Manuscript accepted on :01-01-2026
Published online on: 14-01-2026
Plagiarism Check: Yes
Reviewed by: Dr. Hassan Shora
Second Review by: Dr. Ahmed Salah
Final Approval by: Dr. Anton R Keslav
Shruthi Mittemari Lakshminarayan Jagannath1,2* , Nishita Nitin Joshi1,2
, Nishita Nitin Joshi1,2 , Tanushree Giridharan1,2
, Tanushree Giridharan1,2 and Nirmala Devi Manickam1
and Nirmala Devi Manickam1
1Department of Electronics and Communication Engineering, PES University, Bengaluru, India
2Centre for Healthcare Engineering and Learning, PES University, Bengaluru, India
*Corresponding Author E-mail: shruthimlj@pes.edu
DOI : https://dx.doi.org/10.13005/bpj/3363
Abstract
In real-world scenarios, clinical trials provide imbalanced datasets and limited labelled data for the diagnostic evaluation of various medical conditions. Especially in detection of Diabetes Mellitus (DM), which is globally prevalent, the challenge becomes multi-fold in resource-constrained environments. To address these challenges, the study proposes a hybrid framework combining Condensed Nearest Neighbor (CoNN) with Few-Shot Learning (FSL), designed to improve detection speed and reduce memory usage without compromising diagnostic performance. Using publicly available datasets, the framework’s performance was compared with multiple Machine Learning (ML) approaches with an emphasis on preprocessing techniques such as imputation, oversampling, and feature reduction. Compared to conventional models, the proposed CoNN-FSL framework used 1/15th of the total samples with 2.5 times improvement in terms of training speed. The study offers a comprehensive evaluation of strategies, enhancing the training speed and reducing the storage requirements. Together, these advancements make Machine Learning models more practical and scalable for real-world clinical applications.
Keywords
Condensed Nearest Neighbor; Cross-Sectional Studies; Data Preprocessing; Diabetes Mellitus; Few-Shot Learning; Machine Learning
Download this article as:| Copy the following to cite this article: Jagannath S. M. L, Joshi N. N, Giridharan T, Manickam N. D. Type 2 Diabetes Mellitus Detection using Heterogeneous Machine Learning Models: A Cross-sectional Study. Biomed Pharmacol J 2026;19(1). |
| Copy the following to cite this URL: Jagannath S. M. L, Joshi N. N, Giridharan T, Manickam N. D. Type 2 Diabetes Mellitus Detection using Heterogeneous Machine Learning Models: A Cross-sectional Study. Biomed Pharmacol J 2026;19(1). Available from: https://bit.ly/4sGnSdt |
Introduction
Diabetes Mellitus is a metabolic disease characterized by chronic hyperglycemia resulting from insulin deficiency or resistance, with complications including cardiovascular disease, nephropathy, neuropathy, and retinopathy.1,2 The 2021 International Diabetes Federation (IDF) report estimated 537 million diabetic adults in 2021, forecasting 643 and 783 million cases in 2030 and 2045, respectively.3 The increasing prevalence of diabetes necessitates early diagnosis and cure.
Diabetes is classified into Type 1, Type 2, and gestational diabetes. Type 1 diabetes is an autoimmune condition that destroys pancreatic beta cells, requiring lifelong insulin therapy.4 Type 2 diabetes is linked to obesity, physical inactivity, and genetics.5 Gestational diabetes develops during pregnancy and increases the risk of Type 2 diabetes in both mother and offspring.6 Prediabetes is a condition where plasma glucose is elevated but not considered diabetic, increases the risk of Type 2 diabetes if no precautions are taken.7
Early diagnosis is critical in preventing complications. Traditional diagnostic methods include fasting blood glucose (FBG), oral glucose tolerance tests (OGTT), and HbA1c assessments.8 Physiological indicators such as body mass index (BMI), blood glucose levels, blood pressure, and cholesterol are vital for assessing diabetes risk. Machine learning (ML) in medical diagnosis has revolutionized the field by enabling quick, precise, and evidence-based decision-making.9 Anjana et al. (2011) reported increased diabetes prevalence among Indian urban populations due to sedentary lifestyles, poor diet, and high rates of obesity. The research identified significant diabetic characteristics such as BMI, blood pressure, and fasting glucose levels.10 ML enhances predictive precision, identifies risk factors, and improves clinical outcomes.11 The 2025 American Diabetes Association report highlights the growing role of ML-based glycemic assessments in diabetes care.12
Several studies have demonstrated ML models’ proficiency in diabetes detection. Bhoi et al. applied multiple supervised learning algorithms to predict diabetes in Pima Indian females, identifying glucose level as the most influential factor. Among the models, k-NN achieved the highest accuracy at 83.12%.13 Jian et al. achieved 97.8% accuracy in predicting diabetes complications using ML models.14 Patil et al. compared several classification algorithms for diabetes prediction, including Logistic Regression, k-NN, SVM, and Random Forest(RF). They found that Logistic Regression and Gradient Boosting achieved the highest accuracy of 78.36%.15 Guan et al. employed binomial logistic regression and regression trees, achieving an accuracy of 77.48% with logistic regression.16 The review conducted by Daza et al. on Type 2 Diabetes Mellitus reported Random Forest was identified as the most efficient and accurate algorithm across multiple evaluations. However, the study does not address computational aspects such as training time or resource requirements, an important limitation given the focus on efficiency in deployment contexts.
Advanced methods such as ensemble learning and deep learning (DL) further enhance diabetes prediction. Alghamdi demonstrated XGBoost’s efficacy in classifying high-risk patients with 89% accuracy,17 while Sabejon et al. achieved 99.03% accuracy using XGBoost on clinical data.18 Doğru et al. proposed a hybrid super ensemble model combining logistic regression, decision tree, random forest, and gradient boosting with SVM for early diabetes prediction achieving 92% accuracy across PIMA dataset.19
Naz and Ahuja emphasized DL’s potential in early diabetes detection.20 Sarwar et al. developed an ensemble-based expert system to diagnose type-II diabetes, integrating classifiers such as ANN, SVM, KNN, and Naïve Bayes. Their approach achieved an accuracy of 98.60%, outperforming individual classifiers.21 Similarly, Reza et al. proposed a stacking ensemble method incorporating deep neural networks, reaching 95.5% accuracy.22 Lakhwani et al. implemented a three-layer ANN with the Quasi-Newton training method on PIDD, yielding competitive accuracy.23 Rajni and Amandeep introduced an RB-Bayes algorithm that outperformed NB, SVM, and KNN models by addressing the zero-probability issue in Naïve Bayes classification.24 Shams et al. proposed an RFE-GRU model for diabetes classification, effectively selecting relevant features and handling class imbalances, achieving an AUC of 92.78%.25 Qaraqe et al. developed a convolutional neural network adapted from an FSL model for long-term HbA1c prediction, achieving 93.2% accuracy.26 Study by Sosale et al. also highlights diabetes complications, emphasizing early intervention strategies.27 Fregoso-Aparicio et al. found that the structure and balance of datasets significantly influenced the accuracy of diabetes prediction models. Tree based models mostly RF and DT required data balancing and feature selection and deep learning models were found to perform well on datasets with more than 70,000 samples. The importance of AUC-ROC curve as evaluation metric was highlighted to minimize variability in performance comparison. The study was limited by heterogeneity in sample sizes and populations across models, and by a lack of focus on predicting diabetes complications.28 The review conducted by Daza et al. on Type 2 Diabetes Mellitus reported Random Forest was identified as the most efficient and accurate algorithm across multiple evaluations. However, the study does not address computational aspects such as training time or resource requirements, an important limitation given the focus on efficiency in deployment contexts.29
As diabetes impacts more people globally, ML integration into clinical decision-making can enhance early detection, minimize complications, and improve patient outcomes. This study aims to leverage ML and DL methods based on key physiological parameters to enhance diagnostic accuracy, predict risks, and optimize training efficiency.
Materials and Methods
The workflow diagram as shown in Figure 1 displays the procedure for diabetes prediction using ML. The input data consists of PIDD and DPD.30,31 Data preprocessing includes imputation methods (zero, mean, and median imputation) and re-sampling via SMOTE to address class imbalance. Feature reduction is materialized using heatmaps and correlation analysis. Performance analysis is conducted to evaluate the detection ability of models in identifying diabetic patients.
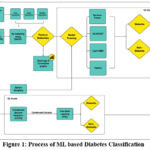 |
Figure 1: Process of ML based Diabetes Classification
|
Traditional ML algorithms are based on large datasets, containing redundant or less informative data and demand high computational resources. Training from such data is usually time-consuming, and iterative model refinement is difficult. To address this, a hybrid framework that integrates CoNN with FSL for detection that significantly reduces training time without compromising decision-making capability has been proposed.
Data Collection
The samples were acquired from publicly available datasets. PIDD has 768 samples (268 diabetic instances as 1 and 500 non-diabetic instances as 0) with eight medical features: Pregnancies, Glucose, Blood Pressure, Skin Thickness, Insulin, BMI, Diabetes Pedigree Function and Age. DPD contains medical records of 100,000 patients from different age groups, that includes eight clinical features commonly associated with diabetes risk factors. These features include categorical and numerical variables like Gender, Age, Hypertension, Heart Disease, Smoking History, BMI, HbA1c Level, Blood Glucose Level, and Diabetes (0,1 for non-diabetic and diabetic individuals respectively).
Data Preprocessing
The data preprocessing methods such as normalization, missing value handling, feature selection, and resampling were incorporated to meet the specific requirements of the analysis and improve model performance. Null values and unwanted information were removed or imputed based on the dataset requirement. These techniques ensure that the dataset remains complete, which can enhance the accuracy of the statistical analysis.
Resampling
The raw dataset included fewer diabetic patients than non-diabetic ones; therefore, the dataset was resampled to balance it. The resampled dataset contains 1.8 Lakh patients with equal attributes. Zero, Mean and median imputation were used to address the missing value issue, resulting in three distinct datasets. Though all these approaches were promising, the median-based method performed the best, as presented in the Results and Discussion section. By applying SMOTE, the model’s ability to learn from minority class patterns will be enhanced, reducing bias while maintaining the data distribution. Several studies have discussed the impact of SMOTE on model performances.32-34
Feature Reduction
To optimize predictive efficacy and accuracy of models, feature selection was performed on PIDD and DPD using correlation heatmaps. It has been proposed to remove features that offered less predictive information or brought redundancy, to enhance the model’s ability to generalize better.
 |
Figure 2: Correlation Heatmap of Features in the Diabetes Prediction Dataset (DPD)
|
The Pearson correlation coefficient (𝑟) was used to calculate the strength and direction of linear relationships between variables in the correlation map that was generated.
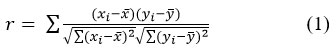
where xi is each value of variable x
yi is each value of variable y
are the means of 𝑥 and 𝑦
This approach involves calculating the covariance between variables and then normalizing it by their individual standard deviations (eq 1). Through this analysis, meaningful patterns and relationships emerged, providing insight into how different variables are correlated, whether positively, negatively, or not at all.
In DPD, features such as ‘heart_disease’, ‘hypertension’, ‘gender’, and ‘smoking_history’ were excluded due to their poor correlation with major predictive variables and distribution patterns. The correlation heatmap shown in Figure 2 indicates that these features have low correlation coefficients with major variables like blood glucose level and HbA1c level, which are major indicators of diabetes. In particular, heart_disease (0.07), hypertension (0.084), gender (0.017), and smoking_history (-0.035) have poor correlation with ‘blood_glucose_level’, indicating limited contribution to the predictive power of the model. Further, their association with HbA1c level is not high, lending additional support towards weak contribution by these variables for diabetes detection.
Similar analysis was followed for PIDD to drop weakly associated features like ‘SkinThickness’, ‘BloodPressure’, and ‘Pregnancies’. This process helps in better generalization, selection of impactful predictors and minimization of redundancy, thereby reducing overfitting.
Model Training
RF, TabNet, XGBoost, LightGBM, and CoNN-FSL models were trained on the datasets and evaluated for accuracy, precision, sensitivity, specificity, F1 scores and training time. The data was split into three sub-sets: 75% for training, 10% for testing, and 15% for validation.
CoNN35 is a prototype selection algorithm that removes redundant training samples while preserving the decision boundary. The goal is to extract a subset 𝑆 ⊂ 𝑋 that approximates the same classification function as the full dataset 𝑋. The algorithm:
Initially selects a random sample x0 from the dataset X and adds it to the prototype set S.
For each remaining sample xi ∈ X, subset generation is performed using 1-Nearest Neighbor (1-NN).
Let d(xi, S) be the Euclidean distance of xi from its nearest neighbour in S
![]()
If ýi ≠ yi (i.e, the predicted label differs from the actual label), then add xi to S.
Hence, the final prototype set S is much smaller than X.
Few-Shot Learning using Prototypical Networks36
The Condensed set is used in training the Prototypical Network to classify new instances by comparing them to class prototypes.
Given a support set Sc of examples from class c the prototype pc is the mean embedding of all examples in that class and is given by:
![]()
where fθ (x) – learned embedding function which is typically a neural network.
For a query sample xq, compute the Euclidean distance for each class prototype:
![]()
The predicted class label ȳq is assigned based on the nearest prototype:
![]()
The model is trained using negative Log-likelihood loss to minimize the negative log probability of the correct class:
![]()
where the probability is computed using a softmax over distances:
![]()
CoNN reduces dataset size while preserving decision boundaries, thus making training more efficient. Prototypical networks leverage these prototypes for FSL by classifying samples based on distance in an embedding space. This combination has proved to be faster and enhances detection, especially in data-scarce scenarios as presented in the Results section.
Results
In health care datasets, failing to observe a positive case (i.e., false negative) will cause serious and long-term patient issues. Therefore, considering recall (sensitivity/true positive rate) becomes essential, thereby decreasing the chance of diseases getting undiagnosed.
Table 1: Impact of Imputation Methods on Model Performance in PIDD
|
Imputation Methods |
Models |
Accuracy |
Precision |
Sensitivity |
Specificity |
F1-Score |
|
Zero Imputation |
RF |
75.32% |
66% |
72% |
77% |
69% |
|
|
XGB |
74.02% |
63% |
63% |
80% |
63% |
|
|
LGM |
72.72% |
62% |
59% |
80% |
60% |
|
|
TabNet |
74.03% |
68% |
48% |
88% |
57% |
|
Mean Imputation |
RF |
72.73% |
62% |
72% |
73% |
67% |
|
|
XGB |
74.02% |
63% |
63% |
80% |
63% |
|
|
LGM |
71.42% |
60% |
56% |
80% |
58% |
|
|
TabNet |
80.52% |
80% |
59% |
92% |
68% |
|
Median Imputation |
RF |
75.32% |
65% |
76% |
75% |
70% |
|
|
XGB |
74.02% |
63% |
63% |
80% |
63% |
|
|
LGM |
74.02% |
63% |
63% |
80% |
63% |
|
|
TabNet |
77.92% |
68% |
70% |
82% |
69% |
Table 1 shows the performance of: RF, XGBoost, LightGBM, and TabNet. Their performance was compared with Zero, Mean and Median imputation methods and the comparative analysis proves that median imputation works best among the three techniques, offering a balanced trade-off across several models and a closer fit for further study. Median imputation achieved greater or equal accuracy for three out of four models with higher sensitivity values reflecting improved detection. Mean imputation is prone to the effect of extreme values, but the median provides a better estimate of central tendency, which leads to better stable feature distributions. Conversely, TabNet performed best with mean imputation achieving the highest accuracy of 80.52% because of its dynamic feature selection architecture. The smoother distribution of mean imputation facilitates better learning of features resulting in improved performance. Regardless, mean imputation showed inconsistent results across other models, making it less suitable for generalization. Zero imputation introduces artificial bias by filling missing values with zeros, making it unfavorable for practical scenarios. Hence, the quantitative analysis validates the use of median imputation as compared to other techniques.
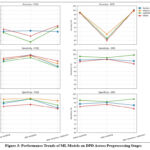 |
Figure 3: Performance Trends of ML Models on DPD Across Preprocessing Stages
|
Figure 3 shows comparative performance evaluation of different data preprocessing methods on the models for the two datasets considered. Training the models without preprocessing resulted in moderate accuracy, ranging from 78.45% to 80.51%, but with significantly lower sensitivity values. The models failed to perform towards increasing true positive rates. Handling the class imbalance notably improved the performance of all the models. They achieved impressive accuracy (93.19% – 97.59%) and higher sensitivity rates. However, RF and TabNet experienced slight variations in their accuracies after resampling, but their true positive rates increased significantly. After feature reduction, the outcome of the model performance was the same, with small variations in precision and accuracy. Although accuracy in a few instances slightly decreased, sensitivity was always high, which means that features removed were not essential for diabetic case detection. Shrinking the feature space also serves to combat overfitting and computational cost at a minimal loss of predictive capability. XGBoost and LightGBM, for which both datasets showed identical trends, outperformed other models as shown in Figure 3, rendering them the more reliable models for primary diabetes detection. Although feature reduction was not highly impacting detection accuracy, the excluded variables can still be valuable for possible future research, especially in risk factor analysis and consequent disease development research.
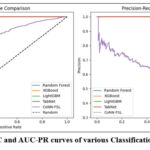 |
Figure 4: AUC-ROC and AUC-PR curves of various Classification models considered
|
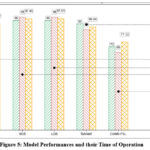 |
Figure 5: Model Performances and their Time of Operation
|
The proposed CoNN-FSL method addresses computational time and storage space constraints effectively. DPD was processed through CoNN which efficiently extracted a subset of 6554 samples from 100,000 samples while preserving the decision boundary. This subset reduced the time required for training the FSL model. The model achieved an accuracy of 77.12% demonstrating its ability to generalize better on smaller data and exhibited 73% sensitivity ensuring that it efficiently identified positive cases.
Table 2: Comparison of Dataset Usage, Memory Footprint, Training Time, and Accuracy Across Models
|
Model |
Dataset Size (Samples) |
Dataset Handling Method |
Memory Usage (kB) |
Training Time |
|
RF |
1,00,000 |
Resampling |
7500 |
488.752 |
|
XG Boost |
1,00,000 |
Resampling |
7500 |
214.471 |
|
Light GBM |
1,00,000 |
Resampling |
7500 |
308.545 |
|
TabNet |
1,00,000 |
Resampling |
7500 |
3318.082 |
|
CoNN-FSL |
1,00,000 |
Reduction |
492 |
83.666 |
CoNN-FSL demonstrated a significant improvement in memory utilization compared to other models. The original dataset, containing 100,000 samples, occupied around 7.5MB (7500 kB) of memory. However, when the sample size was reduced to 6,554 samples using CoNN, the memory footprint shrank to 492 kB approximately 1/15th of the original memory usage as shown in Table 2. The compressed memory profile not only highlights the model’s efficiency but also makes it highly suitable for lightweight deployment scenarios where resource constraints are critical.
Figure 4 shows that CoNN-FSL yields slightly lower ROC and Precision-Recall performance, due to a higher false positive rate and reduced precision at high recall, this is a strategic and expected trade-off. As shown in Figure 5, the proposed CoNN-FSL algorithm outpaces other models with a training time of 83.666 seconds, which is 39% less than the time required by XGBoost, a gradient boosting architecture. A careful analysis projects that, CoNN-FSL utilizes 3% of the training time required by deep learning frameworks like TabNet. The results are promising enough to handle the real-world applications of AI in the medical field.
Discussion
Diabetes prediction using ML has been widely researched, and ensemble models including RF, XGBoost, and LightGBM have exhibited excellent classification performance. A key factor in implementing ML models in clinical practice is ensuring their robustness across several datasets and real-world variability. Patient populations differ widely in demographics and data quality, making it essential for predictive models to generalize well beyond the controlled conditions of benchmark datasets. It is evident from Table 3 that earlier RF models have recorded accuracy ranging from 75.22% to 91%, and recall, 66% to 86.4%, suggesting inconsistency in the ability to effectively classify diabetic cases. Similarly, XGBoost-based models exhibited recall rates between 88% and 95.08%, suggesting a higher ability to identify positive cases. LightGBM models delivered recall scores ranging from 82% to 93.13%, further supporting their accuracy in diabetes classification. Nonetheless, the models in this research produced higher or comparable true positive scores, with better diabetic case identification. In terms of sensitivity, RF yielded 94%, XGBoost a staggering 100%, with all diabetic cases identified correctly, and LightGBM yielded 96%, surpassing previous benchmarks. Although TabNet is not much explored for diabetes prediction, this study analyzed its performance and achieved an accuracy of 96.95%, and a sensitivity of 65%. In spite of overall high accuracy, relatively lower sensitivity of TabNet suggests potential difficulties in identifying positive diabetic cases. It can be due to the model’s sensitivity to data imbalance or insufficient representation of minority classes during training, which could be overcome with more targeted sampling strategies or architecture optimization.
Table 3: Performance Metrics of various models for Diabetes Prediction
|
References |
Model |
Accuracy |
Precision |
Sensitivity |
F1-score |
| [25] |
Random Forest |
86.40% |
86.10% |
86.40% |
85.80% |
| [22] |
Random Forest |
76.81% |
77% |
79% |
78% |
| [39] |
Random Forest |
75.22% |
82.12% |
80.52% |
81.31% |
| [13] |
Random Forest |
75.4% |
75.4% |
75.1% |
75.2% |
| [37] |
Random Forest |
88% |
69% |
66% |
75% |
| [37] |
XGBoost |
90% |
89% |
88% |
86% |
| [37] |
LightGBM |
86% |
83% |
82% |
86% |
| [38] |
XGBoost |
94.18% |
95.51% |
95.08% |
95.30% |
| [38] |
LightGBM |
91.37% |
91.97% |
94.32% |
93.13% |
| [40] |
RandomForest |
91% |
89% |
79% |
84% |
| [40] |
XGBoost |
93% |
89% |
87% |
88% |
|
Experiment 1 |
Random Forest |
93.90% |
90% |
99% |
94% |
|
Experiment 2 |
XGBoost |
98.67% |
97% |
100% |
99% |
|
Experiment 3 |
Light GBM |
95.47% |
92% |
99% |
96% |
|
Experiment 4 |
TabNet |
96.95% |
99% |
65% |
78% |
The data imbalance is handled using SMOTE to enhance true positive rates, allowing the detection of diabetic cases at higher rates without affecting precision. The process of feature reduction also confirmed that the omission of some features did not affect sensitivity, further demonstrating the effectiveness of a less complex predictive model. However, synthetic sampling might not entirely capture actual patient complexity and over-pruning risks discarding clinically valuable features. Although SMOTE aids enhanced sensitivity, it can create synthetic examples that are clinically unbalanced, which might influence the interpretability and clinical acceptability of model predictions. This requires diligent validation prior to deployment.
In contrast to traditional DL models that need massive training data, the proposed CoNN-FSL framework systematically downscales the dataset requirement while maintaining the decision boundaries, which results in reduced time for training. CoNN-FSL exhibits a better memory utilization condensing the original set of 6554 samples maintaining a similar performance compared to the techniques in the literature as shown in Table 3. While the proposed CoNN-FSL framework shows promising results in terms of reducing computational costs and training time, its performance across heterogeneous clinical datasets is yet to be fully established beyond the ones used in this study. As the datasets may not fully capture the demographics and clinical variability in broader populations, sampling bias may be present. Additionally, although the model is optimized to reduce the training time, this optimization could increase the risk of overfitting, especially when learning from condensed or limited data samples. It is equally important to consider that certain observed model performances may be the result of dataset-specific properties, including class distribution, interaction among features, or hyperparameter advantage benefits. Thus, future research should validate these results across broader and diverse datasets to maintain reproducibility.
The outcomes indicate that while conventional ensemble methods are extremely powerful, CoNN-FSL presents a promising way to manage AI-driven medical diagnosis. This technique provides a novel approach for future studies addressing lightweight but effective DL models in healthcare solutions.
Conclusion
The study presents CoNN-FSL, an ML approach that offers faster and more resource-efficient diabetes prediction without compromising diagnostic accuracy. Unlike traditional models that require extensive data and computing power, the design effectively learns from smaller, representative datasets, making it highly suited for real-world clinical settings where time, data, and infrastructure may be limited. By reducing training time and memory demands, this method supports more scalable and adaptable screening solutions. These findings highlight the value of efficient data preprocessing and model optimization in developing AI tools for healthcare. While training time and computational costs have reduced, the current predictive performance of the model remains below the desired threshold, therefore a key focus in future work will be to enhance accuracy without compromising model’s lightweight efficiency. Future work will focus on refining this balance, exploring its integration into clinical workflows and electronic health record systems to enhance its practical utility in time-sensitive diagnostics.
Acknowledgement
The authors would like to thank PES University for granting the Internal Research Funding for carrying out this research work. We would also like to thank the Dept. of ECE, PES University for facilitating the necessary support.
Funding Source
This study was supported by PES University, Bengaluru. Grant number PESUIRF/ECE/2024/07
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to Reproduce Material from other Sources
Not Applicable.
Author Contributions
- Shruthi M L J: Conceptualization, Funding Acquisition, Project Administration, Resources, Supervision, Writing – Review & Editing.
- Nirmala Devi M: Conceptualization, Funding Acquisition, Project Administration, Resources, Supervision, Writing – Review & Editing.
- Nishita N Joshi: Data Collection, Analysis, Methodology, Model Training, Writing – Original Draft, Writing – Review & Editing.
- G Tanushree: Data Collection, Analysis, Methodology, Model Training, Writing – Original Draft, Writing – Review & Editing.
References
- Kharroubi AT. Diabetes mellitus: The epidemic of the century. World Journal of Diabetes. 2015;6(6):850. doi:10.4239/wjd.v6.i6.850
CrossRef - Chaudhary N, Tyagi N. Diabetes mellitus: an overview. Int J Res Dev Pharm Life Sci. 2018;7(4):3030-3033.
CrossRef - International Diabetes Federation. IDF Diabetes Atlas. 10th ed. Brussels, Belgium: International Diabetes Federation; 2021.
- American Diabetes Association. Diagnosis and classification of diabetes mellitus. Diabetes Care. 2021;44(1):1-20.
CrossRef - World Health Organization. Global Report on Diabetes. Geneva, Switzerland: World Health Organization; 2016.
- National Institute of Diabetes and Digestive and Kidney Diseases. Gestational diabetes: causes, risks, and prevention. National Institute of Diabetes and Digestive and Kidney Diseases. Published 2022.
- Maghfiroh AA, Simanjorang C, Karima UQ. Factors Associated with the Incidence of Prediabetes in Bogor, Indonesia: A Cohort Study. J Res Health Sci. 2025;25(1):e00635. doi:10.34172/jrhs.2025.170
CrossRef - American Diabetes Association. Standards of medical care in diabetes—2022. Diabetes Care. 2022;45(1):1-264.
CrossRef - Grundy SM. Obesity, metabolic syndrome, and cardiovascular disease. J Clin Endocrinol Metab. 2004;89(6):2595-2600. doi:10.1210/jc.2004-0372
CrossRef - Anjana RM, Pradeepa R, Deepa M, et al. The Indian Council of Medical Research-India Diabetes (ICMR-INDIAB) study: methodological details. J Diabetes Sci Technol. 2011;5(4):906-914. Published 2011 Jul 1. doi:10.1177/193229681100500413
CrossRef - Dagliati A, Marini S, Sacchi L, et al. Machine Learning Methods to Predict Diabetes Complications. J Diabetes Sci Technol. 2018;12(2):295-302. doi:10.1177/1932296817706375
CrossRef - American Diabetes Association. Glycemic goals and hypoglycemia: standards of care in diabetes—2025. Diabetes Care. 2025;48(Suppl 1):S128-S140.
CrossRef - Bhoi SK, Panda SK, Jena KK, et al. Prediction of diabetes in females of Pima Indian heritage: a complete supervised learning approach. Turk J Comput Math Educ. 2021;12(10):3074-3084.
- Jian Y, Pasquier M, Sagahyroon A, Aloul F. A Machine Learning Approach to Predicting Diabetes Complications. Healthcare (Basel). 2021;9(12):1712. Published 2021 Dec 9. doi:10.3390/healthcare9121712
CrossRef - Patil, R, Tamane, Sharvari. A Comparative Analysis on the Evaluation of Classification Algorithms in the Prediction of Diabetes. International Journal of Electrical and Computer Engineering. 2018;8:3966-3975. doi:10.11591/ijece.v8i5.pp3966-3975.
CrossRef - Guan Y, Tsai C, Zhang S. Research on diabetes prediction model of Pima Indian females. In: Proceedings of the 2024 4th International Symposium on Artificial Intelligence for Medicine Science (ISAIMS); 2024:294-303. doi:10.1145/3644116.3644168.
CrossRef - Alghamdi T. Prediction of diabetes complications using computational intelligence techniques. Appl Sci. 2023;13(5):3030.
CrossRef - J. A. Sabejon, J. B. Rejas, G. S. Lumacad, R. L. Zarate, E. A. D. Mendez and F. M. L. O. Tinoy. XGBoost–Based Analysis of the Early–Stage Diabetes Risk Dataset. 2023 International Conference in Advances in Power, Signal, and Information Technology (APSIT), Bhubaneswar, India. 2023, 19-24. doi: 10.1109/APSIT58554.2023.10201658
CrossRef - Doğru A, Buyrukoğlu S, Arı M. A hybrid super ensemble learning model for the early-stage prediction of diabetes risk. Med Biol Eng Comput. 2023;61(3):785-797. doi:10.1007/s11517-022-02749-z
CrossRef - Naz H, Ahuja S. Deep learning approach for diabetes prediction using PIMA Indian dataset. J Diabetes Metab Disord. 2020;19(1):391-403. Published 2020 Apr 14. doi:10.1007/s40200-020-00520-5
CrossRef - Sarwar, A., Ali, M., Manhas, J., & Sharma, V. Diagnosis of diabetes type-II using hybrid machine learning based ensemble model. International Journal of Information Technology. 2020;12, 419-428.
CrossRef - Reza MS, Amin R, Yasmin R, Kulsum W, Ruhi S. Improving diabetes disease patients classification using stacking ensemble method with PIMA and local healthcare data. Heliyon. 2024;10(2):e24536. Published 2024 Jan 19. doi:10.1016/j.heliyon.2024.e24536
CrossRef - K. Lakhwani, S. Bhargava, K. K. Hiran, M. M. Bundele and D. Somwanshi. Prediction of the Onset of Diabetes Using Artificial Neural Network and Pima Indians Diabetes Dataset. 2020 5th IEEE International Conference on Recent Advances and Innovations in Engineering (ICRAIE), Jaipur, India. Published 2021 Feb 26; 1-6, doi: 10.1109/ICRAIE51050.2020.9358308.
CrossRef - Rajni, Amandeep. RB-Bayes algorithm for the prediction of diabetic in Pima Indian dataset. International Journal of Electrical and Computer Engineering. 2019;9(6), 4866–4872.
CrossRef - Shams MY, Tarek Z, Elshewey AM. A novel RFE-GRU model for diabetes classification using PIMA Indian dataset. Sci Rep. 2025;15(1):982. Published 2025 Jan 6. doi:10.1038/s41598-024-82420-9
CrossRef - Qaraqe, M., Elzein, A., Belhaouari, S. et al. A novel few shot learning derived architecture for long-term HbA1c prediction. Sci Rep. 2024;14,482. Published 2024 Jan 04. doi.org/10.1038/s41598-023-50348-1
CrossRef - Sosale A, Prasanna Kumar KM, Sadikot SM, et al. Chronic complications in newly diagnosed patients with Type 2 diabetes mellitus in India. Indian J Endocrinol Metab. 2014;18(3):355-360. doi:10.4103/2230-8210.131184
CrossRef - Fregoso-Aparicio L, Noguez J, Montesinos L, García-García JA. Machine learning and deep learning predictive models for type 2 diabetes: a systematic review. Diabetol Metab Syndr. 2021;13(1):148. Published 2021 Dec 20. doi:10.1186/s13098-021-00767-9
CrossRef - Alfredo Daza et al. Clinical applications of artificial intelligence in diabetes management: A bibliometric analysis and comprehensive review, Informatics in Medicine Unlocked, Volume 50, 2024, 101567, ISSN 2352-9148. Published 2024 Jan 1. doi: 10.1016/j.imu.2024.101567.
CrossRef - PIDD: UCI Machine Learning Repository. Pima Indians Diabetes Database. Kaggle; 1988 Available from:https://www.kaggle.com/datasets/uciml/pima-indians-diabetes-database.
- DPD: iammustafatz. Diabetes Prediction Dataset. Kaggle; 2025 Available from: https://www.kaggle.com/datasets/iammustafatz/diabetes-prediction-dataset.
- Fernández, A., García, S., Herrera, F., Chawla, N. V. SMOTE for learning from imbalanced data: Progress and challenges, marking the 15-year anniversary. Journal of Artificial Intelligence Research. 2018;61, 863–905.
CrossRef - He, H., Garcia, E. A. Learning from imbalanced data. IEEE Transactions on Knowledge and Data Engineering. 2009; 21(9), 1263–1284.
CrossRef - Buda, M., Maki, A., Mazurowski, M. A. A systematic study of the class imbalance problem in convolutional neural networks. Neural Networks. 2018;106, 249–259.
CrossRef - P. Hart. The condensed nearest neighbor rule (Corresp.). IEEE Transactions on Information Theory, vol. 14, no. 3, pp. 515-516, May 1968. doi: 10.1109/TIT.1968.1054155.
CrossRef - Jake Snell, Kevin Swersky, and Richard Zemel. Prototypical networks for few-shot learning. 31st International Conference on Neural Information Processing Systems. 2017. Curran Associates Inc., Red Hook, NY, USA, 4080–4090.
- Ganie, S. M., Pramanik, P. K. D., Malik, M. B., Mallik, S., Qin, H. An ensemble learning approach for diabetes prediction using boosting techniques. Frontiers in Genetics. 2023;14, 1252159.
CrossRef - Li W, Peng Y, Peng K. Diabetes prediction model based on GA-XGBoost and stacking ensemble algorithm. PLoS One. 2024;19(9):e0311222. Published 2024 Sep 30. doi:10.1371/journal.pone.0311222
CrossRef - Chang V, Bailey J, Xu QA, Sun Z. Pima Indians diabetes mellitus classification based on machine learning (ML) algorithms. Neural Comput Appl. Published online March 24, 2022. doi:10.1007/s00521-022-07049-z
CrossRef - Mohan S, Gowrisankar Reddy D. Enhanced diabetes prediction using Random Forest and XGBoost machine learning classifiers with dual datasets. Int J Sci Res Sci Technol. 2023; 10(5): 434.







