Manuscript accepted on :12-02-2026
Published online on: 06-05-2026
Plagiarism Check: Yes
Reviewed by: Dr. Yerbolat Iztleuov
Second Review by: Dr. Karthikeyan
Final Approval by: Dr. Anton R Keslav
Sandeep Kumar Sunori1* , Shilpa Jain2
, Shilpa Jain2 , Govind Singh Jethi2
, Govind Singh Jethi2 and Pradeep Juneja1
and Pradeep Juneja1
1Department of ECE, Graphic Era Hill University, Bhimtal Campus, India,
2Department of CSE, Graphic Era Hill University, Bhimtal Campus, India
Corresponding Author E-mail:sksunori@gehu.ac.in
Abstract
The stages of chronic infection in hepatitis C Virus (HCV) include Hepatitis and Fibrosis, followed by Cirrhosis, and staging of the diseases must be non-invasive to be effectively used in clinical practices. This research article creates a powerful computational algorithm of multi-class HCV staging with standard serum laboratory biomarkers.A dataset of 12 clinical biomarkers and demographics of 615 subjects has been used. In connection to the intrinsic correlation and high dimensionality of the biomarker panel, Principal Component Analysis (PCA) was used as an essential step in feature engineering as it retained 95% of total data variance. Three supervised machine learning classifiers, Naive Bayes (NB), K-Nearest Neighbors (KNN, k=5) and a multi-class Support Vector Machine (SVM) based on the Error-Correcting Output Codes (ECOC) wrapper with a linear kernel,were trained and compared on the optimal low-dimension set of features obtained through PCA. The SVM-ECOC model has shown better overall predictive performance (highest Accuracy 91 %), Macro-Averaged Precision (0.745) and Macro-Averaged Recall (Sensitivity) of 0.61. The translational usefulness of the SVM model was further validated by further rigorous clinical validation using the Multi-Category Net Reclassification Improvement (MCNRI) measure, which reported a net improvement in proper risk stratification of 8.13 % over Naive Bayes and 4.88 % over K-Nearest Neighbors. This performance justifies the feasibility of PCA in reducing multidimensional biological data to a space of features that can be separated linearly, which boosts the success of classification tremendously. Nevertheless, another significant limitation of the study is pointed out, the difference between the high overall accuracy and moderate Macro-Averaged Recall indicates the insensitivity (high False Negative Rate) of the key minority disease types (Hepatitis, Fibrosis, Cirrhosis) because of the imbalance in the dataset. All models have been simulated on MATLAB. Research in future should focus on the application of data-level methods, such as oversampling, to reduce the bias of the class and determine ethically acceptable, reliable diagnostic sensitivity at all phases of HCV development to be clinically applicable.
Keywords
HCV (Hepatitis C Virus); Liver Fibrosis; MCNRI (Multi-Category Net Reclassification Improvement); Multi-class Classification; PCA (Principal Component Analysis); Predictive modelling; SVM (Support Vector Machine); Serum Biomarkers
| Copy the following to cite this article: Sunori S. K, Jain S, Jethi G. S, Juneja P. Predictive Modelling of Hepatitis C Virus Disease Progression Using PCA and Machine Learning. Biomed Pharmacol J 2026;19(2). |
| Copy the following to cite this URL: Sunori S. K, Jain S, Jethi G. S, Juneja P. Predictive Modelling of Hepatitis C Virus Disease Progression Using PCA and Machine Learning. Biomed Pharmacol J 2026;19(2). Available from: https://bit.ly/42RjzAm |
Introduction
Background on Hepatitis C Virus (HCV) and Disease Progression
Hepatitis C Virus (HCV) infection is a persistent and major challenge to the global population health, mainly because of the high rate of chronicity and further manifestation into the severe liver disease.16Clinical course of chronic HCV is a continuum of pathology, which progresses in order of acute and chronic hepatitis to appearance of liver fibrosis, and finally, cirrhosis. Cirrhosis is an irreversible disorder characterized by massive hepatic scarring and functional dysfunction, which is a severe risk factor contributing to life-threatening conditions and events such as hepatocellular carcinoma and end-stage liver failure. The classic techniques of measuring the extent of liver injury, especially the level of fibrosis and cirrhosis, were based on the invasive liver biopsy. Although traditionally viewed as the gold standard of diagnosis, the biopsy procedure suffers because of its expensive nature, complications, sampling error, and related patient discomfort.17 Such significant weaknesses demand the creation and justification of powerful, non-invasive substitutes.
The Role of Serum Biomarkers in Non-Invasive Diagnosis
Modern non-invasive staging is based on the interpretation of easily available serum biomarkers or Liver Function Tests (LFTs) and related panels. The dataset25 that the research examined in this paper includes a set of ten important laboratory biomarkers-Albumin (ALB), Alkaline Phosphatase (ALP), Alanine Aminotransferase (ALT), Aspartate Aminotransferase (AST), Bilirubin (BIL), Cholinesterase (CHE), Cholesterol (CHOL), Creatinine (CREA), Gamma-Glutamyl Transferase (GGT) and Total Protein (PROT), in combination with demographic ones (Age, Sex), that represent the entire picture of the physiological health of the liver in combination.Although in the clinical sense, these biomarkers are informative, but when combined, their predictive strength is likely to be lost in complexity and high dimensionality in the data structure. As an example, AST and ALT frequently coincide in the occurrence of acute damage, and ALP is very likely to be related to GGT in the cholestatic pattern.18 In order to reliably identify the subtle biochemical patterns that define the five stages of the disease viz. Blood Donor, Suspect Donor, Hepatitis, Fibrosis, and Cirrhosis, there is a need to adopt the complex plans of computation that are able to manage these kinds of relationships and extract the latent patterns associated with the diagnosis.
Combining Dimensionality Reduction and Classification of Diagnostic Modelling
A high feature correlation and dimensionality would present significant challenges, which this study addressed with the help of Principal Component Analysis (PCA) as an upstream feature engineering approach. PCA is a linear transformation of the initial set of correlated biomarker features, which results in a new orthogonal coordinate system, where a set of Lower-dimensional Principal Components (PCs) captures the highest possible amount of original data variance.19 It is an essential dimension reduction to ensure the maximum amount of computational efficiency and avoid redundancy in the features presented to the classifiers.20
The given feature set after a PCA was then applied to the performance of three different supervised machine learning paradigms, namely, Naïve Bayes (NB), which is a probabilistic approach; K-Nearest Neighbors (KNN), which is an instance-based and non-parametric approach; and Support Vector Machine (SVM), which is a maximum-margin geometric classifier.21 A rigorous comparison of these algorithms is crucial to discovering the most appropriate model architecture to use in multi-class classification in the complicated environment of HCV disease staging.
Hypothesis and Objectives
The basic hypothesis upon which this research is based is that when the PCA is used in standardizing and dimensional reduction of the HCV biomarker data, it will produce a low-dimensional feature representation that will significantly improve the predictive capability of the supervised machine learning classifiers. Moreover, the SVM model is also expected to have high classification performance in predicting the different stages of HCV progression than NB and KNN because it has strong generalization abilities and geometrical separation.
The research objectives of this work are:
To use PCA and choose the best number of the principal components necessary to explain 95 per cent of total variance in the standardized biomarker data.
To train, tune and test the relative performance of the NB, KNN and the SVM models on the PCA-reduced set of features.
To ascertain the most effective classification algorithm in general in terms of a critical analysis of multi-class measures, namely Accuracy, Macro-Averaged Precision, Macro-Averaged Recall (Sensitivity) and F1-Score.
In order to conduct MCNRI analysis of the developed models.
Literature Review
Clinical Interpretation of Liver Biomarkers in HCV Staging
Patterns and ratios of LFTs have become essential in the interpretation of liver disease in clinical hepatology, and cannot be interpreted alone. ALT and AST are enzyme indicators of cellular integrity, and a high level of them characterizes hepatocellular damage, which is characteristic of hepatitis. An important discriminating test is the AST:ALT ratio (De Ritis ratio). The strong indication of alcoholic liver disease is a ratio of 2:1, which indicates high levels of AST in the mitochondria. On the other hand, during cases of cholestasis, a lower AST:ALT ratio of < 1.5 tend to indicate extrahepatic obstruction usually where the ALT is significantly higher than AST. Higher concentrations of Alkaline Phosphatase (ALP) and Gamma-Glutamyl Transferase (GGT), often followed by an increase of Bilirubin (BIL), is a cholestatic profile that is the result of the biliary tract obstruction. Enzyme induction that occurs due to chronic alcohol intake or certain medications is especially sensitive to GGT.18 Synthetic capacity markers, which are mainly Albumin (ALB), Total Protein (PROT) and Cholinesterase (CHE), decrease with the severity of the disease. Reduced levels of ALB is a strong indicator of chronic liver failure, which is normally accompanied by progressive Fibrosis and Cirrhosis. The HCV staging is well presented in the form of a flowchart as shown in Fig.1. This flowchart is a summary of natural history and clinical course of Hepatitis C. Once infected, the individuals go through the acute HCV phase (0-6 months) where 15-45 % of them can attain spontaneous viral clearance. Without clearance of the virus, infection turns into chronic HCV which causes progressive liver damage. The progression of the chronic infection is determined by the stages of the METAVIR fibrosis (F0-F4) of no fibrosis to cirrhosis. DAAs can be treated at any stage of fibrosis and usually leads to SVR (viral cure), stops the progression and could partially reverse fibrosis. When the fibrosis is further developed to F4, the patient already has cirrhosis that can be able to be compensated over a period of time and then degenerate into decompensated cirrhosis that is accompanied by jaundice, ascites, and hepatic encephalopathy. Out of decompensation, patients are predisposed to hepatocellular carcinoma (HCC) and end-stage liver disease (ESLD). Highly developed disease either HCC or ESLD can eventually necessitate liver transplantation.
Methodological Precedents for Dimensionality Reduction (PCA)
PCA is a framework method of high-dimensional (HD) clinicaldata analysis in health science applications, which facilitates the discovery of latent data and compression of computational requirements. The approach used in this case required a very strict inclusion criterion: the selection of a minimum of PCs that would explain 95 % of the cumulative variance. This 95 % threshold is very specific and it is selected in health informatics because it offers much better stability and reliability than less deterministic methodslike the Kaiser-Guttman criterion or Cattell’s Scree Test. The dimensionality reduction step is important in that it ensures that much of the variability in the original data (95 %) is retained, thus ensuring that important diagnostic information that pertains to subtle metabolic patterns, which are important in determining diseases stages, is retained, and agood methodological rigor is established.
Leveraging Advanced Diagnostic Techniques in Liver Disease
The growing use of computational intelligence, biomedical data analytics, and sophisticated diagnostic technology has changed modern studies of hepatitis viruses, liver disease progression, and classification sciences. At the methodological level, the accuracy of the classification results is highly determined by the choice of appropriate loss and accuracy measurement metrices especially in situations, which are influenced by a data imbalance, where a metric bias and misinterpretation can be a major determinant of the actual model performance.1 These issues have gained even greater importance due to the implementation of machine learning frameworks in the fields of virology, epidemiology and clinical decision support. Indicatively, to illustrate how computational methods can be used to discover mechanistic insights into the hepatitis C virus (HCV) biology, rotation-forest-based classifiers combining position-specific scoring matrices and two-dimensional PCA have been shown to yield high performance in human-viral protein interactions.2 Simultaneously, the individualized diagnostic modeling has become a highly promising direction, where patient-specific machine learning systems can achieve high diagnostic accuracy and recall of HCV compared to standardized generalized models.3
In addition to traditional statistical learning, new reasoning approaches have been suggested including neutrosophichypersoft mapping to resolve uncertainties in symptom assessment and treatment distribution that allow more consistent mapping of hepatitis symptoms and treatment decisions in clinically ambiguous scenarios.4 Ensemble learning methods have been also found to have a good predictive ability in differentiating between cases of hepatitis C and cirrhosis with some of the critical biomarkers being ALT and AST that have been found to be the major predictors.5 Other works have come up with multiclass HCV detection pipelines using random forest, logistic regression, and oversampling methodsto record better early diagnosis using imbalanced clinical data.6
Simultaneously with the development of models on a patient level, population-based modeling still serves as a significant aspect of the transmission of hepatitis viruses. Compartmental formulations determined deterministically have helped explain the HCV important epidemiological thresholds, including the basic reproductive number, by which HCV would be persistently controlled or eliminated within high-risk communities.7 In order to enable the simulation study of large-scale simulations, the agent-based models accelerated by the parallel sliding-region algorithms make country-level epidemic models based on real demographic data possible without paying a significant computational cost.8 However, systems-level studies based on protein interaction networks have offered new insights into the mechanisms of fibrosis formation, interference with immune pathways, and biomarker identification in hepatitis B and C.9
Machine learning has also been used in clinical risk stratification, where decision trees, regression models, genetic algorithms, and particle swarm optimization have demonstrated significant predictive power of advanced levels of fibrosis, particularly where large populations of patients are considered.10 In addition to these diagnostic innovations, research in the area of biosensing technologies has resulted in portable electrochemical systems that are able to perform HCV immunodetection in rapid and ultrasensitive modes, opening up the possibilities of decentralized and resource-limited screenings.11 Bayesian inference has continued to gain a significant role at the molecular level in defining drug-resistance mechanisms, especially the identification of resistance-associated mutations in the NS5A region of genotype 1a HCV virus.12 As well, more general principles of Bayesian methodology have been used to detect intricate mutations in hepatitis B, hepatitis C and HIV making it possible to gain a more precise picture of how high-dimensional genomic interactions can affect phenotypes.13
These statistical models are further augmented with sophisticated data-mining strategies. The complete HBV genome sequence studies have been performed using clustering, evolution, fuzzy-measure-based classification and information-gain-driven feature selection to determine the potential oncogenic markers in the development of hepatocellular carcinoma.14The development of pharmaceutical analytics including carbon-nanotube-enhanced electrochemical sensors have facilitated the sensitive detection of antiviral drugs like daclatasvir, which can be used to measure the correct dosage and treatment effects of antiviral drugs in HCV treatment .15Machine learning algorithms have also been used in generating decision trees from laboratory data.26
Collectively, these diverse streams of study point to the more multidisciplinary nature of the present-day landscape of the hepatitis virus research, in which the state of the art computational, statistical, epidemiological and biosensing advances intersect to increase disease knowledge, diagnosis, and optimized treatment. Table 1 is summarizing the performance of some existing ML models in hepatitis research.
Comparative research on the hepatology diagnostics topic usually demonstrates mixed outcomes, yet tends to conclude that the more sophisticated machine learning algorithms are effective. Of special interest is the comparison of Naive Bayes (NB) with SVM. NB uses an independence assumption of features, which usually is not the case in complex biological systems where biomarkers interact on a large scale. SVM on the other hand works by projecting data into a high dimensional space in order to detect the best maximum-margin separating hyperplane. Since non-linear interactions among biomarkers are most likely, the SVM framework theoretically stands in a better position to estimate the geometric distance between disease groups than the purely probabilistic nature of NB, and it is in agreement with the existing literature that SVM often yields better results on the more complex biomedical data sets.
In the five category multi-class staging problem with the HCV progression, the SVM was trained with the Error-Correcting Output Codes (ECOC) wrapper. ECOC is a complex method that transforms the multi-class problem of interest into multiple and strong binary classification sub-problems to boost the stability and the ability to generalize the resulting model. The line kernel used in the linear implementation of SVM with application of PCA effectively transforms the data so that the classes of the disease can be effectively separated linearly in the reduced feature space, a result commonly characterized by an effective feature engineering process and a linearly separable data set.
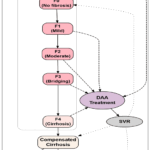 |
Figure 1: HCV staging |
Table 1: Summary of literature review on application of ML in hepatitis research
| Authors | Research Area | Model/Method | Results | Novelty |
| L. Chen et al.3 | HCV Diagnosis (Patient-Specific) | Customized ML Model | Over 99% accuracy and 94% recall | Demonstrates the strength of a patient-specific model over general- purpose model. |
| X. Liu et al.2 | HCV-Human PPI Prediction | RF-PSSM (Rotation Forest-Position-Specific Scoring Matrix) | Accuracy 93.74%; AUC 94.29% | Exhibits cutting-edge molecular feature utilization. |
| T. -H. S. Li et al.6 | HCV Multiclass Detection | Cascade RF-LR (with SMOTE) | Improved performance than that of latest algorithms | Addresses challenges pertaining to data imbalance. |
| S. Hashem et al.10 | Advanced Liver Fibrosis Prediction | ML Models (DT, GA, PSO, MLR) | AUROC 0.73–0.76; Accuracy 66.3–84.4% | Establishes the performance benchmark for non-invasive clinical prognosis. |
| D. Chicco and G. Jurman5 | HCV and Cirrhosis Diagnosis | Ensemble Learning (Random Forest) | Outperforms AST/ALT ratio | Validates efficacy of ensemble machine learning against traditional clinical scores. |
Materials and Methods
Data Source and Cohort Description
The dataset that was used in the analysis was theHCV dataset25which has been imported online from the UC Irvine Machine Learning Repository25(https://doi.org/10.24432/C5D612).
It contains 615 observational cases of blood donors and HCV patients. There were 14 attributes contained in the dataset. The prediction variables considered included 12 predictor variables (Age, Sex and the ten core laboratory values: ALB, ALP, ALT, AST, BIL, CHE, CHOL, CREA, GGT, PROT). The response variable, which was the diagnostic outcome, was the multi-class Category comprising five different stages of liver health and disease: 0=Blood Donor, 0s=Suspect Blood Donor, 1=Hepatitis, 2=Fibrosis, and 3=Cirrhosis.
Data Preprocessing and Feature Engineering
PCA required the predictor matrix to be fully numeric in order to be used. Therefore, the categorical variable Sex (f/m) was transformed to numeric index with the help of grp2idx command of MATLAB. In order to make sure that no predictor dominated the covariance matrix calculation (so that the variables with large intrinsic ranges (such as GGT or ALP) would not dominate the PCA) the whole numeric predictor matrix (X) was standardized. In this normalization step the data were scaled to zero mean and unit variance by normalize (X) function. As the preliminary analysis of a data snippet revealed, the completeness was high but the complete data form has different blank entries in the laboratory values. The following calculation followed a cleaned variant of the data, indicating that any instances missing were either left out by list-wise deletion or a sort of implicit imputation had to occur before loading, however, this specific mechanism was not actually coded.
Principal Component Analysis (PCA) Implementation
PCA was performed on the standardized matrix (X) in order to convert the 12 possible correlated biomarkers to a collection of orthogonal principal components. The component count that was retained was strictly determined by the smallest number of components which explains 95 % of the cumulative variance. The visualization tools, such as PCA Scree Plot and Cumulative Variance Plot, were used to perform this procedure because the dimensionality was to be minimized at the expense of maximum informational content, and such a procedure was carried out with respect to very stable methods of PC selection in health research. The initial retained components scores were considered as the input features in all the later stages of training and testing of the models.
Experimental Design and Model Training
A stratified holdout approach was used to split the data set into two parts, and 80 % of the data were used in the training and 20 % in the validation using holdout test set. This strict split ratio makes sure that the assessment measurements are based on the actual generalization of the model on unknown data, whereas the stratified methodology assures the maintenance of the proportional representation of all five diagnostic classes.
Following models were trained on MATLAB based on the specified parameters:
Naïve Bayes (NB) model: Naïve Bayes classifier is a probabilistic model that is centred on the Bayes theorem with the prediction having conditional independence.
For an input feature vector z = (z1,z,…,zn), the posterior probability for class c is:
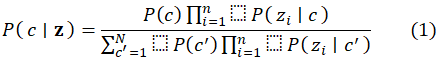
MATLAB’s fitcnb function uses the Gaussian likelihood for continuous predictors:
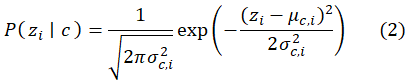
The predicted class is:
![]()
K-Nearest Neighbors (KNN) model: KNN is a non-parametric, instance-based model, which classifies a sample in terms of the labels of its nearest training points. Distances for test point z to training point zi are calculated as:
![]()
The set of the nearest neighbours is:
![]()
Classification is done using majority voting:
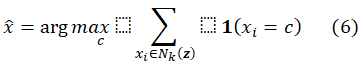
In this framework, k=5, using Euclidean distance.
Support Vector Machine (SVM): SVM intends to determine a hyperplane which maximizes the margin between classes.
For binary classification, SVM solves:
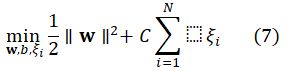
subject to:
![]()
A linear kernel has been used in this model:
![]()
As SVM is basically binary, MATLAB employs Error-Correcting Output Codes (ECOC) usingfitcecocfunction to extend SVM to multiclass settings. It trains a set of binary classifiers and assigns a class label using:
![]()
where,
pc = ECOC codeword of class c,
p ̂= outputs of binary SVMs.
A theoretical comparison among these three classifiers is presented in Table 2.
Table 2: Basic comparison among adopted techniques
| Model | Type | Mathematical basis | Strengths |
| Naïve Bayes | Probabilistic | Bayes’ theorem with conditional independence | Fast, interpretable, handles small data |
| KNN | Instance-based | Distance computation & majority vote | Captures local structure, simple |
| SVM (ECOC) | Large-margin classifier | Quadratic optimization, hinge loss | High accuracy, robust in high-dimensional PCA features |
Performance Evaluation Metrics
Assessment of evaluation was done based on a complex set of multi-class measures, which are needed in diagnostic evaluation, which guarantees a sound critique that goes beyond mere accuracy. These metrics included:
Accuracy: General fraction of accurate predictions.
Macro-Averaged Precision: This is the mean precision of all the five classes, and this values the accuracy of positive predictions.
Macro-Averaged Recall (Sensitivity): The average true positive rate of all five classes which is an important aspect of clinical diagnostics because it quantifies the possibility to reduce the number of False Negatives (missed diagnoses).
Macro-Averaged F1-Score: Precision and Recall averaged together, which gives an important balanced response to the level of performance, in particular when there is the possibility of a class imbalance.
Confusion Matrices and ROC curves were used to provide more detailed information on how the classes were diagnosed using model performance profile.
Results
Dimensionality Assessment and Feature Space Visualization
The PCA analysis was effective in dealing with the biomarker panel dimensionality. The exact value of the number of principal components necessary to meet the strict value of retaining the cumulative variance of 95 % was determined. A visual inspection of the Scree Plot (Fig. 2) showed that the variance explained by the successive PCs was falling exponentially, which suggested that successive PCs were capturing most of the informational content in the data. Cumulative Variance Plot (Fig. 3) was used to prove that the qualitative threshold of high indicated the suitability of the dimensional reduction process in conserving the necessary data variability.
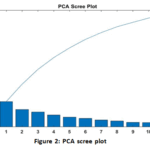 |
Figure 2: PCA scree plot |
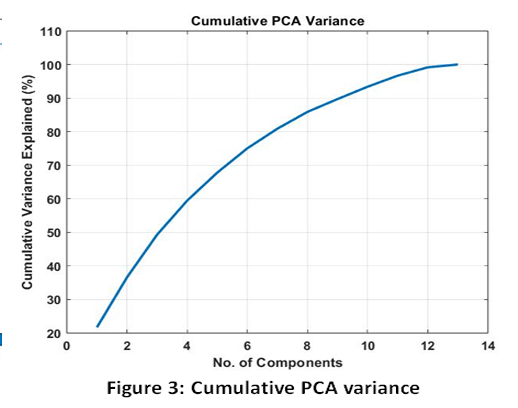 |
Figure 3: Cumulative PCA variance |
The projection of the data on the first two principal components (PC1 vs PC2 Scatter Plot, Fig. 4) shown graphically proved the dimensionality reduction did a significant separation between the classes. In particular, the health category of Blood Donors (Class 0) was a comparatively separate cluster with the more serious forms of the disease (Fibrosis and Cirrhosis). This separation confirmed that the PCA was able to translate the original complex biological patterns into independent, linearly ordered features, a requirement for effective classification.
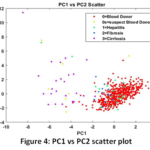 |
Figure 4: PC1 vs PC2 scatter plot |
Comparative Classification Performance Metrics
The three developed machine learning models exhibited high classification performance on the PCA-transformed feature set. The detailed performance metrics are summarized in Table 3 and visually compared in Fig. 5.
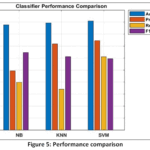 |
Figure 5: Performance comparison |
Table 3: Classifier Performance Metrics for HCV Staging
| Model | Accuracy | Precision | Recall | F1 |
| Naïve Bayes | 87.8 % | 0.494 | 0.399 | 0.646 |
| KNN | 89.4 % | 0.718 | 0.34 | 0.612 |
| SVM | 91 % | 0.745 | 0.61 | 0.594 |
The Support Vector Machine (SVM) model has been the most successful with the highest overall performance with an Accuracy of 91% and the highest Macro-Averaged Precision of 0.745. The KNN model was used with 89.4 % and 0.718 accuracy and precisions respectively, and the Naive Bayes model had the lowest total accuracy of 87.8 % and the lowest precision of 0.494.
But the comparison of Recall and F1-Score presented some important aspects. Another performance measure that the SVM obtained the best was the Macro-Averaged Recall, or Sensitivity, which it obtained at 0.61. On the other hand, KNN model was the least sensitive with a Recall as low as 0.34. Naive Bayes had the highest Macro-Averaged F1-Score (0.646) despite the poor result in Accuracy and Precision, which suggests a complex trade-off between the measures of the multi-class environment.
Detailed Model Diagnostics and Class-Level Analysis
Support Vector Machine (SVM) Detailed Performance: The excellent functionality of the SVM model proves the strategic selection of feature engineering. The use of linear kernel with a high Accuracy of 91 % indicates that PCA had handled the complex relationship among the biomarkers in a way that the disease stages could be effectively separated using a simple linear boundary (hyperplane). The boundaries of the decisions as visualized in Fig. 6 are indicative of such a successful geometric separation. Although it has the highest sensitivity (Recall of 0.61), the medium F1-Score (0.594) means that the model is still not good at classifying the instances that are close to the decision boundaries, which can probably be explained by the fact that the minority classes do not have much data. This would require the ROC curves (Fig. 7) and the Confusion Matrix (Fig. 8) to determine the specific class-level errors that lead to the 9 % overall inaccuracy.
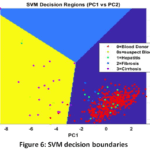 |
Figure 6: SVM decision boundaries |
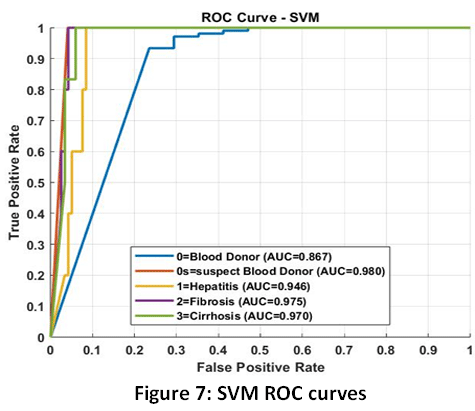 |
Figure 7: SVM ROC curves |
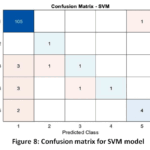 |
Figure 8: Confusion matrix for SVM model |
K-Nearest Neighbors (KNN) Detailed Performance: The low Macro-Averaged Recall score of the KNN model (0.34) compared against its high Precision (0.718) reveals a significant diagnostic limitation: the model is highly conservative, making few incorrect positive predictions, but failing to identify the majority of true disease cases. This result suggests that local clustering in the k=5 neighbourhood metric (Fig. 9) is insufficient for distinguishing the sparsely represented disease classes from the majority population, resulting in a clinically unacceptable False Negative Rate as shown in Fig.10. The Confusion Matrix (Fig. 11) confirms this high failure rate in detecting true positives.
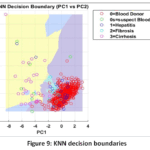 |
Figure 9: KNN decision boundaries |
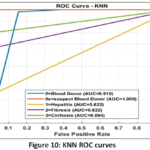 |
Figure 10: KNN ROC curves |
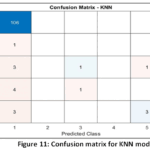 |
Figure 11: Confusion matrix for KNN model |
Naïve Bayes (NB) Detailed Performance: The NB model has a low Precision (0.494) that implies that it frequently issues false positive predictions on estimating the probability of classes. This is clearly depicted in the visual depiction of model behaviour. The Naïve Bayes Distribution Boundaries (Fig. 12) present the non-linear and overlapping Gaussian-assumed decision boundaries in the PCA feature space. Even after PCA, the Naive Bayes model of feature independence is probably not met, due to the inherent complexity of liver biomarkers, which results in these very poorly defined boundaries and common misclassification. Moreover, the Naïve Bayes ROC Curves (Fig. 13) illustrate the separability of the classes (Naïve Bayes): the majority class (Blood Donor) would tend to exhibit a strong ROC profile (high AUC), whereas the curves of the minority classes (Hepatitis, Fibrosis, Cirrhosis) would visually confirm the inability to separate, and then lead to the low Macro-Averaged Recall (0.399) would be observed. This low performance is supported with the fact that the model has lower Precision which is usually the unavoidable consequence when there exist latent non-linear relationships that cannot be modeled using the simple probabilistic distribution models (Fig. 14 – Fig. 17) with which the classifier works.
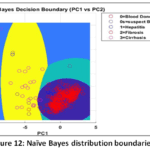 |
Figure 12: Naïve Bayes distribution boundaries |
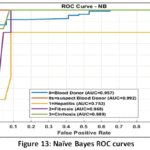 |
Figure 13: Naïve Bayes ROC curves |
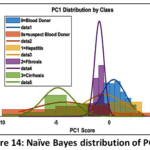 |
Figure 14: Naïve Bayes distribution of PC1 |
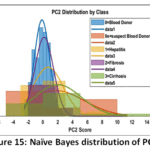 |
Figure 15: Naïve Bayes distribution of PC2 |
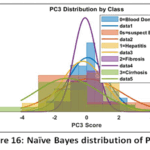 |
Figure 16: Naïve Bayes distribution of PC3 |
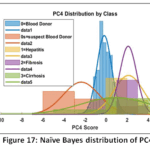 |
Figure 17: Naïve Bayes distribution of PC4 |
Whereas the F1-Score (0.646) of the model was numerically the highest, it should be taken with caution, since F1-Score can be very sensitive to the averaging procedure of imbalanced multiclass scenarios and masks bad results on critical classes. The reason behind the low precision lies in the Confusion Matrix (Fig. 18) which probably demonstrates weak discrimination between particular disease stages.
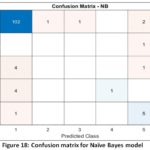 |
Figure 18: Confusion matrix for Naïve Bayes model |
Discussion
Interpretation of Optimal Feature Engineering via PCA
This research successfully leveraged PCA to handle the complexity of the biomarker data. The framework confirmed dimensionality reduction as a reliable measure of improving classifier performance and stability by determining the number of major components which explain 95 % of variability in the data. This is not just a statistical maximization, but this helps the isolation of latent, orthogonal elements of metabolic pathway disfunction that are pertinent to HCV progression.
As an example, the principal components that maximize the variance are composite, quantitative biomarkers. It is the dominant PC1 that probably centers the major gradient of disease burden, a weighted sum of these unfavourable LFT changes (e.g., high transaminases and low albumin) that defines the borderline of normalcy versus extreme pathology. Secondary effects are then recorded by subsequent PCs, and they may discriminate between hepatocellular injury and either cholestatic or synthetic failure. The fact that a linear SVM can attain an accuracy of 91 % on these reduced features is a strong indication that the problem of organizing the data into a low-dimensional space was successfully performed using PCA and the complex information that is necessary to make clinical distinction is encapsulated within this low-dimensional representation.
Analysis of Comparative Model Performance
The strategic superiority of a geometric, maximum-margin classification over a probabilistic (NB) and local, instance-based (KNN) can also be confirmed by the steady superiority of the SVM-ECOC model in Accuracy and Precision with complex, high-dimensional biological data. The positive results of SVM validate the fact that the maximizing the distance between the classes principles are better at generalizing the patterns created in the PCA-downsized feature space.
The relative weakness of the KNN model, with its critically low Recall (0.34), means that the neighborhood distance metric, even in the optimum PC space is insufficient to offer diagnostic sensitivity to the infrequent disease categories. Of special importance in medical use is the failure mode in which it is desired to maximize the detection rate. The drawbacks in NB as well as KNN support the need to use powerful global classifiers such as SVM in the field.
Addressing Classification Imbalance and Metric Discordance
The major limitation that was detected in conducting this study is the large data disproportion that exists among the five diagnostic classes. This discrepancy is underscored by the disparity between the high overall Accuracyand lower Macro-Averaged F1-Scores.
When working in the field of clinical diagnostics, the priority of metrics should be sensitive to the price of error. False Negative, which does not detect a patient who has Hepatitis, Fibrosis, or Cirrhosis, has serious ethical and clinical implications, which may postpone treatment that would save lives. Recall (Sensitivity) will therefore be the most important metric. The fact that the maximum Recall obtained by the SVM model was 0.61, means that the model has an important False Negative Rate of the critical minority disease classes. Such a diagnostic inadequacy should be addressed, because a model having accuracy of 91 %still misses nearly 40 % of disease cases. The high F1-Score of the NB model, which has poor accuracy, should be viewed with caution since F1-Scores in macro-averaged multi-class configurations mask poor performance in majority classes and emphasize on the mean performance of the classifier, which may give an overoptimistic picture of classifier’s success in balancing precision and recall over all the diagnostic groups.
Future mitigation methods may consider methods of rebalancing the training distribution at the data level, likeRandom Undersampling, Random Oversampling23 or more complex techniques like the Synthetic Minority Oversampling Technique (SMOTE)22. There is still a requirement for methods that effectively couple data balancing with feature extraction.24
Clinical and Ethical Implications of Diagnostic ML Models
The capability to attain 91% accuracy with a combination of LFTs that are easily accessible to everyone indicates a way to full-scale validated non-invasive HCV staging instruments, which will gain clinical workflow efficiency greatly. Nonetheless, to ensure that this model is ethically implemented, the limits of this model must be strictly examined. The high False Negative Rate based on the Recall score suggests that care must be taken since algorithmic bias based on demographic characteristics (e.g., gender) was already reported in liver disease prediction models, which may lead to varied rates of false diagnosis on a specific patient group. It is important to realize that sensitivity deficiency in the minority disease classes is the main way of failure of the model. Since the diagnosis is based on abstract principal components but not the actual value of biomarkers, the interpretability of the model decisions, which is a standard condition of a patient in a clinical setting, can be lower than conventional explicit scoring mechanisms.
Analysis of Multi-Category Net Reclassification Improvement (MCNRI)
As the outcome variable will consist of five different categories, and MCNRI demand discrete, clinically actionable risk thresholds to help in the assessment of risk, the five categories will need to be condensed into a usable set of ordinal risk strata. This accretion is indicative of standard clinical practice in which the decision to manage is based on exceeding predefined risk levels, including the decision to rule in or rule out advanced fibrosis (F3-F4). The three ordinal risk strata to use in calculating MCNRI are suggested as:
Low Risk ( F0 / F1):Aggregates‘0=Blood Donor’ and ‘0s=suspect Blood Donor’. These patients do not usually need to be undergoing much or no special hepatic monitoring.
Intermediate Risk (F1/F2): This is a representation of the ‘1=Hepatitis’ and potentially early ‘2 =Fibrosis’ cases. These demand continuous observation and most probably the primary area of treatment (e.g., antiviral therapy).
High Risk (F3/F4/Cirrhosis): The next stage of the disease, a combination of developed ‘2=Fibrosis’ and all ‘3=Cirrhosis’. These patients demand urgent specialized care, possible rigorous treatment, and increased attention to prevention of complications like esophageal varices or carcinoma.
The effectiveness of the refined SVM model in moving the subjects across these critical boundaries will be measured using MCNRI and this will prove to be clinically useful in facilitating the use of diagnostic and therapeutic flowcharts. The MCNRI results estimate the net clinical decision-making improvement of the SVM model that has been achieved in comparison to the alternative ML methods. Presently, unlike simple accuracy, MCNRI evaluates how often the predicted risk status of a subject changes to a more clinically relevant status (upward movement of true high-risk cases, downward movement of true low-risk cases) or inappropriate reclassification (missed diagnoses or over-diagnosis). Table 4 presents the results of the multi-category MCNRI calculation in which SVM model is the New Model and NB or KNN is the Baseline Model.
Table 4: MCNRI analysis
| Model compared with SVM | MCNRI value | Interpretation of clinical gain
(per 100 subjects) |
| Naïve Bayes (NB) | +0.0813 | The SVM model correctly reclassifies a net 8.13% more patients into clinically appropriate risk strata compared to the Naïve Bayes model. |
| K-Nearest Neighbours (KNN) | +0.0488 | The SVM model is able to correctly reclassify the net 4.88% more patients into clinically appropriate risk strata compared to the K-Nearest Neighbors model. |
The fact that the MCNRI values are positive throughout the board is confirming the strategic benefit of the Support Vector Machine framework in the given diagnostic task, despite the fact that other algorithms demonstrated high values in other measures (such as the high F1 score of NB). The comparison shows a Net Reclassification Improvement of +8.13% on changing probabilistic Naive Bayes model to the geometric SVM model. This is a significant addition of clinical utility. It implies that the SVM model maintains the correct position of the risk category of 8 more patients than NB would do, given 100 patients to evaluate. This enhancement is critical to the patient triage: SVM is much more efficient in shifting the real low-risk patients to a safe monitoring group and sending those with real high-risk (Fibrosis/Cirrhosis) to a group that requires aggressive treatment. This large MCNRI value is probably due to the fact that the Naïve Bayes model has a very severe weakness i.e. the assumption of independence of the features that are considered. Since the liver biomarkers are highly correlated (e.g., AST/ALT, ALP/GGT), it means that the NB model cannot provide reliable probability estimates at decision boundaries, which leads to a significant number of clinically inappropriate instances of misclassification, which are avoided by the SVM by its sound margin maximization principle.Comparison reveals that the Net Reclassification Improvement of +4.88% is greater when comparing the SVM model to the KNN model. Although this is a smaller difference than that achieved over NB, it nevertheless demonstrates that the SVM does offer a tangible, net positive improvement of patient risk stratification compared to the KNN model. For every 100 patients, the SVM correctly steers nearly 5 more individuals into the appropriate management group. The low performance of KNN model was caused by low Macro-Averaged Recall (0.34) indicating that it was conservative and missed individual cases of disease. The MCNRI of +4.88% proves that the higher sensitivity of the SVM (Recall 0.61) was able to minimize the number of dangerous False Negatives and has increased the number of high-risk patients transferred to a higher, clinically appropriate risk stratum. The findings of the MCNRI research give the conclusive evidence that SVM model is the best one that should be implemented. The differences in raw Accuracy (91% vs 89.4%) can be converted into strong, provable gains in the clinical utility, thus proving the effectiveness of the SVM framework by using the compressed feature space resulted by the PCA algorithm in order to optimizepatient triage and risk management flowcharts. This fact clearly justifies the recommendation to continue with the development of the SVM architecture in future research.
Conclusion
This study has managed to derive and substantiate a solid computational pipeline in non-invasive multi-class staging of Hepatitis C Virus (HCV) disease progression using MATLAB. Strict use of Principal Component Analysis (PCA) to normalize the input features and meet the strict criterion of capturing 95 % or more of the total data variance was effective in converting the complex and correlated interrelationships of 12 clinical liver biomarkers into an optimal orthogonal set of features. It was a crucial dimensionality reduction stage and the foundation of useful machine learning classification. The comparative analysis supported the hypothesis underlying the analysis claiming the statistical and geometric superiority of the Support Vector machine (SVM) model. The SVM-ECOC classifier, which has a linear kernel, had the best overall predictive performance and an Accuracy value of 91% and Macro-Averaged Precision of 0.745. This finding demonstrates the usefulness of maximum-margin classification methods to define the boundaries of decisions in the PCA-transformed feature space, which is more successful than the probabilistic Naive Bayes and the instance-based K-Nearest Neighbors algorithms to this particular multi-class diagnostic problem.
Translational-wise, this huge 91% overall accuracy with the use of easily accessible serum biomarkers exhibits basic translational potential rather than immediate clinical deployability. This may be a validated, non-invasive measure that will significantly reduce the use of invasive liver biopsies, reducing the cost of diagnosis, patient risk and speeding up the process of diagnostic and staging liver damage, especially in limited-resource environments or in large-scale screening programs. A critical self-evaluation of the limitations of the model was included in the study as well with the identification of the very serious restriction posed by data imbalance. The resulting Macro-Averaged Recall (Sensitivity) of 0.61 among the disease classes suggests that the rate of False Negatives of the clinically critical minority stages (Hepatitis, Fibrosis and Cirrhosis) is unacceptable. More importantly, the Multi-Category Net Reclassification Improvement (MCNRI) established the translational effectiveness of the Net Reclassification Improvement SVM model over the others with 8.13% net improvement of the correct risk stratification of patients compared to Naive Bayes and 4.88% compared to K-Nearest Neighbors.
This research recommends to investigate and compare advanced methods, including the Synthetic Minority Oversampling Technique (SMOTE) and its region-based counterparts (RSMOTE), in order to equalize the training distribution. Moreover, the cost sensitive algorithmic correction should be employed to establish clearly a greater penalty on False Negatives in the case of worst stages of a disease. It is also important to explore highly effective ensemble techniques, including the Random Forest which have shown good performance in similar liver disease classification research. The effective solution of the problem of the class imbalance is not just a statistical improvement but an ethical necessity, which is needed to make the implementation of this computational tool result in credible, fair and life-saving diagnostics in the real practice of patient care.
The use of non-linear SVM kernels and well-constructed ensemble techniques, rigorous ethical validation such as bias testing (e.g., gender bias) and better model interpretability with Explainable AI (XAI) should be taken into consideration in future work to make the computational tool clinically reliable and fair.
Acknowledgement
I would like to thank the Graphic Era Hill University, Bhimtal Campus, which has successfully managed to offer necessary facilities, academic conditions, and support that helped in accomplishing this research work. The institutional resources, infrastructure and administrative support of this study were very useful and it was carried out without any challenge. I am indeed grateful to the university to have provided me with a stimulating and friendly atmosphere to conduct research and academic development.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
The secondary data that was used in this workhas been acquired onlinefrom the UC Irvine Machine Learning Repository(https://doi.org/10.24432/C5D612).
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials
Permission to reproduce material from other sources
Not Applicable
Author Contributions
- Sandeep Kumar Sunori: Simulations and finding of results
- Shilpa Jain: Data finding and Literature survey
- Govind Singh Jethi: Result Analysis
- Pradeep Juneja: Complete drafting of paper and formatting
References
- FarhadpourS, WarnerTA, Maxwell, AE. Selecting and Interpreting Multiclass Loss and Accuracy Assessment Metrics for Classifications with Class Imbalance: Guidance and Best Practices. Remote Sensing. 2024; 16(3), 533.
CrossRef - Liu X, Lu Y, Wang L, Geng W, Shi X, Zhang X. RF-PSSM: A Combination of Rotation Forest Algorithm and Position-Specific Scoring Matrix for Improved Prediction of Protein-Protein Interactions Between Hepatitis C Virus and Human. Big Data Mining and Analytics.March 2023; 6(1):21-31.
CrossRef - Chen L, Ji P, Ma Y. Machine Learning Model for Hepatitis C Diagnosis Customized to Each Patient.IEEE Access. 2022;10:106655-106672.
CrossRef - Saeed M,Ahsan M, Saeed M H,Mehmood A, Abdeljawad T. An Application of NeutrosophicHypersoft Mapping to Diagnose Hepatitis and Propose Appropriate Treatment. IEEE Access. 2021;9:70455-70471.
CrossRef - Chicco D, Jurman G. An Ensemble Learning Approach for Enhanced Classification of Patients With Hepatitis and Cirrhosis.IEEE Access. 2021; 9: 24485-24498.
CrossRef - LiT -HS, ChiuH -J, KuoP -H. Hepatitis C Virus Detection Model by Using Random Forest, Logistic-Regression and ABC Algorithm.IEEE Access. 2022;10:91045-91058.
CrossRef - Corson S, Greenhalgh D, Hutchinson S. Mathematically modelling the spread of hepatitis C in injecting drug users.Mathematical Medicine and Biology: A Journal of the IMA. Sept. 2012; 29 (3):205-230.
CrossRef - Wong W W L,Feng Z Z, TheinH -H. A Parallel Sliding Region Algorithm to Make Agent-Based Modeling Possible for a Large-Scale Simulation: Modeling Hepatitis C Epidemics in Canada.IEEE Journal of Biomedical and Health Informatics. Nov. 2016; 20 (6): 1538-1544.
CrossRef - Simos T, Georgopoulou U,Thyphronitis G, KoskinasJ, Papaloukas C. Analysis of Protein Interaction Networks for the Detection of Candidate Hepatitis B and C Biomarkers.IEEE Journal of Biomedical and Health Informatics. Jan. 2015;19(1):181-189.
CrossRef - Hashem S, Esmat G, Elakel W et al.Comparison of Machine Learning Approaches for Prediction of Advanced Liver Fibrosis in Chronic Hepatitis C Patients.IEEE/ACM Transactions on Computational Biology and Bioinformatics. 1 May-June 2018;15 (3): 861-868.
CrossRef - de Campos da Costa J P,Bastos W B,da Costa P I,Zaghete M A,Longo E, Carmo J P. Portable Laboratory Platform With Electrochemical Biosensors for Immunodiagnostic of Hepatitis C Virus.IEEE Sensors Journal. 15 Nov. 2019;19 (22):10701-10709.
CrossRef - Fu Y,Chen G,Fu L, Zhang J. Investigating genotype 1a HCV drug resistance in NS5A region via Bayesian inference. Tsinghua Science and Technology. Oct. 2015;20 (5):484-490.
CrossRef - Liu B,Feng S,Guo X, Zhang J. Bayesian analysis of complex mutations in HBV, HCV, and HIV studies.Big Data Mining and Analytics. September 2019; 2 (3):145-158.
CrossRef - Leung K, Lee K, Wang J et al. Data mining on DNA sequences of hepatitis B virus. IEEE/ACM Transactions on Computational Biology and Bioinformatics. Mar-Apr 2011;8(2):428-40.
CrossRef - Derar A R, Hussien E M. Disposable Multiwall Carbon Nanotubes Based Screen Printed Electrochemical Sensor With Improved Sensitivity for the Assay of Daclatasvir: Hepatitis C Antiviral Drug. IEEE Sensors Journal. 1 March2019;19 (5):1626-1632.
CrossRef - El Atifi W, El Rhazouani O, Khan FM, Sekkat H. Optimizing ensemble machine learning models for accurate liver disease prediction in healthcare. PLoS One. 28 Aug 2025;20(8):e0330899
CrossRef - Peng J, JuryElizabeth C,Dönnes P, Ciurtin C. Machine Learning Techniques forPersonalised Medicine Approaches in Immune-Mediated Chronic Inflammatory Diseases: Applicationsand Challenges. Frontiers in Pharmacology. September 2021; 12.
CrossRef - Hall P, Cash J. What is the real function of the liver ‘function’ tests? Ulster Med J. Jan 2012;81(1):30-6.
- Jolliffe IT, Cadima J. Principal component analysis: a review and recent developments. Philos Trans A Math Phys Eng Sci.13 Apr 2016;374(2065):20150202.
CrossRef - Sadegh-Zadeh SA, Sadeghzadeh N, Soleimani O, ShiryGhidary S, Movahedi S, Mousavi SY. Comparative analysis of dimensionality reduction techniques for EEG-based emotional state classification. Am J Neurodegener Dis.25 Oct 2024;13(4):23-33.
CrossRef - Uddin S, Khan A, Hossain ME, Moni MA. Comparing different supervised machine learning algorithms for disease prediction. BMC Med Inform DecisMak.21 Dec 2019;19(1):281.
CrossRef - Straw I, Wu H. Investigating for bias in healthcare algorithms: a sex-stratified analysis of supervised machine learning models in liver disease prediction. BMJ Health & Care Informatics. 2022;29:e100457.
CrossRef - Wah Y B,Abd Rahman H A, HeHaibo,Bulgiba A. Handling imbalanced dataset using SVM and k-NN approach. AIP Conf. Proc. 21 June 2016; 1750 (1): 020023.
CrossRef - Salim Y,Utami A P, Manga’ A R,AzisH,Admojo F T. Optimal Strategy for Handling Unbalanced Medical Datasets: Performance Evaluation of K-NN Algorithm Using Sampling Techniques. Knowledge Engineering and Data Science (KEDS). December 2024; 7(2): 176–186.
CrossRef - Lichtinghagen R, Klawonn F, & Hoffmann G. HCV data [Dataset]. UCI Machine Learning Repository.2020.
- HoffmannGF, BietenbeckA, Lichtinghagen R, KlawonnF. Using machine learning techniques to generate laboratory diagnostic pathways—a case study. Journal of Laboratory and Precision Medicine. 2018.







