Manuscript accepted on :02-03-2026
Published online on: 08-05-2026
Plagiarism Check: Yes
Reviewed by: Dr. Sandeep sharma
Second Review by: Dr. Hisham Abdel Aziz Orban
Final Approval by: Dr. Achyut Shankar
Gaurav Singh , Narander Kumar*
, Narander Kumar* and Shishir Kumar
and Shishir Kumar
Department of Computer Science, Babasaheb Bhimrao Ambedkar University (A Central University), Vidya Vihar, Raebareli Road, Lucknow, Uttar Pradesh, India.
Corresponding Author E-mail:nk_iet@yahoo.co.in
Abstract
A condition where the thyroid gland functions normally but the level of hormones is imbalanced, known as Sick Euthyroid Syndrome. SES is frequently observed in patients suffering from serious infections or chronic conditions and may mimic hypothyroidism. Machine Learning (ML) can assist doctors in diagnosing the disease because of the complex nature of SES. This paper proposed a Stacked RSL model that uses Random Forest, Support Vector Machine, and Logistic Regression ML classifiers with Sequential Forward and Sequential Backward Feature Selection mechanisms to select the best features and an Over-sampling technique to balance the dataset. The proposed model performs better with RandomOver sampling and feature selection methods, achieving 99.55% accuracy using 3,163 patient data taken from the UCI-ML repository. By addressing class imbalance and removing irrelevant attributes, the improved ML model enhances feature quality and model generalization.These technical improvements enable the algorithm to find the necessary diagnostic patterns more effectively than existing approaches, which enhances the overall predictive performance.
Keywords
Data balancing; Euthyroid; Feature selection; Machine learning; Stacking classifier
| Copy the following to cite this article: Singh G, Kumar N, Kumar S. Early Detection of Sick Euthyroid Syndrome using Stacked RSL Machine Learning Model. Biomed Pharmacol J 2026;19(2). |
| Copy the following to cite this URL: Singh G, Kumar N, Kumar S. Early Detection of Sick Euthyroid Syndrome using Stacked RSL Machine Learning Model. Biomed Pharmacol J 2026;19(2). Available from: https://bit.ly/4ddeTcS |
Introduction
The thyroid is a gland that present in neck, produces hormones in our body which plays a very important role in regulating metabolism, growth, and overall development.1Common states of thyroid including Hyperthyroidism, Hypothyroidism and Euthyroid state. When thyroid gland produces excessive amount of hormones, it is called Hyperthyroidism and when thyroid gland produces insufficient amount of hormones, it is called Hypothyroidism. Apart from hyperthyroidism and hypothyroidism, when thyroid gland functions normally in context of hormone level, it is called Euthyroid state. This ensures the proper functioning of various physiological processes, including metabolism, body temperature regulation, energy production, and cardiovascular health.
In certain cases, where no thyroid disease presented, but the thyroid hormone level still be detected abnormal due to the other sever illnesses. This situation of the patient is known as Euthyroid_Sick_Syndrome (ESS), also known as Sick-Euthyroid_Syndrome(SES)or Non-Thyroidal_Illness_Syndrome. SES is frequently observed in patients suffering from severe infections, chronic conditions like heart, kidney, or liver disease, and in those undergoing major surgeries or admitted to intensive care units. In such situations, the illness itself affects hormone levels, though the thyroid gland remains structurally and functionally intact.23Diagnosis of SES complicatesbecause its symptoms might be similarto hypothyroidism or overlap with other illnesses.This variability creates diagnostic challenges, as differentiating SES from primary thyroid dysfunction requires careful clinical judgment and exclusion of hypothyroidism.4 Also interpreting test results in acutely ill patients remains complex without advanced diagnostic tools.5
To make diagnosis of thyroid disease more accurate, Machine Learning (ML) techniques can be very useful.6 These methods help computers to learn from medical data and spot important patterns that might be missed otherwise. One of the important step in this process is feature selection, which means picking out the most important piece of informationlike specific test results or symptomsthat are closely linked to different thyroid conditions. When ML is combined with feature selection, it becomes a powerful tool that can help doctors to detect thyroid problems at earlier stage. This early detection can lead to faster treatment and better health outcomes for patients. Feature Selectionis particularly important as it helps reduce data dimensionality, prevent overfitting, and improve model interpretability, leading to more efficient and reliable predictive models.The application of ML algorithms, with a feature selection mechanism, provides a promising method for the early detection of ESS. By implementing advanced data analysis methods, these techniques can identify patterns and relationships within patient data that may not be noticed through traditional diagnostic approaches.7 8 9
The prediction of thyroid disorders in diabetes patients is presented by using six ML algorithms, which are RF, DT, Logistic Regression (LR), K-Nearest Neighbours (KNN), Naïve Bayes (NB), and SVM. RandomUnderSampler and manual balancing are used to balance the dataset, which consists of 44,534 diabetic patients dataset. The highest accuracy of 95% was achieved with RF and the manual balancing technique, and achieved 84% with RF and the RandomUnderSampler technique.10
To identify thyroid sickness,the fine-tuned LGBM (Light Gradient Booster Machine) is used11, while for data balancing, the nominal continuous SMOTE technique was used. The approach was outperformed by the SMOTE-NC-LGBM approach, achieving high-accuracy performance scores.
SMOTE oversampling techniques is usedto balance the hypothyroid dataset while SVM, LR, RF, KNN, and XgBoost algorithms are implemented to train the model. Among these techniques RF performs better.12
SMOTE oversampling technique is used to balance the dataset. SVM, DT, RF,Adaptive Boosting, and KNN were implemented to classify the chronic kidney disease using Kaggle dataset. Among these techniques SVM achieved better accuracy of 99.3% with 6 features.13
A CNN based model is proposed14to classify thyroid nodule by using boundary detection techniques. Hyperparameters such as learning rate and dropout factor were fine-tuned. For Thyroid Ultrasound Image Database (TDID) dataset model achieved 93.75% accuracy and for collected datasets from scanning centre it achieved 96.89% accuracy. Proposed model shows [2, 5]% improvement in comparison of some deep learning techniques.
The Decision Tree classifier is used to detect heart disease,and SMOTE oversampling technique is used to balance the dataset.15 With the use of SMOTE, they achieved 11% more accuracy in comparison of without using SMOTE technique.
ML classifiers, including RF, Gradient Boosting, DT, XGBoost , LR, NB, and SVM, are used to predict the diabetes disease. With feature selection techniques, LGBM performed better with 98.99% accuracy for DMS (Diabetes Mellitus Dataset) and 92.5% accuracy for Pima Indians Diabetes Dataset.16
Variousmethods were employedto overcome the issues associated with imbalanced and limited training datasets. The OhioT1DM public dataset, which was obtained from BID Medical Centre, was used to predict type 1 diabetes. MLP outperformed with 86.08% accuracy,and LSTM achieved 87.26%.17
The ML algorithms were used to predict the trend of levothyroxine (LT4) treatment in hypothyroid patients. The dataset containing patients’ medical histories was taken from the AOU-Federico II hospital. Among various mechanisms, the highest accuracy of 84% obtained by the Extra-Tree Classifier. The findings aim to assist endocrinologists in optimizing treatment and improving patient care.18
The research19 focuses on classifying thyroid disorders using four models: NB, DT, Multilayer Perceptron (MLP), and Radial Basis Function Network (RBFN). They used datasets from a Romanian source and the UCI ML repository and obtained the highest accuracy of 97.35% with the Decision Tree classifier.
The study develops ML models to classify seven thyroid disease categories using a comprehensive multi-feature thyroid dataset. Ensemble-based algorithms (RF, GB, DT) were trained with cross-validation and hyperparameter tuning. Gradient Boosting achieved the best performance with 0.97 F2 score, 94.7% accuracy, and strong precision–recall values.20
Table 1 represents the summary of related work of newly published research papers, presented in tabular form.
Table 1: Summary of Related Work
| Reference | Method | Abridgment | Strength | Weakness |
| [10] | RF, DT, LR, KNN, Naïve Bayes, and SVM | RF achieved the best performance with 95% accuracy (manual) and 84% (under-sampled). | Robust evaluation using Stratified K-Fold CV. | Performance drop with RandomUnderSampler, may lack a comparison with other sampling methods. |
| [11] [12] [15] | LGBM SVM, LR, RF, KNN, XgBoost, and DT | ML techniques with SMOTE Oversampling are used, in which LGBM, RF, and DT perform better with different datasets. | SMOTE over sampling, enabling the model to learn minority-class patterns more accurately and resulting in enhanced overall performance. | In addition to oversampling, feature selection may improve theperformance of the model that focuses on the prediction goal. |
| [13] [16] | SVM, DT, RF,Adaptive Boosting, KNN,
Gradient Boosting, DT, XGBoost, LR, and NB. |
SVM achieved an accuracy of 99.3% using Kaggle dataset, and LGBM achieved 98.99% accuracy for the DMS dataset using feature selection techniques. | Provides a comprehensive evaluation of resampling methods for imbalanced medical data. | lacks external validation |
| [17] | RF, MLP, and LR for classification. LSTM, MA, and LR for predictive analysis. | Using PIMA dataset, MLP achieved 86.08% accuracy, and LSTM achieved 87.26%. Also proposes IoT-based BG monitoring concept. | Uses multiple ML predictive models; integrates IoT concept; covers classification & forecasting. | accuracy could improve with optimized or ensemble techniques. |
| [19] | NB, DT, MLP, RBF Network | ML classifiers and ANN models are compared, in which DT achieved highest accuracy of 97.35%. | Detailed experimental comparison as well as evaluating diverse attributes with two datasets. | Accuracy could increase further with more suitable ML models or advanced ensemble strategies to stabilize predictions. |
| [20] | RF, Gradient Boosting, DT | The study presents machine learning models for classifying seven categories of thyroid disease using a multi-feature dataset. Ensemble-based techniques including RF, Gradient Boosting, and Decision Trees were trained. The best performance was achieved by Gradient Boosting with 94.7% accuracy. | comprehensive cross-validation and detailed analysis using a confusion matrix. | Accuracy could be further improved through the careful selection of more suitable algorithms or by adopting advanced ensemble strategies. |
Although various approaches have been discussed, each comes with certain limitations. We found that there is a need of hybrid technique which combine the most suitable feature selection methods with the best data balancing strategies.As research in this field continues to advance, the integration of feature elimination techniques and ML algorithms has great potential for improving the early detection and management of euthyroid sick syndrome, leading to better outcomes and more efficient healthcare delivery. This paper explores the application of feature selection and ML techniques for the early detection of sick euthyroid disease, focusing on improving diagnostic accuracy, reducing healthcare costs, and enhancing patient outcomes through timely interventions.
Materials and Methods
This paper proposed a Stacked RSL (R: Random Forest, S: Support Vector Machine, L: Logistic Regression) Machine Learning model (as shown in Figure 1) that used three ML classifiers in such a way, that the predictions of two classifiers work as input for the third classifier.The system incorporates two feature selection mechanismsi)Sequential Forward Feature Selection (SFFS) technique and ii) Sequential Backward Feature Selection (SBFS) technique. Since the dataset is highly imbalanced, we used the over-sampling techniques to balance the dataset. Figure 5 represents the proposed Stacked RSLML model with feature selection and the oversampling technique. The very first step is preprocessing, where feature selection mechanisms are applied to determine the most pertinent features, reducing noise in dataset. After that, two oversampling techniques, RandomOverSampler and SMOTE, are applied to solve class imbalance issue and improve the model performance. Following this, two base models (or base classifier) SVM and RF are trained with the help of stacking technique. These models work independently and generating predictions using refined input. After obtaining the prediction of base models, a meta model or meta classifier LR is introduced. Instead of working directly with the input data, the LR (meta model) works on the predictions obtained from the base models to produce more accurate results.
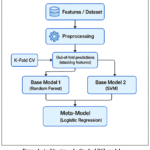 |
Figure 1: Architecture of a Stacked RSL model. |
Preprocessing
In this step, our main objective is to clean the dataset by handling any missing or unnecessary data. More precisely, we detect and exclude attributes (or columns) that have a significant proportion of null or missing values, as they might distort the outcomes and reduce the accuracy of the model. In addition, we remove features that are considered non-essential or irrelevant to the analysis, such as those that do not provide valuable information to the model for prediction.
Feature Selection
Selecting the most ideal features from the dataset is known as FeatureSelection. It helps to eliminate redundant, irrelevant, or noisy features to enhance the model performance.
Sequential-Forward Feature Selection (SFFS)
SFFS is a feature selection technique that begins with no features and progressively adds them one by one. At each step, it adds a feature that enhance the model performance most. Repeat this process until there is no improvement after adding the new feature. One of the main advantages of SFFS is that it reduces computational complexity by incrementally selecting relevant features from the dataset instead of working on all possible subsets.
Algorithm 1 SFFS
Require: Data set of all features
Ensure: Select best feature set
Step 1: Initialize the set of selected features:
S = ∅
Step 2: Repeat until the required number of features is selected:
For each feature f not in S:
Create a temporary subset S’ = S ∪ {f}.
Train the model using S’.
Evaluate the model performance.
Identify the feature f* that provides the best performance.
Update the selected set S = S ∪ {f*}.
Step 3: Return the final selected feature subset S.
Sequential Backward Feature Selection (SBFS)
SBFS is a type of SequentialFeature Selection technique. It takesall available features and removes the least important one by one at each step based on the model performance. This process continues until the removal of feature(s) impacts negatively on the performanceof the model. One of the importantadvantages of SBFS is its ability toidentify the minimum number of subset of features that provides the best performance, this ensure that only the most relevant variables are retained.
Algorithm 2 SBFS
Input: Complete feature set
Output: Reduced optimal feature subset
Step 1: Initialize the selected feature set:
S = {all features}
Step 2: Repeat while the number of features in S is greater than the desired count:
For each feature f in S:
Create a temporary subset by removing the feature: S’ = S − {f}.
Train the model using S’.
Evaluate the model’s performance.
Identify the feature whose removal results in the least performance drop or the greatest performance gain.
Update the selected set: S = S − {f*}.
Step 3: If further removal significantly degrades model performance:
Stop the elimination process early.
Step 4: Return the final selected feature subset S.
Dataset Balancing
RandomOver Sampler:
RandomOver Sampler is a simple oversampling technique. In this, instances of the minority class are randomly duplicated until a balanced class distribution is achieved.
Procedure
Identify the minorityclass (Rmin) and majority class (Rmaj) in the taken dataset.
Randomly select instances from Rmin with replacements and add them to the dataset.
Continue sampling until the number of instances in Rmin matches or approaches Rmaj.
Figure 2. shows the unbalanced and Figure 3.shows the balanced data by using RandomOver Sampler.
 |
Figure 2: Unbalanced Dataset |
 |
Figure 3: Balanced dataset with ROS technique |
SMOTE (Synthetic Minority Over-sampling Technique)
It is a data balancing technique that generates synthetic data points between existing data samples, instead of duplicating minority class samples.
Procedure
Select a Minority Class Instance: Randomly choose a data point X from the minority class.
Find KNN: Identify the k-nearest neighbours of X within the minority class using Euclidean distance.
Randomly Select One Neighbour: Randomly choose one of the k-nearest neighbours XNN.
Generate a Synthetic Sample: Create a new synthetic sample using the formula:
Xnew = X + λ × (XNN – X)
where, λ ϵ [0,1]
Repeat the Process: Continue generating synthetic samples until the desired class balance is achieved.
Figure 2. shows the unbalanced and Figure4.shows the balanced data by using SMOTE.
 |
Figure 4: Balanced dataset of sick-euthyroid and negative patients with the SMOTE technique |
Algorithm 3: Proposed Model
Input: Training data (Xrsltrain,yrsltrain), Test data Xrsltest; Base classifiers B1,B2,…,Bn
Output: Final predicted class labels
Step 1: Select the base classifiers B1,B2,…,Bn.
Step 2: For each base classifier Bi:
![]()
Step 3: Generate prediction outputs from all base classifiers on the training set:
![]() .
.
Step 4: Construct a new training dataset using these predictions as features.
Step 5: Train the meta-classifier using the new dataset.
Step 6: Use each trained base classifier to generate predictions on Xrsltest:
![]() .
.
Step 7: Provide these test predictions as input to the trained meta-classifier to obtain final outputs.
Step 8: Return the final predicted class labels.
 |
Figure 5: Proposed Stacked RSLML model with feature selection and the over sampling technique |
Results
With the use of RandomOver Sampler technique, Table2 and Figure 6 describe the comparison of the performanceof the proposed Stacked RSL (S-RSL) model with SVM, LR, and RF. Our proposed model outperformed with these classifiers. Furthermore, two feature selection techniques are used to select the minimum features with the best accuracy.
Table 3 showsthe S-RSL model’s performance using the SFFS andRandomOverSampler techniques. With 5-fold cross-validation (CV) and stratified 5-fold CV, the S-RSL model achieved 99.34% and 99.53% accuracy, respectively. With 10-fold CV and stratified 10-fold CV, it achieved99.55%accuracy.Table 4 shows the S-RSL model’s performance using SBFS and ROS. With 5-fold CV and stratified 5-fold CV, the S-RSL model achieved 99.42% and 99.46% accuracy, respectively.With 10-fold CV and stratified 10-fold CV, it achieved 99.49% and 99.55% accuracy, respectively.Table 5 shows theS-RSL model’s performance using the combined feature of SFFS and SBFS techniques along with the RandomOverSampler (ROS) technique. With 5-fold CV and stratified 5-fold CV, the model achieved99.37 % and 99.55% accuracy, respectively. With 10-fold CV and stratified 10-fold CV, it achieved99.55% and 99.55% accuracy, respectively. Figure9 show the performance comparison of theS-RSL model using ROS techique and Figure10 show the performance metrics across k-folds.
With the use of SMOTE and SFFS technique Table6 shows theS-RSL model’s performance with accuracy of 99.30%and with SBFS technique Table 7 showsthe S-RSL model’s performance with accuracy of 98.67% and when we take the union of features of SFFS and SBFS techniques, the S-RSL model shows 98.60% accuracy, which is mentioned in Table8.
Table 2: Performance of different classifiers, including Proposed model
| S. No. | Over-Sampling Method | Technique Used | Cross-Validation Technique | Accuracy (%) |
| 1. | Random Over Sampler | SVM | 5-fold | 63.53 |
| 2. | Random Over Sampler | SVM | 10-fold | 70.92 |
| 3. | Random Over Sampler | LR | 5-fold | 89.52 |
| 4. | Random Over Sampler | LR | 10-fold | 90.83 |
| 5. | Random Over Sampler | RF | 5-fold | 99.26 |
| 6. | Random Over Sampler | RF | 10-fold | 99.25 |
| 7. | Random Over Sampler | (RF+SVM+LR) | 5-fold | 99.35 |
| 8. | Random Over Sampler | (RF+SVM+LR) | 10-fold | 99.49 |
 |
Figure 6: Performance of different classifiers using Random Over Sampler |
 |
Figure 7: Cross-validation score graph using SFFS |
 |
Figure 8: Cross-validation score graph using SBFS |
Table 3: Performance of Proposed model by using SFFS and RandomOver Selection techniques (6 features)
| Over-Sampling Method | Technique Used | Cross-Validation Technique | No. of features | Accuracy (%) |
| Random Over Sampler | (RF+SVM+LR) | 5-fold | 6 | 99.3379 |
| Random Over Sampler | (RF+SVM+LR) | 10-fold | 6 | 99.5470 |
| Random Over Sampler | (RF+SVM+LR) | Stratified 5-fold | 6 | 99.5296 |
| Random Over Sampler | (RF+SVM+LR) | Stratified 10-fold | 6 | 99.5470 |
Table 4: Performance of the Proposed model Using SBFS and RandomOver Selection techniques (8 features)
| Over-Sampling Method | Technique Used | Cross-Validation Technique | No. of features | Accuracy (%) |
| Random Over Sampler | (RF+SVM+LR) | 5-fold | 8 | 99.42 |
| Random Over Sampler | (RF+SVM+LR) | 10-fold | 8 | 99.46 |
| Random Over Sampler | (RF+SVM+LR) | Stratified 5-fold | 8 | 99.49 |
| Random Over Sampler | (RF+SVM+LR) | Stratified 10-fold | 8 | 99.55 |
Table 5: Performance of Proposed model by using Union of SFFS AND SBFS with RandomOver Selection techniques (11 features)
| Over-Sampling Method | Technique Used | Cross-Validation Technique | No. of features | Accuracy (%) |
| Random Over Sampler | Stacking Classifier
(RF+SVM+LR) |
5-fold | 11 | 99.37 |
| Random Over Sampler | Stacking Classifier
(RF+SVM+LR) |
10-fold | 11 | 99.55 |
| Random Over Sampler | Stacking Classifier
(RF+SVM+LR) |
Stratified 5-fold | 11 | 99.55 |
| Random Over Sampler | Stacking Classifier
(RF+SVM+LR) |
Stratified 10-fold | 11 | 99.55 |
 |
Figure 9: Performance comparison of the proposed model using the ROS technique |
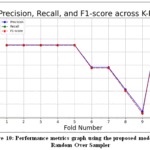 |
Figure 10: Performance metrics graph using the proposed model and Random Over Sampler |
Table 6: Performance of proposed model by using Sequential Forward Selection and SMOTE Techniques (6 Features)
| Over-Sampling Method | Technique Used | Cross-Validation Technique | No. of features | Accuracy (%) |
| SMOTE | Stacking Classifier
(RF+SVM+LR) |
5-fold | 6 | 96.5679 |
| SMOTE | Stacking Classifier
(RF+SVM+LR) |
10-fold | 6 | 97.0557 |
| SMOTE | Stacking Classifier
(RF+SVM+LR) |
Stratified 5-fold | 6 | 97.2996 |
| SMOTE | Stacking Classifier
(RF+SVM+LR) |
Stratified 10-fold | 6 | 97.2299 |
Table 7: Performance of Stacking Classifier by using SBFS and SMOTE techniques (8 features)
| Over-Sampling Method | Technique Used | Cross-Validation Technique | No. of features | Accuracy (%) |
| SMOTE | Stacking Classifier
(RF+SVM+LR) |
5-fold | 8 | 98.2055 |
| SMOTE | Stacking Classifier
(RF+SVM+LR) |
10-fold | 8 | 98.2404 |
| SMOTE | Stacking Classifier
(RF+SVM+LR) |
Stratified 5-fold | 8 | 98.6759 |
| SMOTE | Stacking Classifier
(RF+SVM+LR) |
Stratified 10-fold | 8 | 98.6236 |
Table 8: Performance of the Proposed model by using Union of SFFS and SBFS with SMOTE techniques (11 features)
| Over-Sampling Method | Technique Used | Cross-Validation Technique | No. of features | Accuracy (%) |
| SMOTE | Stacking Classifier
(RF+SVM+LR) |
5-fold | 11 | 98.1881 |
| SMOTE | Stacking Classifier
(RF+SVM+LR) |
10-fold | 11 | 98.3275 |
| SMOTE | Stacking Classifier
(RF+SVM+LR) |
Stratified 5-fold | 11 | 98.4843 |
| SMOTE | Stacking Classifier
(RF+SVM+LR) |
Stratified 10-fold | 11 | 98.6062 |
 |
Figure 11: Performance comparison of the proposed model using the SMOTE technique |
 |
Figure 12: Performance metrics graph using proposed model and SMOTE |
Discussion
The dataset sick-euthyroid consists of 3,163 patient data, which was taken from the UCI-ML repository and has 26 features. Initially, seven features were discarded, six due to irrelevance and one because of a high percentage of missing values. Table 2 and Figure 6 describe the performance comparison of theStacked RSL (S-RSL) model with SVM, LR, and RF. The proposed model performed best with these classifiers.
After that, SFFS and SBFS techniques were applied to select the minimal number of features that show the highest accuracy. SFFS selects six features, and SBFS selects eight features. Figure 7 shows the performance of the SFFS method, with the model achieving optimal accuracy withsix features. After this, the performance remains relatively stable with slight fluctuations. Figure 8 shows the performance of the SBFS method, with the best accuracy observed with eight features selected. Table 3 showsthe S-RSL model’s performance using the SFFS and RandomOverSampler techniques. After SFFS, we implement the SBFS and ROS techniques with our proposed model. Table 4 shows the performance of the S-RSL model using SBFS and ROS. The proposed model is now implemented using 11 features obtained by the union of SFFS and SBFS. Table 5 shows the performance of the S-RSL model using these combined features along with the RandomOverSampler (ROS) technique. Figure9 shows the performance comparison of proposed S-RSL model using the ROS technique, and Figure10 shows the performance metrics across k-folds.
Instead of using RandomOverSampler, SMOTE was implemented to handle class imbalance. All previously applied techniques, including SFFS, SBFS, and the union of selected features, were re-implemented using SMOTE. The performance of the S-RSL model under these configurations is presented in Tables 6, 7, and 8. Among these, the highest accuracy of 98.67% was achieved using SBFS in combination with Stratified 5-Fold Cross-Validation, demonstrating the model’s effectiveness under SMOTE-based oversampling. Figure 11 shows the performancecomparison of the Stacked RSL model using SMOTE technique, and Figure 12 shows the performance metrics across k-folds.
Conclusion
Various research works in the field of thyroid classification based on different ML techniques are mentioned in the literature.To build a robust classifier, there is a need for an ensemble technique which combines the most suitable feature selection methods with the best data balancing strategies. In this work, a Stacked RSL Machine Learning model is proposed using RF, SVM, and LR techniques, followed by Sequential Feature Selection and Oversampling techniques. This research paper highlights the impact of integrating feature selection and ML techniques. The SFFS and SBFS techniques helped in selecting the most relevant features, while the combination of RF, SVM, and LR provided the strengths of multiple models. Multiple cross-validation techniques are implemented to validate the model. Random Over Sampler (ROS) achieved the highest accuracy of 99.55% among various balancing techniques. Also, applying multiple cross-validation techniques shows our proposed model is more stable with 11 features. Future research can investigate deep learning architectures, including neural networks and transformer-based models, to enhance feature extraction and improve prediction accuracy.
Acknowledgement
The author(s)would like to express their gratitude to the Babasaheb Bhimrao Ambedkar University for providing the opportunity for research workand to all others who contributed to its successful completion.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
The dataset used in this manuscript is taken from the open-source UCI ML repository.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable.
Author Contributions
- Gaurav Singh: Analysis, Conception, Technique, and wrote the manuscript;
- Narander Kumar: Visualisation,Supervision, Review & Editing;
- Shishir Kumar: Critical review andSupervision.
References
- Cuadros A, La Torre D, Ticona W. Machine Learning Predictive Model for Thyroid Disease Detection. In: Silhavy R, Silhavy P, eds. Software Engineering Methods Design and Application. Vol 1118. Lecture Notes in Networks and Systems. Springer Nature Switzerland; 2024:267-282. doi:10.1007/978-3-031-70285-3_20
CrossRef - Muñoz-Ortiz Sierra-Cote MC, Zapata-Bravo E, et al. Prevalence of hyperthyroidism, hypothyroidism, and euthyroidism in thyroid eye disease: a systematic review of the literature. Syst Rev. 2020;9(1):201. doi:10.1186/s13643-020-01459-7
CrossRef - Indunil Karunarathna, K Gunawardana, S Rajapaksha, et al. The Euthyroid Sick Syndrome A Diagnostic Challenge in Clinical Practice. Published online 2025. doi:10.13140/RG.2.2.34642.06083
- Da Silveira CD, De Vasconcelos FP, Moura EB, et al. Thyroid Function, Reverse Triiodothyronine, and Mortality in Critically Ill Clinical Patients. Indian Journal of Critical Care Medicine. 2022;25(10):1161-1166. doi:10.5005/jp-journals-10071-24001
CrossRef - Hobbs A, Jack M, Benitez‐Aguirre P, Srinivasan S, Wotton T, Cho YH. Challenges in management of abnormal thyroid function tests in unwell infants: A tertiary centre real‐world experience. J Paediatrics Child Health. 2025;61(2):185-190. doi:10.1111/jpc.16733
CrossRef - Sudar KM, Rohan M, Vignesh K. Detection of adversarial phishing attack using machine learning techniques. Sādhanā. 2024;49(3):232. doi:10.1007/s12046-024-02582-0
CrossRef - Jian W, Li JP, Haq AU, et al. Feature elimination and stacking framework for accurate heart disease detection in IoT healthcare systems using clinical data. Front Med. 2024;11:1362397. doi:10.3389/fmed.2024.1362397
CrossRef - Chaganti R, Rustam F, De La Torre Díez I, Mazón JLV, Rodríguez CL, Ashraf I. Thyroid Disease Prediction Using Selective Features and Machine Learning Techniques. Cancers. 2022;14(16):3914. doi:10.3390/cancers14163914
CrossRef - Ram Kumar RP, Sri Lakshmi M, Ashwak BS, Rajeshwari K, Md Zaid S. Thyroid Disease Classification using Machine Learning Algorithms. Tummala SK, Kosaraju S, Bobba PB, Singh SK, eds. E3S Web Conf. 2023;391:01141. doi:10.1051/e3sconf/202339101141
CrossRef - Sayyid HO, Mahmood SA, Hamadi SS. Comparison of Machine Learning Algorithms for Predicting Thyroid Disorders in Diabetic Patients. IJCAI. 2025;49(12). doi:10.31449/inf.v49i12.6927
CrossRef - Raza A, Eid F, Montero EC, Noya ID, Ashraf I. Enhanced interpretable thyroid disease diagnosis by leveraging synthetic oversampling and machine learning models. BMC Med Inform Decis Mak. 2024;24(1):364. doi:10.1186/s12911-024-02780-0
CrossRef - Sajjan VN, S V, S S. Machine Learning Models for Predicting Hypothyroidism: Utilizing Synthetic Data for Improved Accuracy. In: 2024 Asia Pacific Conference on Innovation in Technology (APCIT). IEEE; 2024:1-5. doi:10.1109/APCIT62007.2024.10673702
CrossRef - Salau AO, Markus ED, Assegie TA, Omeje CO, Eneh JN. Influence of Class Imbalance and Resampling on Classification Accuracy of Chronic Kidney Disease Detection. MMEP. 2023;10(1):48-54. doi:10.18280/mmep.100106
CrossRef - Srivastava R, Kumar P. Optimizing CNN based model for thyroid nodule classification using data augmentation, segmentation and boundary detection techniques. Multimed Tools Appl. 2023;82(26):41037-41072. doi:10.1007/s11042-023-15068-8
CrossRef - Albert AJ, Murugan R, Sripriya T. Diagnosis of heart disease using oversampling methods and decision tree classifier in cardiology. Res Biomed Eng. 2022;39(1):99-113. doi:10.1007/s42600-022-00253-9
CrossRef - Ahamed BS, Arya MS, Nancy AOV. Diabetes Mellitus Disease Prediction Using Machine Learning Classifiers with Oversampling and Feature Augmentation. Troussas C, ed. Advances in Human-Computer Interaction. 2022;2022:1-14. doi:10.1155/2022/9220560
CrossRef - Butt UM, Letchmunan S, Ali M, Hassan FH, Baqir A, Sherazi HHR. Machine Learning Based Diabetes Classification and Prediction for Healthcare Applications. Espino D, ed. Journal of Healthcare Engineering. 2021;2021:1-17. doi:10.1155/2021/9930985
CrossRef - Aversano L, Bernardi ML, Cimitile M, et al. Thyroid Disease Treatment prediction with machine learning approaches. Procedia Computer Science. 2021;192:1031-1040. doi:10.1016/j.procs.2021.08.106
CrossRef - Ionita I, Ionita L. Prediction of Thyroid Disease Using Data Mining Techniques. Broad Research in Artificial Intelligence and Neuroscience. 2016;7(3):115-24.ISSN 2067-3957 (online), ISSN 2068 – 0473 (print)
- Singh GA, Mane VV, Anand R, Sakoglu U. Thyroid Disease Multi-Class Classification. Procedia Computer Science. 2025; 258: 3649-3658. ISSN 1877-0509, doi.org/10.1016/j.procs.2025.04.620.
CrossRef







