Manuscript accepted on :25-03-2026
Published online on: 12-05-2026
Plagiarism Check: Yes
Reviewed by: Dr. Ilya Nikolaevich Medvedev
Second Review by: Dr. Rajendran Susai
Final Approval by: Dr. Anton R Keslav
Kamal Upreti1* , Jossy George1
, Jossy George1 , Bosco Paul Alapatt1
, Bosco Paul Alapatt1 , Rituraj Jain2
, Rituraj Jain2 and Ganeshavishwaa Veluswwamy Radhakrishnan3
and Ganeshavishwaa Veluswwamy Radhakrishnan3
1Department of Computer Science, Chrıst University, Delhi NCR Campus, Ghaziabad, India
2Department of Information Technology, Marwadi University, Rajkot, Gujarat, India
3Department of Economics and Finance, Kalinga Institute of Industrial Technology, Bhubaneswar, India
Corresponding Author E-mail: kamalupreti1989@gmail.com
Abstract
Early identification of abnormal cardiac activity through electrocardiogram (ECG) screening is essential for improving clinical outcomes and enabling timely intervention. Manual ECG interpretation is labor-intensive, subject to inter-observer variability, and difficult to scale for continuous monitoring, highlighting the need for automated screening support. This study presents a deep learning–assisted ECG screening framework based on a hybrid Convolutional Neural Network–Long Short-Term Memory (CNN–LSTM) architecture for automated detection of abnormal cardiac activity. The proposed framework explicitly models short-term temporal dependencies by processing multi-step sequential ECG segments, allowing LSTM layers to learn the evolution of abnormal patterns across consecutive samples. CNN layers extract clinically relevant morphological features related to P-waves, QRS complexes, and T-waves, while LSTM layers capture their temporal progression across three- and six-step windows. Evaluation on a benchmark ECG dataset demonstrates strong screening performance, achieving an accuracy of 99.10%, precision of 99.38%, recall of 99.38%, and an F1-score of 99.38%. Although real-time clinical deployment was not assessed, the lightweight architecture and low inference latency indicate suitability for future wearable and IoT-enabled cardiac monitoring systems.
Keywords
Abnormal Cardiac Activity; Cardiac Signal Analysis; Clinical Decision Support; Deep Learning; Electrocardiogram Screening; Wearable Health Monitoring
| Copy the following to cite this article: Upreti K, George J, Alapatt B. P, Jain R, Radhakrishnan G. V. Deep Learning–Assisted ECG Screening for Detection of Abnormal Cardiac Activity. Biomed Pharmacol J 2026;19(2). |
| Copy the following to cite this URL: Upreti K, George J, Alapatt B. P, Jain R, Radhakrishnan G. V. Deep Learning–Assisted ECG Screening for Detection of Abnormal Cardiac Activity. Biomed Pharmacol J 2026;19(2). Available from: https://bit.ly/49qNB1y |
Introduction
Artificial intelligence (AI) has become a relevant facilitating technology in contemporary healthcare, aiding clinical decision-making, disease screening, and mass health surveillance. Cardiovascular diseases (CVDs) are the foremost cause of death in the world and the cause of death in these diseases amounts to approximately 17.9 million individuals every year. The most common type is coronary heart disease, which leads to the narrowing of blood vessels in the coronary arteries caused by the deposition of atherosclerotic plaque, which reduces the supply of blood to the heart and predisposes it to myocardial infarction and sudden cardiac arrest.1 Coronary heart disease, which is the most common type, is caused by the deposition of atherosclerotic plaque that leads to the limitation of blood flow to the heart, predisposes the heart to myocardial infarction and sudden cardiac arrest Long-term hypertension, ischemic injury, or cardiomyopathies leads to heart failure, and cardiac arrhythmias (e.g. atrial fibrillation) are disturbances in electrical conductivity that can elevate morbidity and mortality substantially.2, 3
Early detection of irregular heart rate is essential in the prevention of the disease development and better patient outcome. ECG is an essential, noninvasive instrument of clinical care that functions to record the electrical activity of the heart and identify disturbances to rhythm, conduction abnormalities, and ischemic alterations.4 Normal cardiac electrical conduction begins in the sinoatrial node and spreads by the Bundle of His, Purkinje fibers, and other standards to generate coordinated contractile activity in the heart. The impairments in any part of this conduction pathway can present as abnormal waveforms of the ECG and usually as a sign of underlying cardiac pathology.4-6 Consequently, ECG-based screening has continued to be a pillar of cardiovascular testing in acute and non-acute care units.
Traditional ECG interpretation is based on the assumptions that visual evaluation of morphology of the waveform is done by an expert clinician using P-waves, QRS complexes and T-waves. This is a time-consuming, inter-observer variant of a clinically effective, but manual process, and it is difficult to scale to continuous or population-wide screening, especially in limited resource settings.7 The mentioned difficulties are exacerbated by the case of long-term ambulatory monitoring and wearable health technologies, where huge amounts of ECG data are obtained on a regular basis. As a result, clinical demands towards automated ECG screening systems have increased in the recent past, which can help medical workers to detect abnormal heart activity and prioritize a case to be examined by specialists in more detail.8
The latest developments in AI and deep learning have significantly enhanced automated ECG analysis, allowing systems to identify complex morphological and temporal features using raw signal data, and assist in early cardiac risk detection. Convolutional Neural Networks (CNNs) are especially useful in estimating spatial and morphological features of the ECG signal, whereas recurrent networks like Long Short-Term Memory (LSTM) networks are less adequate in clinical settings but are useful in continuous cardiac monitoring and early intervention.9,10
Inspired by these advancements, this paper introduces a deep learning-based ECG screening system developed to aid in the identification of abnormal cardiac activity. The suggested method uses a hybrid CNN-LSTM model, where CNN blocks acquire clinically significant morphological characteristics of ECG signals and LSTM nodes learn short-term temporal dynamics among samples in a sequence. It aims to develop a powerful and computationally efficient screening model that would be capable of supporting clinical decision-making and at the same time be applicable in wearable and IoT-enabled healthcare settings in the future. Moreover, this research will give a comparative analysis of the traditional machine learning models such as support vector machines, random forests, and XGBoost with deep learning architectures to frame the performance of the offered hybrid model. Classification accuracy, sensitivity, specificity, and computational efficiency are some of the key performance indices that are analyzed to determine whether the framework is suitable to use in ECG screening issues and not in definitive diagnosis. The rest of the current paper is structured in the following way: Section 2 will conduct a review of related work regarding AI-assisted ECG analysis and uncover gaps in the current research. Section 3 has the description of the materials and methods such as the characteristics of the data set and model architecture, as well as, preprocessing. Section 4 shows results and performance analysis of the experiment. Section 5 covers the implications in clinical setting, limitations, as well as the research directions.
Related Work
Artificial intelligence and machine learning techniques have substantially advanced automated ECG analysis for cardiac abnormality detection and clinical screening. Prior studies have demonstrated that deep learning architectures can outperform traditional signal-processing approaches in identifying abnormal cardiac patterns. For instance, ensemble-based generative adversarial network–LSTM frameworks have shown superior performance compared to classical classifiers, particularly on benchmark ECG datasets.11 Convolutional neural network–based approaches have consistently reported high diagnostic accuracy, exceeding 98% in detecting cardiac abnormalities, including coronary and rhythm-related conditions.12,13 Automated anomaly detection systems have also been applied to support heart disease identification, while hybrid signal-processing and machine learning pipelines, such as wavelet decomposition combined with advanced classifiers, have improved detection of congestive heart failure and myocardial infarction under noisy conditions.14,15
Optimization-assisted and hybrid deep learning models have further enhanced ECG analysis performance.16,17 CNN-based architectures optimized using metaheuristic algorithms have demonstrated accuracies above 99% for ECG abnormality detection.18 Wearable ECG acquisition systems integrated with AI models have enabled physiological and psychological monitoring in ambulatory environments.19 Hybrid CNN–LSTM frameworks have been successfully applied to arrhythmia, congestive heart failure, and normal sinus rhythm detection, highlighting the benefit of combining spatial and temporal feature learning.20 Time–frequency–focused wavelet filter banks and AI-assisted coronary artery disease screening frameworks have also reported strong performance across multiple ECG datasets.21,22 In addition, multimodal pipelines combining ECG with other clinical modalities, such as echocardiography, have demonstrated high discriminative ability for complex cardiac conditions.
More recent studies have explored ensemble learning, multimodal modeling, and large-scale deep learning approaches to extend ECG-based screening capabilities. Ensemble-based deep learning strategies have achieved high accuracy for arrhythmia detection, while dual-input neural networks integrating ECG and phonocardiogram signals have improved coronary artery disease classification.21,22 Variational autoencoder–based ECG reconstruction methods have been applied for myocardial infarction detection using multi-lead ECG recordings.23 Other investigations have focused on heart rate variability analysis derived from wearable photoplethysmography signals for early cardiovascular risk prediction,23,24 as well as energy-efficient ECG monitoring using convolutional and spiking neural networks.25 Large-scale deep learning models trained on extensive ECG datasets have further enabled event prediction, risk stratification, and cardiac phenotyping, achieving high classification accuracy for arrhythmia detection.3,26,27Dual-channel ECG and derivative input improves detection of subtle morphology.28 Cardiac wall motion abnormalities (WMA) are strong predictors of mortality, but current screening methods using Q waves from electrocardiograms (ECGs) have limited accuracy and vary across racial and ethnic groups.29
Collectively, these studies demonstrate the growing role of AI in enhancing ECG-based cardiac screening and decision support. A wide range of approaches, including CNNs, LSTM-based models, hybrid CNN–LSTM architectures, variational autoencoders, and optimized time–frequency analysis techniques, have been reported to improve sensitivity and specificity across diverse cardiac conditions. These methods are summarized in Table 1 with respect to their modeling strategies, target conditions, performance metrics, and datasets.
Table 1: Comparative table of ECG-based disease detection models
| Ref. | Disease Target | Methodology | Performance | Dataset | Research Gap Identified |
| 11 | Heart Disease Detection | GAN + LSTM | Best accuracy & F1-score in simulations | PTB-ECG Dataset | No explicit modeling of short sequential windows; heavy architecture unsuitable for deployment |
| 12 | Cardiac Disorder Diagnosis | CNN | 98.33% Accuracy, Sensitivity 98.33%, Specificity 98.35% | 4000 ECG samples (47 subjects) | Only spatial features learned; no temporal dependency captured |
| 13 | Cardiac Disorder Detection | Lightweight Deep Learning | 98% Precision | 11,148 12-lead ECG scans | Not optimized for single-lead wearable ECG; no hybrid temporal modeling |
| 15 | Congestive Heart Failure | Wavelet Decomposition + QSVM | High precision, sensitivity, specificity | Normal & CHF ECG sets | Traditional ML pipeline; handcrafted features; no end-to-end temporal learning |
| 16 | Myocardial Infarction Detection | SVM + PCA | Sensitivity/Specificity/Accuracy: 96.66% | 60 MI + 60 healthy | Lacks spatial–temporal representation; limited generalization on raw ECG |
| 17 | Myocardial Infarction Detection | Two-band optimal biorthogonal filter | Precision: 99.62% (noisy), 99.74% (clean) | ECG signals | No deep feature extraction; unsuitable for evolving temporal patterns |
| 18 | Cardiac Disorder Detection | CNN + Grapevine Optimization Algorithm | 99.58% Accuracy, 0.42% error | MIT-BIH Arrhythmia | No temporal modeling; optimization adds computational overhead |
| 20 | Arrhythmia, CHF, NSR | 2D-CNN + LSTM | ARR: 98.7%, CHF: 99%, NSR: 99% | Live ECG readings | Handles temporal data but uses 2D formats, not optimized for 1×140 wearable signals |
| 21 | Coronary Artery Disease | Time–frequency wavelet filter bank | 99.53% Accuracy, Sensitivity 98.64%, Precision 99.70% | ECG signals | Requires handcrafted features; lacks real-time deployment feasibility |
Despite these advances, several clinically relevant gaps remain in the existing literature. First, explicit temporal modeling of short ECG sequences is limited, as many approaches treat individual ECG segments as independent samples without capturing how abnormalities evolve across consecutive time steps, particularly in compact single-lead recordings typical of wearable devices. Second, there is insufficient integration of spatial and temporal learning within lightweight architectures optimized for single-lead ECG screening, with many studies emphasizing either morphological or temporal features alone. Third, deployment-oriented considerations such as inference latency, computational efficiency, and suitability for resource-constrained environments are often underreported, despite their importance for real-time or near–real-time clinical screening. Finally, a gap persists between benchmark performance and practical smart-healthcare deployment, as relatively few studies explicitly address integration with wearable sensors, IoT platforms, or edge-based clinical monitoring systems. These limitations highlight the need for clinically oriented, computationally efficient deep learning frameworks that combine spatial and temporal ECG feature learning while remaining suitable for real-world screening applications.
Materials and Methods
The proposed ECG screening framework employs a hybrid Convolutional Neural Network–Long Short-Term Memory (CNN–LSTM) architecture designed to support automated identification of abnormal cardiac activity in wearable and remote monitoring contexts (Figure 1). ECG segments undergo signal quality assessment, morphology-preserving denoising, and normalization to ensure clinically meaningful input. During model development, class imbalance is addressed using the Synthetic Minority Over-Sampling Technique (SMOTE) applied exclusively to training data. CNN layers automatically extract morphological features related to P-waves, QRS complexes, and T-waves, while LSTM units model short-term temporal dependencies across consecutive ECG samples. The resulting spatiotemporal representations are refined using fully connected layers with dropout regularization and classified using a sigmoid-based output to generate a probabilistic screening score. Model performance is evaluated using clinically relevant metrics, including accuracy, sensitivity, specificity, and area under the receiver operating characteristic curve (AUC), supporting assessment of the framework’s suitability as a clinical decision-support tool for ECG screening.
Dataset Description
The dataset used in this model came from Devavrat A. Tripathy’s “ECG Dataset,” which is a publicly accessible Kaggle repository.30 Each row in the dataset represents a complete ECG recording consisting of 140 time-series data points from a single patient, and the same format has also been used in reference,31 which supports our selection of this structure. In this repository, each ECG record is stored as 140 consecutive, uniformly spaced samples of the ECG amplitude collected within a fixed-length time window. Concretely, one row corresponds to the denoised ECG voltage trace sampled at indices inside that window; the 140 columns are therefore raw time-domain samples rather than pre-computed descriptors. Models consuming sequences (CNN/LSTM) operate directly on this 1×140 vector, learning P–QRS–T morphology and temporal dependencies end-to-end. The dataset card does not report the nominal sampling frequency or lead configuration; hence, each row is treated as a fixed-duration segment with uniform sample spacing (140 points per segment) and document this as a limitation. These data points are floating-point numbers that show the heart’s electrical activity throughout time. Another column has a binary label telling us whether the ECG recording is normal (0) or not (1).
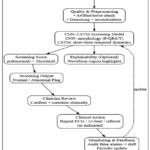 |
Figure 1: Block diagram for the proposed system architecture |
There is no personal identifiable information presented in the dataset, which is completely anonymized. The dataset utilized in this study has 140 time series features corresponding to the electric heart activity of 4,997 ECG recordings (in total, 140 time series features were used). These features play a significant role in the measurement of cardiac functions and can be applied to detect cardiac disease abnormalities. The target variable identifies ECG examples as either abnormal or normal whereby the abnormal cases associate with numerous cardiac abnormalities but no disease is cited. Before modeling, every 140-sample segment is (i) band-limited/denoised, (ii) z-score normalized across segments to zero mean and unit variance (or min-max scaled to [0,1]) and (iii) padded/truncated only when doing so is necessary to maintain a length of 140. This maintains relative morphology and eliminates amplitude offsets between segments.
The data utilized in this research is found publicly on Kaggle and is completely anonymized. The contributors of the dataset were the first to offer the labeling process (normal vs. abnormal), stating that the ECG recordings were acquired in clinically validated equipment and standardized acquisition procedures. The data on the documentation of the dataset did not provide an in-depth annotation or a diagnostic confirmation for each sample on the physician level. To reduce this shortcoming, this study has extensively verified the integrity of the data set by statistical analysis, waveform inspection and cross-linking with literature to identify consistency with clinically accepted ECG patterns.
The ECG recordings were acquired using clinically validated monitoring equipment, ensuring reliable signal quality. Each 140-sample segment preserves the key cardiac waveform components, the P-wave, QRS complex, and T-wave, which are essential for identifying abnormal cardiac activity. The P-wave reflects atrial depolarization and appears as a small positive deflection before the QRS complex in normal ECGs. The QRS complex, representing ventricular depolarization, is a short, high-amplitude peak with a typical duration of 80–120 ms. The T-wave, indicating ventricular repolarization, is normally upright and smoothly returns to baseline. Abnormal ECGs may show inverted or missing P-waves, widened or irregular QRS complexes, or inverted/elevated T-waves. Because these morphological patterns are preserved in each record, the dataset enables the model to learn both normal waveforms and clinically relevant deviations. In order to understand the target variable distribution, figure 2 presents the class frequencies of normal (0) and abnormal (1) ECG recordings. The dataset shows a moderate class imbalance, with abnormal recordings occurring more frequently. This imbalance can bias the classifier toward the majority class; therefore, SMOTE was applied to ensure balanced learning. It also focuses on the imbalance between classes since the number of abnormal ECG recordings exceeds the normal one. Synthetic Minority Over-Sampling Technique (SMOTE) was used in order to tackle the class imbalance. This method can be used to create new synthetic examples of the minority group by interpolating between the existing samples and enhances the capacity of the model to learn the minority classes. SMOTE was chosen compared to other sampling approaches, including ADASYN and Tomek Links, because it is more successful in maintaining the boundaries of the classes decisions and avoiding overfitting, especially in more or less imbalanced clinical data such as the one used in this study. This method facilitated more balanced training and much better classification performance between the traditional and deep learning models.
The publicly available dataset relied upon in the present study has one binary target (0 = normal, 1 = abnormal) and no condition specific labels (e.g., AFib, MI, tachycardia). Therefore, our model screens anomalies, as opposed to doing a differential cardiac diagnosis. Although this option is congruent with the precedent benchmark literature with the same repository, it limits clinical granularity and downstream decision making.
While this study employed a publicly available dataset, these datasets are often used for proof-of-concept validation and benchmarking due to their accessibility and reproducibility, but they may not always capture local demographic or cultural variations. This therefore highlighted as a limitation and note that future work will include testing the proposed framework on real, clinically collected datasets from hospital partners to strengthen the generalizability and clinical adoption of our approach.
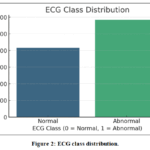 |
Figure 2: ECG class distribution. |
Data Preprocessing
To ensure that the ECG signals provided to the hybrid CNN–LSTM model retained full diagnostic integrity, a carefully controlled preprocessing workflow was applied that focused strictly on cleaning and standardizing the data without removing any medically relevant temporal information. Missing values arising from brief acquisition interruptions were addressed conservatively using simple mean imputation only when required, ensuring that no distortion of the underlying waveform occurred. Mild, morphology-preserving denoising was applied to suppress baseline drift and extreme high-frequency artifacts while explicitly avoiding aggressive filtering procedures that could attenuate key temporal features such as QRS amplitude, ST-segment deflections, or T-wave morphology. Amplitude variability across subjects was normalized using z-score scaling or min–max scaling to [0,1], which harmonizes signal ranges while preserving relative ECG morphology.
To address the imbalance between normal and abnormal recordings, the Synthetic Minority Over-Sampling Technique (SMOTE) was used to generate minority-class samples in feature space without modifying or deleting any time-series samples of the ECG waveform itself. Figure 3 presents the correlation heatmap of the 140 time-domain features, included only for exploratory understanding of feature relationships; however, in line with reviewer guidance and medical-diagnostic best practices, no dimensionality reduction, feature elimination, PCA, or RFE was applied. The model therefore processes the entire 1×140 ECG vector for every recording, ensuring that all clinically relevant temporal characteristics are fully preserved for downstream anomaly detection.
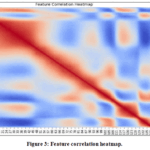 |
Figure 3: Feature correlation heatmap. |
Feature Engineering
To enhance the performance of the model, it must be featured engineered to be better at converting raw ECG signals into useful representations. To improve the accuracy of classification, statistical, frequency-domain, and morphological features were computed in the study. Signal distribution and variability was measured using Statistical Features. Parameters like mean, median, variance, standard deviation, skewness and kurtosis showed ECG signal characteristics. Moreover, since the aberrant cardiac patterns generally possess a greater entropy than the normal rhythms, entropy was calculated to determine the randomness of ECG data. The feature importance plot as shown in figure 4 generated with the help of RF demonstrates the most crucial ECG aspects in order to diagnose cardiac issues. The importance score is used to identify which characteristics impact on classification the most. This has been found valuable in feature engineering as it enables one to pick the most salient features and drop the less important ones to help the model be more interpretable and perform better. In addition to time-domain morphology, frequency-domain characteristics were extracted to capture spectral patterns associated with arrhythmic activity. The Fast Fourier Transform (FFT) was computed on each 140-sample ECG segment to obtain the amplitude spectrum within the physiologically relevant ECG frequency range of 0.5–40 Hz. From the 140-point FFT output, the first 70 unique coefficients (up to the Nyquist limit) were retained as spectral features.
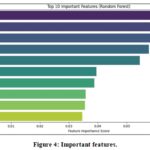 |
Figure 4: Important features. |
To analyze transient, non-stationary components in the signal, the Discrete Wavelet Transform (DWT) was also applied using a Daubechies-4 (db4) mother wavelet, which is widely used in ECG analysis. The ECG waveform was decomposed into approximation (A1) and detail (D1–D3) coefficients, capturing low-frequency P-wave/T-wave content and high-frequency QRS components. This resulted in a total of (A1 + D1 + D2 + D3) features per segment, representing multi-resolution frequency information. These spectral features complement the time-domain patterns learned by the CNN–LSTM model and help characterize both stationary and transient abnormalities.
Power Spectral Density (PSD) was also done to determine the energy distribution across the various frequency bands which gave more informative data on cardiac conditions. Morphological Features were found to represent the characteristics of ECG waveforms in terms of their structure. Significant variables including T-wave morphology, QRS complex width, and P-wave duration and amplitude were examined as alterations in these variables are trusted indicators of cardiac issues. RR interval records, a measure of cardiac variability of heart rate (HRV), were also included to measure abnormal pattern of heartbeats that is related to arrhythmia. In order to ensure the consistency of high-dimensional ECG data, PCA and autoencoders were used as forms of dimensionality reduction in order to retain the basic features and remove the less significant ones, which enhanced the effectiveness and understanding of models.
Figure 5 indicates the percentage of information that was removed with regard to the original ECG data as each additional component to include in principle component modeling was added. The graph shows that the variance explained improves with addition of more components but at some stage the ratio becomes insignificant. This helps to establish the optimal quantity of components that the researcher needs to reduce the dimensions without distorting any crucial features of the ECG signals. The model is able to operate efficiently without losing valuable information useful in diagnostics by identifying this balance, which increases interpretability and computational efficiency.
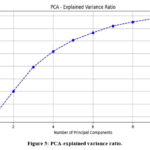 |
Figure 5: PCA-explained variance ratio. |
In addition to using the raw 140 ECG signal samples as input for deep learning models, several engineered feature groups were derived to support classical machine learning approaches. First, 11 statistical features were computed to summarize the global characteristics of the ECG signal, including mean, median, variance, standard deviation, coefficient of variation, minimum, maximum, range, skewness, kurtosis, and sample entropy. Second, 12 temporal and morphological features were extracted to describe the shape and rhythm of the signal, such as number of peaks, maximum peak amplitude, average inter-peak distance, estimated QRS complex width, rise and fall time, mean and variance of slope, zero-crossing count, baseline wander index, T-wave segment energy, and outlier fraction. Third, 16 frequency and time–frequency features were derived using Fast Fourier Transform (FFT) and Wavelet Transform to capture spectral patterns, including band energies, spectral centroid, bandwidth, flatness, dominant frequency, PSD, and multi-level wavelet coefficients. Finally, 10 principal components obtained through PCA were used to retain the most informative signal representations while reducing dimensionality. Together, these engineered features provide complementary statistical, morphological, and spectral information about ECG signals, enabling classical machine learning models to better differentiate normal and abnormal patterns, while the deep learning models learn directly from the 140 raw time-series samples.
Model Selection
To identify the most effective architecture for ECG-based cardiac abnormality detection, several machine learning and deep learning models were systematically evaluated. Traditional ML models such as SVM, RF, and XGBoost were first tested to establish baseline performance. SVM was considered for its ability to form non-linear decision boundaries through kernel transformations, although its performance required careful tuning of kernel type and regularization parameters. RF offered robustness through ensemble aggregation and provided interpretable feature-importance insights, while XGBoost demonstrated strong handling of imbalanced data through gradient boosting and regularization.
Following the evaluation of baseline models, deep learning approaches were explored for their capacity to learn discriminative features directly from raw ECG sequences. CNNs effectively extracted spatial waveform patterns such as peaks and segment transitions, whereas LSTM networks were capable of modeling sequential dependencies across the 140-sample ECG window. As illustrated in figure 6, the comparative model selection workflow involved training, validation, and performance assessment based on predefined criteria. Through this structured selection process, the hybrid CNN–LSTM model emerged as the most suitable, as it combines spatial morphology extraction with temporal dependency learning, consistently meeting all selection thresholds.
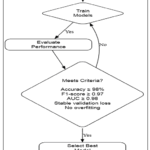 |
Figure 6: Flowchart of the model selection process based on training, performance evaluation, and predefined selection criteria. |
Model Evaluation and Selection
The models have undergone testing with a strict process applying move-validation strategies to ensure stability across exceptional sets of data. Hyper parameter tuning became utilized through Grid Search and Random Search, optimizing mastering fee, batch length, and community depth parameters for exceptional performance. CNN layers improved classification accuracy by automatically detecting important waveform features including peaks and segments, eliminating the need for manually created feature extraction. LSTM Networks are used in ECG data detection due to their ability to manage sequential dependencies. Unlike other recurrent neural networks, LSTMs minimize the vanishing gradient problem, preserving long-term dependencies. The LSTM model has two stacked layers with 128 and 64 hidden units, while the hybrid CNN-LSTM design connects two convolutional layers. The last parameters are the 64-batch size, Adam optimizer, 100 training epochs, the learning rate of 0.001 and a dropout rate of 0.5. Best-performing weights are kept in model checkpoints and early pausing prevents overfitting.
The accuracy, precision, recall, F1-score, and the ROC curve were used to measure the performance of a deep ECG model. In this study, 10-fold cross-validation was used for model evaluation because it offers an appropriate trade-off between training sample availability and validation stability. Although 5-fold and 8-fold configurations were initially tested, they exhibited greater variance in validation accuracy and F1-score. The 10-fold setting consistently produced more stable and reliable performance estimates while ensuring suitable representation of both normal and abnormal ECG segments within each fold. The hybrid CNN-LSTM version was chosen for accurate extraction of spatial and temporal ECG functions. This methodical approach to model selection ensures that the chosen version is highly optimized for implementation in actual worldwide clinical settings, promoting AI-assisted cardiac prediction and better outcomes for impacted individuals.
Results
This section presents a comprehensive evaluation of machine learning and deep learning models for ECG-based screening of abnormal cardiac activity. Model performance was assessed using clinically relevant metrics, including accuracy, precision, recall (sensitivity), and F1-score, to examine the ability of each approach to differentiate normal from abnormal ECG recordings. In addition to classification performance, practical considerations such as model robustness, interpretability, and computational feasibility were evaluated to assess suitability for ECG screening and decision-support applications.
Descriptive Analysis of the Dataset and ECG Signals
The ECG dataset consisted of 4,997 single-lead recordings, each recording being represented by 140 time-domain samples, i.e. sequential points of ECG waveform. ECG records were classified into two groups, which included normal (0) and abnormal (1). Visual and statistical analysis of ECG recordings showed that normal recordings tended to have well-defined P-waves, QRS complex and T-waves with uniform morphology. On the contrary, abnormal records showed irregularities in waveforms, such as changed amplitudes, long or distorted QRS complexes, and changes in shape of waveforms, which are indicative of abnormal cardiac activity. The morphological differences were identified by comparative visualization of normal and abnormal ECG segments representing the norm. Moreover, the exploratory analysis of amplitude, duration, and frequency distributions also helped to confirm that there were two distinct signal properties in the two classes.
Screening Performance of Machine Learning and Deep Learning Models
Many machine learning models (support vector machines, random forests, and XGBoost) and deep learning models (CNN, LSTM, and their hybrid CNN-LSTM) were compared. In general, deep learning models showed high results in screening because they are able to take advantage of spatial and temporal features of ECG signals. Specifically, the hybrid CNN-LSTM model demonstrated a good trade-off between the sensitivity and specificity by combining the morphological feature extraction with the short-term temporal dependency modeling. Although the traditional machine learning models also obtained high classification accuracy, the CNN-LSTM architecture was more robust and showed more consistent results across validation folds. Even though the accuracy enhancement of the hybrid model compared to the standalone CNN model was insignificant and not statistically relevant at the 0.05 level, the hybrid model had lower variability and was more stable, which makes it suitable to ECG screening applications where it is clinically relevant to consistently detect abnormal cardiac activity.
Table 2 provides a summary of the screening performance of machine learning and deep learning models based on multiple clinically relevant evaluation metrics of identifying abnormal cardiac activity using ECG recordings. In all models, all high values of accuracy (98.8%) show that there is a high overall capability to differentiate between normal and abnormal ECG pattern. In a clinical screening situation, however, different metrics, including sensitivity, specificity, and control of false-negative, are more important than accuracy itself. The old models of machine learning, such as support vector machines (SVM), random forests (RF), and XGBoost, showed high levels of accuracy in classification, with the RF reaching the highest overall accuracy (0.996) and close to perfect AUC-ROC (0.999). Such findings demonstrate high levels of discriminative power but it is possible that such models that are accuracy-dominant do not necessarily optimize the sensitivity-specificity relationship that is needed to assure safe clinical screening.
Table 2: Screening Performance of Evaluated Models.
| Model | Accuracy | Precision | Recall (Sensitivity) | Specificity | F1-Score | Cohen’s Kappa | MCC | AUC-ROC |
| SVM | 0.994 | 1.000 | 0.990 | 1.000 | 0.995 | 0.988 | 0.988 | 0.998 |
| RF | 0.996 | 0.995 | 0.998 | 0.993 | 0.997 | 0.992 | 0.992 | 0.999 |
| XGBoost | 0.994 | 0.993 | 0.997 | 0.990 | 0.995 | 0.988 | 0.988 | 0.998 |
| CNN | 0.993 | 0.991 | 0.997 | 0.988 | 0.994 | 0.986 | 0.986 | 0.997 |
| LSTM | 0.988 | 0.983 | 0.997 | 0.976 | 0.990 | 0.975 | 0.975 | 0.996 |
| CNN-LSTM | 0.990 | 0.988 | 0.995 | 0.983 | 0.991 | 0.979 | 0.979 | 0.998 |
The CNN-LSTM model had a clinically desirable sensitivity (0.995) and specificity (0.983) showing a reliable ability to recognize abnormal ECG records and a lower rate of false alarms. The large AUC-ROC (0.998) and Matthews Correlation Coefficient (0.979) are also evidence of much and solid discrimination of decision thresholds, even in the case of class-imbalanced problems. Though the standalone CNN showed a slightly higher accuracy, the hybrid CNN-LSTM showed elevated consistency and temporal strength when morphological and sequential features of ECG were combined. These results indicate that a number of models give high screening accuracy, but the hybrid CNN-LSTM framework gives a more balanced and clinically viable screening profile, indicating its future use as a decision-support tool to detect abnormalities using ECGs and not as a diagnostic system.
Discussion
This study explored the use of a deep learning-assisted model to ECG-based screening of abnormal heart activity with particular focus on clinical reliability, robustness, and decision-support (not definitive diagnosis) application. The findings indicate that it is possible to screen with high performance using automated analysis of small single-lead ECG segments with a hybrid CNN-LSTM architecture and retain the clinically relevant morphology and temporal dynamics of waveforms.
One of the major results of the given study is the high sensitivity of the suggested framework, which means that the abnormal ECG recordings can be identified reliably. The sensitivity and minimizing false-negatives are of interest in a situation of clinical screening where missing abnormal cases can postpone further diagnosis and proper clinical treatment. CNN-LSTM model demonstrated a positive balance on sensitivity and specificity, which indicates that it can be effectively used to detect abnormal cardiac activity and reduce false alarms, which would not be necessary. The large AUC-ROC and Matthews Correlation Coefficient also demonstrates that discrimination remains consistent at the various decision-thresholds, and that discrimination is also effective in class-imbalanced situations which occur with real-world ECG screening datasets.
Convolutional and recurrent learning components have their benefits when integrated to complement each other in the ECG analysis. CNN layers allow extracting morphological features related to P-waves, QRS complexes, and T-waves automatically, and LSTM units are able to capture short-term time correlations that indicate changes in rhythm patterns between successive samples. Whereas the accuracy improvement of the hybrid architecture over a single CNN model was relatively small, there was an enhanced consistency between validation folds, which is why the hybrid architecture can be useful in screening applications where reliability and robustness are clinically relevant. The results of these findings are consistent with the results of earlier studies that suggested that combined spatial-temporal modeling can be more effective to detect ECG abnormalities, especially with small single-lead recordings.
Clinically speaking, the proposed framework can best be understood as a screening/triage support device and not a substitution of a skilled interpretation of ECG. The binary normal/abnormal output fits well with early-stage screening processes where automated systems are applied to assign ECG segments to additional clinical examination. This method is especially applicable in the realms of ambulatory monitoring, wearable devices and telemedicine platforms, where large amounts of ECG data are produced, and it is practically impossible to manually review all the segments. The framework can help earlier detect the possible abnormal recordings and allocate clinical resources more efficiently by helping clinicians.
Although such promising outcomes are evident, there are some limitations that should be recognized. The data that has been utilized in this research gives binary labels without annotation to diseases, which restricts clinical specificity of the screening results. This, therefore, means that the framework is unable to distinguish between particular cardiac conditions, including atrial fibrillation or myocardial infarction and should not be applied as an independent diagnostic system. Also, despite the publicly available and commonly used dataset to benchmark, lack of details about patient demographics, the settings of acquisition, and clinical settings may influence generalizability to different populations. Until it can be applied in a real world clinical setting, external validation with clinically annotated multi-center ECG data must be done.
Future directions The framework should be expanded to multi-class classification with clinician-verified labels, the performance should be assessed on a variety of patient populations, and the explainability methods should be incorporated to increase clinician trust and interpretability. The clinical relevance of the proposed approach would be further supported by validation in a prospective or real-world monitoring environment especially in a wearable and remote care setting. Properly validated and governed, deep learning-complemented ECG screening systems like that introduced in this paper can potentially contribute to scalable, timely, and clinically relevant cardiac screening.
Conclusion
This work provided a deep learning-based ECG screening model that relies on a hybrid CNN-LSTM model to detect abnormal heart activity. The proposed method that combined CNN-based morphological feature extraction with LSTM-based temporal pattern learning demonstrated a high screening high performance on short single-lead ECG segments. The findings suggest that automated ECG signal analysis can be used reliably in helping to identify abnormal cardiac patterns without compromising clinically useful waveform properties. Though the traditional machine learning models had less computational requirements, they do not have the capacity to efficiently represent temporal dependencies that have significance in ECG rhythm interpretation. However, the hybrid CNN-LSTM offers a compromise that offers better screening activities, and at the same time, it is feasible to implement in the context of wearables and remote monitoring.
Clinically, the proposed framework is mandated as a screening/decision-support tool and not as a system of its own diagnosis. Binary normal/abnormal design declares reduce diagnostic specificity at the point of use, because this model fails to differentiate between each distinct cardiac condition. Further development of the framework will include additional classification into multi-class with clinical-annotated data, external validation on various population groups and acquisition environments, and the assessment of model calibration and fairness concerning demographic subgroups. The clinically verified ECG datasets require additional validation to improve real-world reliability, as well as in order to facilitate safe integration into telemedicine and mobile health ecosystems.
Acknowledgment
The authors gratefully acknowledge Devavrat A. Tripathy for providing the ECG Dataset, which is publicly available through the Kaggle platform (https://www.kaggle.com/datasets/devavratatripathy/ecg-dataset) and was used in this study.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials
Permission to reproduce material from other sources
Not Applicable
Author Contributions
- Kamal Upreti: Conceptualization, Formal analysis, Methodology, Validation, Visualization, Writing – original draft, Writing – review & editing.
- Jossy George: Conceptualization, Formal analysis, Methodology, Validation, Visualization, Writing – original draft, Writing – review & editing.
- Bosco Paul Alapatt: Conceptualization, Formal analysis, Methodology, Validation, Visualization, Writing – original draft, Writing – review & editing.
- Rituraj Jain: Conceptualization, Formal analysis, Methodology, Validation, Visualization, Writing – original draft, Writing – review & editing
- Ganeshavishwaa Veluswwamy Radhakrishnan: Conceptualization, Formal analysis, Methodology, Validation, Visualization, Writing – original draft, Writing – review & editing
References
- Ullah A, Anwar SM, Bilal M, Mehmood RM. Classification of arrhythmia by using deep learning with 2-D ECG spectral image representation. Remote Sensing. 2020;12(10):1685. doi:10.3390/rs12101685.
CrossRef - Mamun MMRK, Elfouly T. AI-enabled electrocardiogram analysis for disease diagnosis. Applied System Innovation. 2023;6(5):95. doi:10.3390/asi6050095.
CrossRef - Pandey SK, Janghel RR. Automatic detection of arrhythmia from imbalanced ECG database using CNN model with SMOTE. Australasian Physical & Engineering Sciences in Medicine. 2019;42(4):1129-1139. doi:10.1007/s13246-019-00815-9.
CrossRef - Bibi A, Rahman JSU. Machine learning-enabled in-home ECG: A review. Medinformatics. 2025. doi:10.47852/bonviewmedin42024336.
CrossRef - Hannun AY, Rajpurkar P, Haghpanahi M, et al. Cardiologist-level arrhythmia detection and classification in ambulatory electrocardiograms using a deep neural network. Nature Medicine. 2018;25(1):65-69. doi:10.1038/s41591-018-0268-3.
CrossRef - M G, Ravi V, Vishvanathan S, GopalakrishnanEA, KP S. Explainable deep learning-based approach for multilabel classification of electrocardiogram. IEEE Transactions on Engineering Management. 2021;70(8):2787-2799. doi:10.1109/TEM.2021.3104751.
CrossRef - Gupta V, Mittal M, Mittal V. R-peak detection-based chaos analysis of ECG signal. Analog Integrated Circuits and Signal Processing. 2019;102(3):479-490. doi:10.1007/s10470-019-01556-1.
CrossRef - Chen TM, Huang CH, Shih ESC, Hu YF, Hwang MJ. Detection and classification of cardiac arrhythmias by a challenge-best deep learning neural network model. iScience. 2020;23(3):100886. doi:10.1016/j.isci.2020.100886.
CrossRef - Jahmunah V, Ng EYK, Tan RS, Oh SL, Acharya UR. Explainable detection of myocardial infarction using deep learning models with Grad-CAM technique on ECG signals. Computers in Biology and Medicine. 2022;146:105550. doi:10.1016/j.compbiomed.2022.105550.
CrossRef - Ko WY, Siontis KC, Attia ZI, et al. Detection of hypertrophic cardiomyopathy using a convolutional neural network-enabled electrocardiogram. Journal of the American College of Cardiology. 2020;75(7):722-733. doi:10.1016/j.jacc.2019.12.030.
CrossRef - Rath A, Mishra D, Panda G, Satapathy SC. Heart disease detection using deep learning methods from imbalanced ECG samples. Biomedical Signal Processing and Control. 2021;68:102820. doi:10.1016/j.bspc.2021.102820.
CrossRef - Avanzato R, Beritelli F. Automatic ECG diagnosis using convolutional neural network. Electronics. 2020;9(6):951. doi:10.3390/electronics9060951.
CrossRef - Khan AH, Hussain M, Malik MK. Cardiac disorder classification by electrocardiogram sensing using deep neural network. Complexity. 2021;2021:5512243. doi:10.1155/2021/5512243.
CrossRef - Elliott PM, Anastasakis A, Borger MA, et al. 2014 ESC guidelines on diagnosis and management of hypertrophic cardiomyopathy. European Heart Journal. 2014;35(39):2733-2779. doi:10.1093/eurheartj/ehu284.
CrossRef - Bhurane A, Sharma M, San-Tan R, Acharya UR. Efficient detection of congestive heart failure using frequency-localized filter banks with ECG signals. Cognitive Systems Research. 2019;55:82-94. doi:10.1016/j.cogsys.2018.12.017.
CrossRef - Dohare K, Kumar V, Kumar R. Detection of myocardial infarction in 12-lead ECG using support vector machine. Applied Soft Computing. 2017;64:138-147. doi:10.1016/j.asoc.2017.12.001.
CrossRef - Sharma M, Tan RS, Acharya UR. Automated diagnostic system for myocardial infarction classification using optimal biorthogonal filter banks. Computers in Biology and Medicine. 2018;102:341-356. doi:10.1016/j.compbiomed.2018.07.005.
CrossRef - Tyagi A, Mehra R. Heartbeat classification model for heart disease diagnosis using hybrid CNN with grasshopper optimization algorithm. SN Applied Sciences. 2021;3(2). doi:10.1007/s42452-021-04185-4.
CrossRef - Heyat MBB, Akhtar F, Abbas SJ, et al. Wearable flexible electronics-based cardiac electrode for mental stress detection using machine learning on single-lead ECG. Biosensors. 2022;12(6):427. doi:10.3390/bios12060427.
CrossRef - Madan P, Singh V, Singh DP, Diwakar M, Pant B, Kishor A. Hybrid deep learning approach for ECG-based arrhythmia classification. Bioengineering. 2022;9(4):152. doi:10.3390/bioengineering9040152.
CrossRef - Sharma M, Acharya UR. Identification of coronary artery disease using ECG signals and time-frequency concentrated antisymmetric biorthogonal wavelet filter bank. Pattern Recognition Letters. 2019;125:235-240. doi:10.1016/j.patrec.2019.04.014.
CrossRef - Kayamk S, Vulasala R. Coronary artery blockage detection using artificial intelligence algorithms. International Journal of Research in Pharmaceutical Sciences. 2020;11(1):471-479. doi:10.26452/ijrps.v11i1.1844.
CrossRef - Cho Y, Kwon JM, Kim KH, et al. Artificial intelligence algorithm for detecting myocardial infarction using six-lead electrocardiography. Scientific Reports. 2020;10(1). doi:10.1038/s41598-020-77599-6.
CrossRef - Lan KC, Raknim P, Kao WF, Huang JH. Toward hypertension prediction based on PPG-derived HRV signals: A feasibility study. Journal of Medical Systems. 2018;42(6). doi:10.1007/s10916-018-0942-5.
CrossRef - Yan Z, Zhou J, Wong WF. Energy-efficient ECG classification with spiking neural networks. Biomedical Signal Processing and Control. 2020;63:102170. doi:10.1016/j.bspc.2020.102170.
CrossRef - Hazratifard M, Gebali F, Mamun M. Machine learning for dynamic authentication in telehealth: A tutorial. Sensors. 2022;22(19):7655. doi:10.3390/s22197655.
CrossRef - Ammar K, Fraihat S, Al-Naymat G, Sanjalawe Y. ECG-CBA: End-to-end deep learning model for ECG anomaly detection using CNN, Bi-LSTM, and attention mechanism. Algorithms. 2025;18(11):674. doi:10.3390/a18110674.
CrossRef - Bashar, Syed Khairul, et al. “Premature Atrial and Ventricular Contraction Detection Using Deep Learning and Short ECG: A Multi-dataset Evaluation.” Biomedical Signal Processing and Control, vol. 113, Oct. 2025, p. 108956, doi:10.1016/j.bspc.2025.108956.
CrossRef - Rogers, Albert J., et al. “Identification of Cardiac Wall Motion Abnormalities in Diverse Populations by Deep Learning of the Electrocardiogram.” Npj Digital Medicine, vol. 8, no. 1, Jan. 2025, p. 21, doi:10.1038/s41746-024-01407-y.
CrossRef - Tripathy D. ECG dataset. Kaggle.com. Published 2021. Accessed December 31, 2025. https://www.kaggle.com/datasets/devavratatripathy/ecg-dataset/data
- Sliti W, Abdelali SEB, Yahyaoui A, Mosbah A, Djebbi O. Cardiovascular anomaly detection using deep learning techniques. In: Lecture Notes in Computer Science. 2023:286–299. doi:10.1007/978-3-031-49333-1_21.
CrossRef







