J., Kroke, A., Möhlig, M., Bergmann, M. M., Ristow, M., Pfeiffer, A. F., & Boeing, H. The role of genetic variants in type 2 diabetes. J of Clin End & Meta. 2004; 89(4):1833-1838. https://doi.org/10.1210/jc.2003-031059.
- Dupuis J, Langenberg C, Prokopenko I, et al. Diabetes Genetics Replication and Meta-analysis Consortium (DIAGRAM). New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nature Genetics. 2010;42(2): 105-116. https://doi.org/10.1038/ng.520.
CrossRef - Pascoe L, Tura A, Patel K, et al. Common variants of the novel type 2 diabetes genes CDKAL1 and HHEX/IDE are associated with decreased pancreatic beta-cell function. Diabetes 2007;56(12); 3101-3104. https://doi.org/10.2337/db07-0634.
CrossRef - Hu M, Kim I, Morán I, et al. Multiple genetic variants at the SLC30A8 locus affect local super-enhancer activity and influence pancreatic β-cell survival and function. The FASEB journal. 2024;30(38). https://doi.org/ 10.1096/ fj.202301700RR
CrossRef - Jing L, Sun M, Bi Y, Shen M, Zhu L. SLC30A8 polymorphism and type 2 diabetes risk: evidence from 27 study groups. Nutr Metab Cardiovasc Dis. 2011;21(6):398-405. doi:10.1016/j.numecd. 2009.11.00410.1016/ j.numecd.2009.11.004 14.
CrossRef - Huang Q, Du J, Merriman C, Gong Z, Genetic, functional, and immunological study of ZnT8 in diabetes. Int J Endo, 2019; Feb 26;2019: https://doi.org/10.1155/2019/1524905.
CrossRef - Chimienti F, Devergnas S, Favier A, Seve M, Identification and cloning of a beta-cell-specific zinc transporter, ZnT-8, localized into insulin secretory granules. 2004;53(9):2330–2337.
CrossRef - Chimienti F, Devergnas S, Pattou F, et al. In vivoexpression and functional characterization of the zinc transporter ZnT8 in glucose-induced insulin secretion. J Cell Sci. 2006;119 (20):4199–4206. doi: 10.1242/jcs.03164.
CrossRef - Lemaire K, Ravier A, Schraenen A, et al. . Insulin crystallization depends on zinc transporter ZnT8 expression, but is not required for normal glucose homeostasis in mice. Proc Natl Acad Sci U S A. 2009;106(35):14872-7. doi:10.1073/pnas.0906587106.
CrossRef - Xu J, Wang J, Chen B, SLC30A8 (ZnT8) variations and type 2 diabetes in the Chinese Han population. Genet Mol Res. 2012;11(2):1592-1598. doi:10.4238/2012.May.24.1.
CrossRef - Rutter A, Chimienti F, SLC30A8 mutations in type 2 diabetes. Diabetologia. 2015; 58(1):31-36. doi:10.1007/s00125-014-3405-7 5.
CrossRef - Tamaki M, Fujitani Y, Hara A, et al. The diabetes-susceptible gene SLC30A8/ZnT8 regulates hepatic insulin clearance. J Clin Invest. 2013;123:4513–24.
CrossRef - Billings K, Jablonski A, Ackerman J, et al. The influence of rare genetic variation in SLC30A8 on diabetes incidence and β-cell function. J Clin Endocrinol Metab. 2014;99(5):E926-30. doi: 10.1210/jc.2013-2378. Epub 2014 Jan 28.
CrossRef - Kifagi C, Makni K, Boudawara M, et al. Association of genetic variations in TCF7L2, SLC30A8, HHEX, LOC387761, and EXT2 with Type 2 diabetes mellitus in Tunisia. Genet Test Mol Biomarkers. 2011;15(6):399-405. doi: 10.1089/gtmb.2010.0199.
CrossRef - Boesgaard W, Zilinskaite J, Vänttinen M, et al. The common SLC30A8 Arg325Trp variant is associated with reduced first-phase insulin release in 846 non-diabetic offspring of type 2 diabetes patients—the EUGENE2 study. Diabetologia. 2008;51(5), 816-820. https://doi.org/10.1007/s00125-008-0955-6.
CrossRef - Cheng L, Zhang D, Zhou L, Zhao J, Chen B. Association between SLC30A8 rs13266634 Polymorphism and Type 2 Diabetes Risk: A Meta-Analysis. Med Sci Monit. 2015;27;21:2178-89. doi: 10.12659/MSM.894052.
CrossRef - Tan T, Ng P, Nurbaya S, et al. Polymorphisms identified through genome-wide association studies and their associations with type 2 diabetes in Chinese, Malays, and Asian-Indians in Singapore. J Clin Endocrinol Metab. 2010;95(1):390-7. doi: 10.1210/jc.2009-0688.
CrossRef - Sladek R, Rocheleau G, Rung J, et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature. 2007;445(7130):881-885. https://doi.org/10.1038/nature05530.
CrossRef - Cauchi S, Meyre D, Durand E, et al. Post genome-wide association studies of novel genes associated with type 2 diabetes show gene–gene interaction and high predictive value. PLoS One 2008;3:2031–2041.
CrossRef - Hu C, Zhang R, Wang C, et al. PPARG, KCNJ11, CDKAL1, CDKN2A-CDKN2B, IDE-KIF11- HHEX, IGF2BP2 and SLC30A8 are associated with type 2 diabetes in a chinese population. PLoS One 4. 2009;4(10) https://doi.org/10.1371/journal.pone.0007643.
CrossRef - Mashal S, Khanfar M, Al-Khalayfa S, et al. SLC30A8 gene polymorphism rs13266634 associated with increased risk for developing type 2 diabetes mellitus in Jordanian population. Gene. 2021;5;768:145279. doi: 10.1016/j.gene.2020.145279.
CrossRef - Statistical Package for Social Sciences Program (SPSS) for Windows; v.17, SPSS Inc., Chicago. USA, 2008.
- Weijers M, Three-dimensional structure of b-cellspecific zinc transporter, ZnT8, predicted from the type 2 diabetes: associated gene variant SLC30A8 R325W. Diabetol Metab Syndr, 2010;2:33–40.
CrossRef - Mtiraoui N, Turki A, Nemr R, et al. Contribution of common variants of ENPP1, IGF2BP2, KCNJ11, MLXIPL, PPARγ, SLC30A8 and TCF7L2 to the risk of type 2 diabetes in Lebanese and Tunisian Arabs. Diabetes Metab. 2012; 38(5):444-9. doi:10.1016/j. diabet.2012.05.002 9.
CrossRef - Cotsapas C, Prokunina-Olsson L, Welch C, et al. Expression analysis of loci associated with type 2 diabetes in human tissues. 2010; 53(11):2334-2339. doi:10.1007/s00125-010-1861-2
CrossRef - Chauhan G, Spurgeon J, Tabassum R, et al. Impact of common variants of PPARG, KCNJ11, TCF7L2, SLC30A8, HHEX, CDKN2A, IGF2BP2, and CDKAL1 on the risk of type 2 diabetes in 5,164 Indians. Diabetes 2010;59:2068–2074.
CrossRef - Mohaddes M, Karami F, Gharesouran J, Bahrami A, The soluble carrier 30 A8 (SLC30A8) gene polymorphism and risk of diabetes mellitus type 2 in Eastern Azerbijan population of Iran. J Sci Islam Repub Iran .2012;23:15–20.
- Xu K, Zha M, Wu X, et al. Association between rs13266634 C/T polymorphisms of solute carrier family 30 member 8 (SLC30A8) and type 2 diabetes, impaired glucose tolerance, type 1 diabetes–a meta-analysis. Diabetes Res Clin Pract. 2011;91(2):195-202. doi:10.1016/j. diabres.2010.11.012 13.
CrossRef - Faghih H, Khatami R, Azarpira N, Foroughmand M, SLC30A8 gene polymorphism (rs13266634 C/T) and type 2 diabetes mellitus in south Iranian population. Biol. Rep. 2014;41:2709–2715. https://doi.org/10.1007/ s11033-014-3158-x.
CrossRef - Huang Q, Yin Y, Dai P, et al. Association analysis of SLC30A8 rs13266634 and rs16889462 polymorphisms with type 2 diabetes mellitus and repaglinide response in Chinese patients. J. Clin. Pharm. 2010;66:1207–1215. https://doi.org/10.1007/s00228-010-0882-6.
CrossRef - Lewis P, Palmer D, Hicks P, et al. Association analysis in African Americans of european-derived type 2 diabetes single nucleotide polymorphisms from whole-genome association studies. Diabetes. 2008;57:2220–2225. https://doi.org/10.2337/db07-1319.
CrossRef - Rong R, Hanson RL, Ortiz D, et al. Association analysis of variation in/near FTO, CDKAL1, SLC30A8, HHEX, EXT2, IGF2BP2, LOC387761, and CDKN2B with type 2 diabetes and related quantitative traits in Pima Indians. Diabetes 2009;58:478–488.
CrossRef - Sanghera K, Ortega L, Han S, et al.) Impact of nine common type 2 diabetes risk polymorphisms in Asian Indian Sikhs: PPARG2 (Pro12Ala), IGF2BP2, TCF7L2 and FTO variants confer a significant risk. BMC Med Genet 2008;9:59–67.
CrossRef - Tabara Y, Osawa H, Kawamoto R, et al. Replication study of candidate genes associated with type 2 diabetes based on genome-wide screening. Diabetes. 2009; 58:493– 498.
CrossRef - Lee H, Kang E, Kim H, et al. Association between polymorphisms in SLC30A8, HHEX, CDKN2A/B, IGF2BP2, FTO, WFS1, CDKAL1, KCNQ1 and type 2 diabetes in the Korean population. Hum. Genet. 2008;53(11-12), 991-998. https://doi.org/10.1007/s10038-008-0341-8.
CrossRef - Han X, Luo Y, Ren Q, et al. Implication of genetic variants near SLC30A8, HHEX, CDKAL1, CDKN2A/ B, IGF2BP2, FTO, TCF2, KCNQ1, and WFS1 in type 2 diabetes in a Chinese population. BMC Med Genet 12010;1:81–89.
- Hertel K, Johansson S, Raeder H, et al. Genetic analysis of recently identified type 2 diabetes loci in 1,638 unselected patients with type 2 diabetes and 1858 control participants from a Norwegian population based cohort (the HUNT study). Diabetologia 2008;51:971–977
CrossRef - Potapov A, Chistiakov, A, Shamkhalova, S, Shestakova V, Nosikov V, TCF7L2 rs12255372 and SLC30A8 rs13266634 confer susceptibility to type 2 diabetes in a Russian population. Diabetes Metab. Syndr. Clin. Res. Rev. 2009;3:219–223. https://doi.org/10.1016/j.dsx.2009.07.005.
CrossRef - Ng M, Park K, Oh B, et al. Implication of genetic variants near TCF7L2, SLC30A8, HHEX, CDKAL1, CDKN2A/B, IGF2BP2, and FTO in type 2 diabetes and obesity in 6,719 Asians. Diabetes 2008;57:2226–2233.
CrossRef - Saxena R, Voight B, Lyssenko V, et al. Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. 2007;316(5829):1331-1336. doi: 10.1126/science.1142358.
CrossRef - Turki A, Al-Zaben G, Khirallah M, et al. Gender-dependent associations of CDKN2A/2B, KCNJ11, POLI, SLC30A8, and TCF7L2 variants with type 2 diabetes in (North African) Tunisian Arabs. Diabetes Res Clin Pract. 2014;103:e40–43. doi: 10.1016/j.diabres.2013.12.040.
CrossRef







