Manuscript accepted on :18-12-2025
Published online on: 28-01-2026
Plagiarism Check: Yes
Reviewed by: Dr. Vikrant Dandekar
Second Review by: Dr. Arif Ansori
Final Approval by: Dr. Mariia Shanaida
Anak Agung Ayu Putri Permatasari1* , Putu Angga Wiradana1
, Putu Angga Wiradana1 , I Gede Widhiantara1
, I Gede Widhiantara1 , Ni Kadek Yunita Sari1
, Ni Kadek Yunita Sari1 , I Made Gde Sudyadnyana Sandhika1
, I Made Gde Sudyadnyana Sandhika1 , Mochammad Aqilah Herdiansyah2
, Mochammad Aqilah Herdiansyah2 , Win Darmanto2
, Win Darmanto2 and Novaria Sari Dewi Panjaitan3
and Novaria Sari Dewi Panjaitan3
1Department of Biology, Faculty of Health and Science, Universitas Dhyana Pura, Jalan Raya Padangluwih, Dalung, North Kuta, Badung Regency, Bali Province, Indonesia.
2Department of Biology, Faculty of Science and Technology, Universitas Airlangga, Surabaya, East Java, Indonesia
3Center for Biomedical Research, Research Organization for Health, National Research and Innovation Agency (BRIN), Bogor, West Java, Indonesia
Corresponding Author E-mail:putripermatasari@undhirabali.ac.id
DOI : https://dx.doi.org/10.13005/bpj/3360
Abstract
Hepatocellular carcinoma (HCC) is the most common type of primary liver cancer worldwide. Several signaling pathways are involved in the development and apoptosis of cancer cells. Brown macroalgae, Sargassum aquifolium, is a marine functional food source known to have anticancer activity by interfering with carcinogenesis and acting as a tumor suppressor. This study aims to screen phytoconstituents from ethanol extract of S. aquifolium and the anticancer effects of its compounds on five protein receptors involved in the development and apoptosis of HCC in silico approach. Phytochemical screening was performed by GC-MS and molecular docking was performed on selected compounds from S. aquifolium with growth receptors such as VEGFR2, EGFR; apoptotic proteins such as Bcl-2, Caspase-3, and Caspase-9. GC-MS analysis, by comparing spectra to the NIST database, identified nineteen bioactive compounds. Among these, only four exhibited potential as candidates for HCC therapy. This study showed that better docking scores were obtained by linoleic acid when compared to palmitic acid, dodecanoic acid, formic acid, and phytol. Linoleic acid demonstrates promising binding affinity for receptors associated with HCC, suggesting its utility as a therapeutic inhibitor. To fully realize this potential, however, future studies should focus on the isolation of this phytoconstituent and the evaluation of its bioactivity through in vivo models. Such research would yield more comprehensive insights and further illuminate its pharmacological prospects.
Keywords
ADME-Tox; GC-MS; Molecular docking;, Pharmacology; Phytochemical; Sargassum aquifolium
Download this article as:| Copy the following to cite this article: Permatasari A. A. P, Wiradana P. A, Widhiantara I. G, Sari N. K. Y, Sandhika I. M. G. S, Herdiansyah M. A, Darmanto W, Panjaitan N. S. D. Phytoconstituents and Molecular Docking of Ethanol Extract of Sargassum aquifolium J. Agardh with Different Apoptosis Proteins Involved in Hepatocellular Carcinoma. Biomed Pharmacol J 2026;19(1). |
| Copy the following to cite this URL: Permatasari A. A. P, Wiradana P. A, Widhiantara I. G, Sari N. K. Y, Sandhika I. M. G. S, Herdiansyah M. A, Darmanto W, Panjaitan N. S. D. Phytoconstituents and Molecular Docking of Ethanol Extract of Sargassum aquifolium J. Agardh with Different Apoptosis Proteins Involved in Hepatocellular Carcinoma. Biomed Pharmacol J 2026;19(1). |
Introduction
Hepatocellular carcinoma (HCC) is a primary liver cancer that develops from abnormal hepatocytes, and accounts for approximately 80% of liver cancer cases worldwide.1 HCC is the fifth most common cancer worldwide and the most common malignancy in adult males.2 Reports suggested that approximately 7.5 Lakh new cases of HCC are diagnosed globally each year.3 The development of HCC has been linked to hepatitis infection and exposure to aflatoxin produced by Aspergillus flavus.4 A case of liver cancer patients in Guangxi, China, showed high levels of aflatoxin B1 in serum and tumor tissue.5 HCC is most commonly found in patients with chronic hepatitis B and C.6 Treatments carried out to treat HCC are through surgery, transplantation, heating of cancer cells, and chemotherapy.7
Surgery is now the main treatment for HCC as it produces the best outcome of all treatments with a 5-year survival rate of 60-80%. Liver excision and transplantation are the initial options for individuals with early malignancies with curative intent.8 Besides the surgical excision, patients with HCC are treated with chemotherapy drugs such as fluorouracil,9 adriamycin,10 and cisplatin.11 Although targeted drugs can be given to treat HCC, some patients develop drug resistance towards those and have a poor prognosis after long-term therapy. Cellularly, several protein receptors involved in HCC development have been studied such as VEGFR-2 and EGFR, and their apoptotic proteins such as B-cell lymphoma-2 (Bcl-2), caspase-3, and caspase-9. The transmembrane receptor tyrosine kinase VEGFR-2/KDR can be overexpressed through vascular endothelial growth in advanced HCC. EGFR is one of the first cancer-causing genes discovered and a major target in cancer therapy including HCC.12,13 Likewise, apoptosis-related proteins such as Bcl-2 are involved in apoptosis, necrosis and autophagy that regulate cell death14 and caspases are proteins that regulate cell development, differentiation and apoptosis with caspase-3 acting as the final apoptosis executor.15
Certain compounds used in traditional medicine have the potential to to treat HCC with relatively lower cytotoxicity than the cytotoxicity due to clinical drugs treatment.16 For example, the astaxanthin compound is a member of xanthophylls, because it contains not only carbon and hydrogen atoms, but also oxygen atoms. As an anticancer, the astaxanthin compound has a mechanism to inhibit HCC growth factors such as VEGFR-2 and EGFR, as well as apoptosis proteins such as Bcl-2, caspase-3, and caspase-9.17 Sources of traditional medicines, such as the astaxanthin compound, can be found from marine sources, especially seaweed which is rich in bioactive compounds with potential anticancer abilities.18,19 Brown seaweed containing fucoidan polymers has been reported as a source of natural compounds 20 that could treat oral cancer.21 Fucoidan from brown macroalgae and carrageenan from red macroalgae was reported to exhibit anti-metastatic, apoptotic, and antiviral properties that can inhibit cancer growth.22,23 Previous studies reported sodium alginate produced from brown seaweed as a potential biopolymer in drug delivery,19 a source of antioxidants,24 antibacterial activities,25 and its combination with Blumea balsamifera leaves extract as an anti-hypercholesterolemia.26
However, until now the mechanism of action played by S. aquifolium compounds in inhibiting the development of HCC has not been reported. This study aims to analyze the phytoconstituents of the ethanol extract of S. aquifolium and its potential targets that can underlie the mechanism of action of its active compounds in the development of HCC. To obtain optimal drug efficacy, the accuracy of active compounds in interacting with receptors is very necessary through analysis of signaling pathways related to HCC by in silico molecular docking.17
Materials and methods
Seaweed collection and extraction
Brown algae, S. aquifolium, was collected from Sindhu Beach, Denpasar City in the intertidal zone at the lowest ebb and flow conditions and transported to the research laboratory, Universitas Dhyana Pura. Authentication of the brown algae samples was submitted to the botanist of the National Research and Innovation Agency (BRIN) and the specimens were preserved in the laboratory for future supporting references according to the method described in previous studies with specimen number: 25498.19,24 1.5 kg of wet weight of S. aquifolium was dried using an oven at a temperature of 50℃ for three days, coarsely ground using a grinder, filtered with a No. 40 mesh sieve. A total of 100 grams of S. aquifolium simplicia was extracted in 1,000 mL of ethanol solution (Sigma Aldrich Co. USA) by the maceration method. The extraction was carried out for three days and stirred every day to increase the absorption of the compound. After three days, the sample was filtered and the filtrate obtained was concentrated and dried using a rotary vacuum evaporator. The final extract yield was determined based on the dry weight of powdered S. aquifolium.27
GC-MS/MS of seaweed extract
The sample was filtered using a 0.45 μm micro membrane and the sample was injected into the GC-MS instrument as much as 1 μL. The optimization conditions of the GC-MS instrument specifications are the instrument used is the Agilent 6980N Network GC system with an autosampler with an Agilent 5973 inert MSD detector, J&W Scientific Column, HP-5MS, 0.25mm × 30m × 0.25 µm, Split inlet 1/100, 250, Over programmed 50 (5 minutes) 10 280 (15 minutes), column flow 1 mL/min (constant), aux of 250, MS Quad = 150 MS Source = 230 Scan mode = 20-600 amu Solvent delay = 0 minutes Library = Wiley version 7.0 Injection volume = 1 µL.
ADME-Tox study
Compounds identified in GC-MS were evaluated for their absorption, distribution, metabolism, and excretion using the SwissADME module, accessible on the SIB (Swiss Institute of Bioinformatics) website (https://www.sib.swiss) , as well as partition coefficients, solubility, and various other characteristics. Furthermore, Osiris DataWarrior (Version 5.2.1) was used to estimate the toxicity and drug-like properties of the compounds.28
Molecular docking analysis
Biological dataset
These computational experiments were conducted using a series of tools including Ligprep, Sitemap, Grid Generation, Grid XP Dock, MMGB-SA analysis, and other computational techniques.
Ligand optimization
A set of phytoconstituents was obtained from the GC-MS table of S. aquifolium extract. For docking purposes, compounds were tested for their drug-like properties based on Lipinski’s Rule of Five screening. The selected compounds with drug-like properties were visualized using ChemSketch (Freeware) in .sdf and mol formats.29 The 3D structures of selected compounds were retrieved in SDF format from the PubChem chemical database of NCBI. Next, using Pymol, these SDF format structures were converted to PDB format and prepared for docking by removing water molecules, forming hydrogen bonds, and adding required charges AutodockTools-1.5.7. These structures were saved in PDBQT format for docking.
Protein optimization
RCSB Protein Data Bank provides the structures of VEGFR-2 kinase (PDB ID. 3VHK), EGFR kinase (PDB ID. 5UGB), BCL-2 (PDB ID. 6O0K), Caspase-3 (PDB ID. 2XYG), and human caspase-9 (PDB ID. 3V3K). Caspase is a hydrolase receptor and Bcl-2 is an apoptosis regulatory receptor. Meanwhile, VEGFR-2 and EGFR act as HCC transferase receptors. All water molecules were removed from the protein structures using AutodockTools-1.5.7, and polar hydrogen atoms and charges were added. Bond angles, bond sequences, and topologies were fixed to optimize the structures. In detail, the list of receptor proteins used in this test is shown in Table 1. For docking, these structures were saved in PDBQT format.30
Table 1: List of HCC receptor proteins used in molecular docking trials
| Number | Receptors | PDB ID | Classification of Receptor | Mutation |
| 1. | Crystal structure of the VEGFR2 kinase domain in complex with a back pocket binder | 3VHK | HCC Transferase | No |
| 2. | Crystal structure of the EGFR kinase domain in complex with 4-(4-{[2-{[(3S)-1-acetylpyrrolidin-3-yl]amino}-9-(propan-2-yl)-9H-purin-6-yl]amino}phenyl)-1-methylpiperazin-1-ium | 5UGB | HCC Transferase | No |
| 3. | Crystal structure of BCL-2 with venetoclax | 6O0K | Apoptosis | No |
| 4. | Caspase-3:CAS329306 | 2XYG | Hydrolase | No |
| 5. | Human caspase 9 in complex with bacterial effector protein | 3V3K | Hydrolase | No |
Receptor grid generation
A three-dimensional grid box was created to allow the ligand to dock with the receptor. To create the grid box, a protein and a ligand in pdbqt format are selected. In addition to adjusting the dimensions of the grid box to match X, Y, and Z, a spatial adjustment of 0.5 A—almost a quarter of the length of a single-bonded carbon atom—was made. In the center of the grid box, the ligand was present and correctly positioned within the active site of the receptor protein.28
Molecular docking
Vina 1.2.5 docking software was used to convert each receptor and its native ligand into a .pdb file for redocking technique. The mean square deviation (RMSD) was calculated by redocking with the native ligand. To validate the grid box coordinates, we set a limit of RMSD <2 Å. As recommended in previous studies, AutoDock Vina 1.2.5 was used to perform three-dimensional interaction simulations between ligands and receptors selected in this study.31
Molecular dynamics simulations
The docking results were verified using molecular dynamics simulations using CABS-flex 2.0 (https://biocomp.chem.uw.edu.pl/CABSflex2/index) to ensure the stability of the molecular complex. The RMSD value, which serves as an indicator of stability, was used to select the ligand-protein combinations evaluated in the simulations. Protein rigidity, constraints, number of cycles, C-alpha weight and side chain constraints, temperature range, trajectory, and random number generator seed were among the characteristics of the molecular dynamics simulations.32
Results
Phytoconstituents of ethanol extract of S. aquifolium
GCMS screening showed the presence of abundant secondary metabolites in S. aquifolium that may provide some therapeutic effects. Gas chromatogram was performed for about 32 min and the fractions from GC were examined by mass spectrometry. The detected bioactive compounds served as ligands for docking analysis. S. aquifolium extract contained 19 bioactive compounds with some of the same compounds and were therefore excluded from the data (Table 2). GC-MS study of the ethanol extract of S. aquifolium revealed a total of 19 different compounds with diverse phytochemical activities. Table 2 lists the chemical components, including retention time (RT) and peak area (%).
Table 2: Results of GC-MS chromatogram analysis of the ethanol extract fraction of brown macroalgae S. aquifolium
| No. | Retention Time | Name of Compound | Peak Area % |
| 1 | 2.335 | Formic Acid | 12.11 |
| 2 | 3.130 | Hydrochloric acid | 12.85 |
| 3 | 3.478 | Acetic acid | 0.03 |
| 4 | 3.843 | Acetyl carbinol | 0.01 |
| 5 | 4.2.17 | Vinylformic acid | 0.02 |
| 6 | 11.383 | Glycinamide | 0.00 |
| 7 | 5.417 | Formamide | 0.02 |
| 8 | 6.429 | 2-Furancarboxaldehyde | 0.01 |
| 9 | 8.975 | Disiloxane | 0.01 |
| 10 | 9.673 | Phosphonic acid | 0.00 |
| 11 | 9.716 | Ethanedioic acid | 0.01 |
| 12 | 9.828 | Hexanoic acid | 0.01 |
| 13 | 9.969 | Ethanediimidic acid | 0.00 |
| 14 | 23.816 | Palmitic Acid | 0.95 |
| 15 | 19.034 | Dodecanoic Acid | 0.03 |
| 16 | 24.044 | Linoleic Acid | 0.07 |
| 17 | 24.170 | Oleic Acid | 0.03 |
| 18 | 24.889 | Stearic acid | 0.02 |
| 19 | 25.266 | Phytol | 0.09 |
Selection of drug-likeness properties
The results of phytochemical screening of S. aquifolium extract were then selected for their drug-like properties based on screening with the Lipinski Rule of Five. A total of three compounds were screened for their drug-like properties with a bioavailability value of 0.55, namely palmitic acid, dodecanoic acid, and linoleic acid are shown in Table 3. Lipinski’s Rule of Five was utilized in distinguishing between drug-like and non-drug-like molecules. This rule predicts a high probability of success or failure due to drug-likeness for molecules that obey 2 or more rules i.e., Molecular mass <500 Daltons, High Lipophilicity (expressed as LogP <5), <5 hydrogen bond donors,<10 hydrogen bond acceptors, and Molar refractivity should be between 40-130.
Table 3: Drug-likeness parameter measurement results based on Lipinski’s Rule of Five Screening
| Compound | Drug-likeness Parameters | ||||
| Mass | Hydrogen Bond Donor | Hydrogen Bond Acceptors | LogP (High Lipophilicity) | Molar Refractivity | |
| Palmitic acid | 256 | 1 | 2 | 5.55 | 77.94 |
| Dodecanoic acid | 200 | 1 | 2 | 3.99 | 59.47 |
| Linoleic acid | 280 | 1 | 2 | 5.88 | 86.99 |
| Formic acid | 312 | 5 | 6 | -0.05 | 77.14 |
| Phytol | 296 | 1 | 1 | 6.36 | 95.56 |
In silico analysis and validation
The results of molecular docking using PyRx software showed that VEGFR-2 (PDB ID: 3VHK) had the best binding affinity to compounds in S. aquifolium. Linoleic acid had the best docking results against all receptor proteins with binding values of VEGFR2 (-6.3 kcal/mol), EGFR (-5.5 kcal/mol), BCL-2 (6.0 kcal/mol), caspase-3 (-4.9 kcal/mol), and caspase-9 (-5.3 kcal/mol) (shown in Table 4).
Table 4: Molecular docking results
| Number | Compound types | Docking results with several tested receptors (kcal/mol) | ||||
| VEGFR2(PDB ID: 3VHK) | EGFR(PDB ID: 5UGB) | BCL-2(PDB ID: 6O0K) | Caspase-3(PDB ID: 2XYG) | Caspase-9 (PDB ID: 3V3K) | ||
| 1. | Palmitic acid | -5.5 | -4.9 | -5.0 | -4.0 | -4.3 |
| 2. | Dodecanoic acid | -6.3 | -4.8 | -4.5 | -4.2 | -3.8 |
| 3. | Linoleic acid | -6.3 | -5.5 | -6.0 | -4.9 | -5.3 |
| 4. | Formic acid | -3.0 | -2.9 | -2.9 | -2.8 | -2.8 |
| 5. | Phytol | -5.2 | -5.3 | -6.1 | -4.3 | -5.0 |
Linoleic acid docking on VEGFR-2 showed hydrogen interactions with six amino acids involved, namely Ile888(A), His1026(A), Leu1019(A), Val899(A), Val898(A), Val916(A) (Figure 1). Linoleic acid docking with EGFR showed hydrogen bonds with five amino acids involved, namely Leu718(A), Leu844(A), Val726(A), Ala743(A), Leu792(A). The bond of linoleic acid with Bcl-2 showed hydrogen interactions with five amino acids, namely Arg146(A), Phe104(A), Tyr108(A), Leu137(A), Val148(A). Linoleic acid with caspase-3 shows hydrogen interactions involving three amino acids, namely Lys82(A), Leu81(A) His277(B) and linoleic acid with caspase-9 shows hydrogen bonds involving one amino acid, namely Arg87(B) (Table 5).
Table 5: Diagram of protein-ligand complex interactions (linoleic acid with receptor proteins related to HCC).
| Interaction Complez | Type of Interaction | Amino Acids Involved in Interaction |
| (a) Linoleic acid-VEGFR2 | vdw | Ile892(A), Ile1044(A), Asp1046(A), Glu885(A), Phe1047(A), Leu1035(A), Val848(A) |
| HI | Ile888(A), His1026(A), Leu1019(A), Val899(A), Val898(A), Val916(A) | |
| PHI | Cys1045(A), Lys868(A) | |
| (b) Linoleic acid-EGFR | vdw | Thr854(A), Thr790(A), Asp855(A), Met766(A), Phe856(A), Leu788(A), Met793(A), Gly796(A), Cys797(A), Gly719(A) |
| HI | Leu718(A), Leu844(A), Val726(A), Ala743(A), Leu792(A) | |
| PHI | Glu762(A), Lys745(A) | |
| (c) Linoleic acid-BCL-2 | vdw | Gly145(A), Trp144(A), Phe198(A), Ala100(A), Glu152(A), Phe153(A) |
| HI | Arg146(A), Phe104(A), Tyr108(A), Leu137(A), Val148(A) | |
| PHI | Ala149(A) | |
| (d) Linoleic acid-caspase-3 | vdw | Pro42(A), Glu43(A), Met39(A), Ser112(A), Thr77(A), Asp40(A), Leu223(B), Ala227(B), Lys224(B), Asp228(B), Tyr276(B), Phe275(B) |
| HI | Lys82(A), Leu81(A) His277(B) | |
| PHI | Met44(A) | |
| (e) Linoleic acid-caspase-9 | vdw | Leu179(B), Glu89(B), Phe84(B), Val86(B), Arg99(B), Glu85(B), Val86(H), Arg99(H), Asn81(H), Glu149(H), Lys183(H), Glu85(H) |
| HI | Arg87(B) | |
| PHI | Asn81(B), Arg80(B), Lys183(B) |
Note: vdw= Van der Walls interaction; HI=hydrophobic interaction; PHI=polar hydrogen interaction.
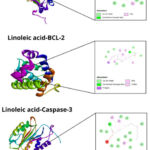 |
Figure 1: Chemical interaction of ligand-receptor complexes. |
Molecular dynamic simulations of the linoleic acid compound against the five protein receptors are shown in Table 6 and Figure 2. The highest root mean square deviation (RMSD) value was shown by the bond of linoleic acid – caspase-9 (2.175), followed by the linoleic acid – caspase-3 bond (2.157), linoleic acid – EGFR (1.971), linoleic acid – VEGFR-2 (1.940), and linoleic acid – Bcl-2 (1.709).
Table 6: Molecular dynamics simulation results
| Complex Name | Root Mean Square Deviation (RMSF)(Å) |
| Linoleic acid – VEGFR-2 | 1.940 |
| Linoleic acid– EGFR | 1.971 |
| Linoleic acid –Bcl-2 | 1.709 |
| Linoleic acid–caspase-3 | 2.157 |
| Linoleic acid–caspase-9 | 2.175 |
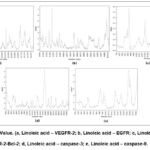 |
Figure 2: RMSF Value. (a, Linoleic acid – VEGFR-2; b, Linoleic acid – EGFR; c, Linoleic acid – VEGFR-2-Bcl-2; d, Linoleic acid – caspase-3; e, Linoleic acid – caspase-9. |
Discussion
Hepatocellular carcinoma (HCC) is a common cancer with a high mortality rate. Growing evidence suggests that gut dysbiosis and metabolic abnormalities contribute to the development of HCC. Drugs such as sorafenib, pembrolizumab, and nivolumab are currently used to treat HCC, but their long-term use may be limited due to side effects such as dermatotoxic responses, gastrointestinal reactions, systemic reactions, and vascular dysfunction.2,33 Several crystal structures of domains such as VEGFR-2, EGFR, BCL-2, caspase-3, and caspase-9 have been studied elsewhere previously.
Previous studies have explained that among 19 compounds of Ocimum basilicum extract, 8 compounds including palmitic acid were selected because they met Lipinski’s Rule of Five.34 Palmitic acid is a saturated fat containing 16 carbon atoms which is generally abundant in animal and vegetable fats.35,36 Dodecanoic acid or lauric acid is a medium-chain fatty acid molecule with twelve carbon atoms found in virgin coconut oil in concentrations ranging from 46 – 52%.37 Lauric acid is also found in foods such as human milk, fruits, palm oil, and including seaweed from the types Sargassum sp. and Ulva sp..38–40
Lauric acid has good antioxidant properties in cells, so it can protect cells and inhibit neuroinflammation, kill human colon cancer cells up to 93% by producing oxidative stress and apoptosis in cells, and can protect the liver from hepatotoxicity caused by ethanol induction by increasing HNF4α regulation and apoptosis.41–43 Linoleic acid can be converted through chain elongation and desaturation into long-chain fatty acids including arachidonic acid.44,45 Research showed that chitosan nanoparticles containing Melastomastrum capitatum leaf extract can deliver linoleic acid to OV7 cancer cells compared to MCF-7 cell lines. Linoleic acid loaded in chitosan also showed the highest selective index and increased caspase-3 activity.46
Molecular docking is a major theoretical tool used in molecular modeling to predict molecular binding, interaction intensity, and signal types in signal transduction and drug production processes. Molecular docking plays an important role in structural molecular biology and computer-aided drug design by predicting important binding models between ligands and known protein structures.47 Our in silico analysis showed that the binding affinity value to VEGFR-2 which is similar to dodecanoic acid is also shown by linoleic acid (-6.3 kcal/mol). In theory, angiogenesis involving vascular endothelial growth factor (VEGF) and its receptor VEGFR-2 is very important in the development of cancer and cardiovascular disease. VEGFR-2 is able to encourage vascular and regulate proliferation, migration, and survival of endothelial cells in vasculogenesis and angiogenesis.48 Targeting VEGFR-2 played by linoleic acid allows for a new therapeutic strategy. This tethering is intended to be able to suppress VEGFR-2 which is a tyrosine kinase receptor required in tumor cells.49 A previous study tested the effectiveness of astaxanthin extract in silico showed a high binding affinity value to the VEGFR-2 receptor (-6.46 kcal/mol when compared to positive drugs (Sorafenib) in general (-4.92 kcal/mol).17
Bcl-2 is a protein expressed by the bcl-2 gene that regulates cell death. Overexpression of bcl-2, especially when paired with certain genes such as myc, can trigger aggressive B-cell malignancies by suppressing apoptosis.50 Interestingly, we found that linoleic acid has a high binding affinity value to the Bcl-2 receptor. Caspase-3 and caspase-9, are required for cell apoptosis, but have the potential to increase cancer cell proliferation and invasion in certain cases.51 EGFR is a cell signaling molecule whose mutation or overexpression is associated with cancer, making it an important target for cancer treatment.52
In summary, this study demonstrates that the phytoconstituents from the ethanol extract of S. aquifolium are closely associated with the inhibition of protein receptors related to HCC. In silico analysis suggests that linoleic acid may act as a key regulator in the energy metabolism of HCC. These findings not only provide a novel perspective for understanding the mechanism of action of seaweed-derived bioactive compounds in regulating the metabolism of HCC development, but also offer a theoretical foundation for the development of novel functional foods aimed at specifically modulating HCC metabolic pathways and their regulatory networks. Subsequent in vitro and in vivo experiments are warranted to further explore the specific functions of these compounds.
Our results showed that the ethanol extract of S. aquifolium is a promising candidate for use in several anticancer therapeutic techniques, especially hepatocellular carcinoma (HCC). However, detailed in vitro and in vivo studies are still needed as an addition to increase the validity of its efficacy and safety profiles before considering the use of S. aquifolium in clinical applications. From the above study, five compounds were found to be potential candidates for HCC inhibitors with the highest value shown by linoleic acid as a potent VEGFR-2 inhibitor.
Conclusion
Molecular docking study between growth factors and apoptosis proteins from HCC with S. aquifolium compounds clearly demonstrated the binding and interaction of amino acids at the active site between ligands and receptors. Since linoleic acid was found to bind growth and apoptosis proteins with the least free energy compared to the other four compounds, linoleic acid might activate apoptosis proteins, which in turn could inhibit the growth of HCC through inhibition of VEGFR-2, EGFR, and Bcl-2 in HCC, thus acting as a potent anticancer agent. To confirm the anti-HCC potential of the drug candidates discovered through the limited computational analysis, further benchworks and clinical studies are needed. The further investigations will provide a more comprehensive understanding of the safety and efficacy of the compounds discovered in treating HCC. On the other hand, exploring the efficacy of these key molecules in HCC animal models may pave new avenues for future clinical trials and the development of novel treatments for degenerative diseases. These phytoconstituents are useful from S. aquifolium source for the development of new HCC drugs targeting five HCC-related ligands, as they can be further modified and improved for better HCC treatment.
Acknowledgement
The authors like to express their gratitude to thank the Research Group of Biological Health, Department of Biology, Faculty of Health and Science, Universitas Dhyana Pura, Department of Biology, Faculty of Science and Technology, Universitas Airlangga, and Research Center for Biomedical, Health Research Organization, National Research and Innovation Agency (BRIN) who have assisted in implementing this research.
Funding Source
This research was funded by the Institute for Research and Community Service (LPPM) Universitas Dhyana Pura through the Internal Grants of Universitas Dhyana Pura 2024 with Contract Number: Contract No. 62/UNDHIRA-LPPM/ST/VIII/2024.
Conflict of Interest
The author(s) do not have any conflict of interest
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials
Permission to reproduce material from other sources
Not Applicable
Author Contributions
- Anak Agung Ayu Putri Permatasari and Putu Angga Wiradana: Conceptualization, Methodology, Writing – Original Draft, Project Administration, Funding Acquisition.
- I Gede Widhiantara: Supervision, Writing – Review & Editing, Project Administration.
- Ni Kadek Yunita Sari and I Made Gde Sudyadnyana Sandhika: Data Collection and Analysis.
- Mochammad Aqilah Herdiansyah, Win Darmanto, and Novaria Sari Dewi Panjaitan: Data Collection, Analysis, Visualization, and Supervision.
References
- Chidambaranathan-Reghupaty S, Fisher PB, Sarkar D. Hepatocellular carcinoma (HCC): Epidemiology, etiology and molecular classification. In: ; 2021:1-61.
CrossRef - Yang JD, Hainaut P, Gores GJ, et al. A global view of hepatocellular carcinoma: trends, risk, prevention and management. Nat Rev Gastroenterol Hepatol. 2019;16(10):589-604.
CrossRef - Kim DY. Changing etiology and epidemiology of hepatocellular carcinoma: Asia and worldwide. J Liver Cancer. 2024;24(1):622-670.
CrossRef - Wiradana PA, Darmayasa IBG, Wiratmin NI. The Test of Saccharomyces sp. Potential Filtrate to Inhibit The Growth of Aspergillus flavus FNCC6109 Broiler Chicken Concentrate Feed Model. In: Proceedings of the 2nd International Conference Postgraduate School. SCITEPRESS – Science and Technology Publications; 2018:532-536.
CrossRef - Zhou R, Liu M, Liang X, et al. Clinical features of aflatoxin B1-exposed patients with liver cancer and the molecular mechanism of aflatoxin B1 on liver cancer cells. Environ Toxicol Pharmacol. 2019;71:103225.
CrossRef - El-Serag HB. Epidemiology of Viral Hepatitis and Hepatocellular Carcinoma. Gastroenterology. 2012;142(6):1264-1273.e1.
CrossRef - Daher S, Massarwa M, Benson AA, et al. Current and Future Treatment of Hepatocellular Carcinoma: An Updated Comprehensive Review. J Clin Transl Hepatol. 2018;6(1):1-10.
CrossRef - Marrero JA, Kulik LM, Sirlin CB, et al. Diagnosis, Staging, and Management of Hepatocellular Carcinoma: 2018 Practice Guidance by the American Association for the Study of Liver Diseases. Hepatology. 2018;68(2):723-750.
CrossRef - Shaaban S, Negm A, Ibrahim EE, et al. Chemotherapeutic agents for the treatment of hepatocellular carcinoma: efficacy and mode of action. Oncol Rev. Published online May 28, 2014.
CrossRef - Mudd TW, Guddati AK. Management of hepatotoxicity of chemotherapy and targeted agents. Am J Cancer Res. 2021;11(7):3461-3474.
- Hamaya S, Oura K, Morishita A, et al. Cisplatin in Liver Cancer Therapy. Int J Mol Sci. 2023;24(13):10858.
CrossRef - Liu X, Wang P, Zhang C, et al. Epidermal growth factor receptor (EGFR): A rising star in the era of precision medicine of lung cancer. Oncotarget. 2017;8(30):50209-50220.
CrossRef - Azorin P, Bonin F, Tariq Z, et al. Kindlin‐1 modulates the EGFR pathway and predicts sensitivity to EGFR inhibitors across cancer types. Clin Transl Med. 2022;12(4).
CrossRef - Ouyang L, Shi Z, Zhao S, et al. Programmed cell death pathways in cancer: a review of apoptosis, autophagy and programmed necrosis. Cell Prolif. 2012;45(6):487-498.
CrossRef - Elaut G, Vanhaecke T, Heyden Y Vander, et al. Spontaneous apoptosis, necrosis, energy status, glutathione levels and biotransformation capacities of isolated rat hepatocytes in suspension: Effect of the incubation medium. Biochem Pharmacol. 2005;69(12):1829-1838.
CrossRef - Widiastuti NK, Virginia NM, Fery YIM, et al. Cytotoxicity Evaluation of Erythrina lithosperma Miq. Leaf Extract against Vero Cell Lines: In Vitro Study. Res J Pharm Technol. 2023;16:153-158.
CrossRef - Suganya V, Anuradha V. In silico molecular docking of astaxanthin and sorafenib with different apoptotic proteins involved in hepatocellular carcinoma. Biocatal Agric Biotechnol. 2019;19:101076.
CrossRef - Widhiantara IG, Putri Permatasari AAA, Rosiana IW, et al. The role of biopolymers as candidates for promoting health agents: A review. J Appl Pharm Sci. 2022;13(1):42-055.
CrossRef - Permatasari AAAP, Rosiana IW, Wiradana PA, et al. Extraction and characterization of sodium alginate from three brown algae collected from Sanur Coastal Waters, Bali as biopolymer agent. Biodiversitas J Biol Divers. 2022;23(3):1655-1663.
CrossRef - Gutiérrez-Rodríguez AG, Juárez-Portilla C, Olivares-Bañuelos T, et al. Anticancer activity of seaweeds. Drug Discov Today. 2018;23(2):434-447.
CrossRef - Madasamy S, Ganapathy D, Easwaran M, et al. Fucoidan a sulphated polysaccharide of brown seaweeds: a potential anticancer agent to treat oral cancer. Nat Prod Res. Published online February 10, 2024:1-3.
- Shannon E, Conlon M, Hayes M. Seaweed Components as Potential Modulators of the Gut Microbiota. Mar Drugs. 2021;19(7):358.
CrossRef - Cotas J, Leandro A, Monteiro P, et al. Seaweed Phenolics: From Extraction to Applications. Mar Drugs. 2020;18(8):384.
CrossRef - Permatasari AAAP, Wiradana PA, Sari NKY, et al. Antioxidant capacity, cytotoxicity, and bacterial contamination of brown macroalgae simplicia. J Pengolah Has Perikan Indones. 2024;27(10).
CrossRef - Waruwu E, Permatasari AAAP, Wiradana PA, et al. In Vitro Evaluation of Brown Algae Extract from Sargassum aquifolium in Inhibiting Aeromonas hydrophila, the Causative Agent of Motile Aeromonas Septicemia (MAS). J Aquac Sci. 2024;9(2):117-127.
CrossRef - Widhiantara IG, Wiradana PA, Permatasari AAAP, et al. Blumea balsamifera and Sargassum aquifolium extracts reduce fatty liver damage through lipid metabolism signalling pathways. Indones J Med Lab Sci Technol. 2024;6(2):131-142.
CrossRef - Jubilee R, Komala M. Phytochemical and isolated compound speciocide from Kigelia africana fruit. Int J Health Sci (Qassim). Published online September 27, 2022:3944-3956.
CrossRef - Pakyntein CL, Thabah D, Rai AK, et al. Identification of phytoconstituents from Houttuynia cordata Thunb. as dipeptidyl peptidase-IV and sodium glucose cotransporter 2 inhibitors guided by molecular docking. Phytomedicine Plus. 2024;4(3):100590.
CrossRef - Hadni H, Mazigh M, Charif E, Bouayad A, Elhallaoui M. Molecular Modeling of Antimalarial Agents by 3D-QSAR Study and Molecular Docking of Two Hybrids 4-Aminoquinoline-1,3,5-triazine and 4-Aminoquinoline-oxalamide Derivatives with the Receptor Protein in Its Both Wild and Mutant Types. Biochem Res Int. 2018;2018:1-15.
CrossRef - Manogar P, Vijaya Prabhu S, Durairaj P, John Abel MM, Prakash N, Jayanthi S. Molecular docking interaction of bioactive molecules from Kigelia africana (lam.) benth., revealed potential inhibitors of penicillin-binding protein 2 (PBP2). Asp Mol Med. 2024;4:100051.
CrossRef - Eberhardt J, Santos-Martins D, Tillack AF, Forli S. AutoDock Vina 1.2.0: New Docking Methods, Expanded Force Field, and Python Bindings. J Chem Inf Model. 2021;61(8):3891-3898.
CrossRef - Anandan S, Gowtham HG, Shivakumara CS, et al. Integrated approach for studying bioactive compounds from Cladosporium spp. against estrogen receptor alpha as breast cancer drug target. Sci Rep. 2022;12(1):22446.
CrossRef - Schutz FAB, Je Y, Richards CJ, et al. Meta-Analysis of Randomized Controlled Trials for the Incidence and Risk of Treatment-Related Mortality in Patients With Cancer Treated With Vascular Endothelial Growth Factor Tyrosine Kinase Inhibitors. J Clin Oncol. 2012;30(8):871-877.
CrossRef - Duraisamy S, Backiam ADS, Raju A, et al. In silico and in vitro analysis of bioactive compounds extracted from Ocimum basilicum against vancomycin-resistant enterococci. Chem Phys Impact. 2024;8:100499.
CrossRef - Mohanty B, Pal K, Quereshi D, et al. Oleogels Based on Palmitic Acid and Safflower Oil: Novel Formulations for Ocular Drug Delivery of Voriconazole. Eur J Lipid Sci Technol. 2020;122(4).
CrossRef - Shepardson RP, Harvatine KJ. Effects of fat supplements containing different levels of palmitic and stearic acid on milk production and fatty acid digestibility in lactating dairy cows. J Dairy Sci. 2021;104(7):7682-7695.
CrossRef - Ghani NAA, Channip AA, Chok Hwee Hwa P, et al. Physicochemical properties, antioxidant capacities, and metal contents of virgin coconut oil produced by wet and dry processes. Food Sci Nutr. 2018;6(5):1298-1306.
CrossRef - Wallace TC. Health Effects of Coconut Oil—A Narrative Review of Current Evidence. J Am Coll Nutr. 2019;38(2):97-107.
CrossRef - Khan HU, Aamir K, Jusuf PR, et al. Lauric acid ameliorates lipopolysaccharide (LPS)-induced liver inflammation by mediating TLR4/MyD88 pathway in Sprague Dawley (SD) rats. Life Sci. 2021;265:118750.
CrossRef - Chin YY, Chang KA, Ng WM, et al. A comparative evaluation of nutritional composition and antioxidant properties of six Malaysian edible seaweeds. Food Chem Adv. 2023;3:100426.
CrossRef - Ramya V, Shyam KP, Kowsalya E, Balavigneswaran CK, Kadalmani B. Dual Roles of Coconut Oil and Its Major Component Lauric Acid on Redox Nexus: Focus on Cytoprotection and Cancer Cell Death. Front Neurosci. 2022;16.
CrossRef - Fauser JK, Matthews GM, Cummins AG, et al. Induction of Apoptosis by the Medium-Chain Length Fatty Acid Lauric Acid in Colon Cancer Cells due to Induction of Oxidative Stress. Chemotherapy. 2013;59(3):214-224.
CrossRef - Namachivayam A, Valsala Gopalakrishnan A. Effect of Lauric acid against ethanol-induced hepatotoxicity by modulating oxidative stress/apoptosis signalling and HNF4α in Wistar albino rats. Heliyon. 2023;9(11):e21267.
CrossRef - Alagawany M, Elnesr SS, Farag MR, et al. Omega-3 and Omega-6 Fatty Acids in Poultry Nutrition: Effect on Production Performance and Health. Animals. 2019;9(8):573.
CrossRef - Mori TA, Hodgson JM. Fatty acids: Health Effects of Omega-6 Polyunsaturated Fatty Acids. In: Encyclopedia of Human Nutrition. Elsevier; 2013:209-214.
CrossRef - Ukwubile CA, Ahmed A, Katsayal UA, Ya’u J, Nettey H. Chitosan nanoparticle-mediated drug delivery for linoleic acid isolated from Melastomastrum capitatum Fern. leaf extract against MCF-7 and OV7 cancer cells. Pharmacol Res – Nat Prod. 2024;5:100105.
CrossRef - Fan J, Fu A, Zhang L. Progress in molecular docking. Quant Biol. 2019;7(2):83-89.
CrossRef - Wang X, Bove AM, Simone G, Ma B. Molecular Bases of VEGFR-2-Mediated Physiological Function and Pathological Role. Front Cell Dev Biol. 2020;8.
CrossRef - Farghaly TA, Al-Hasani WA, Abdulwahab HG. An updated patent review of VEGFR-2 inhibitors (2017-present). Expert Opin Ther Pat. 2021;31(11):989-1007.
CrossRef - Timucin AC, Basaga H, Kutuk O. Selective targeting of antiapoptotic BCL‐2 proteins in cancer. Med Res Rev. 2019;39(1):146-175.
CrossRef - Jiang M, Qi L, Li L, Li Y. The caspase-3/GSDME signal pathway as a switch between apoptosis and pyroptosis in cancer. Cell Death Discov. 2020;6(1):112.
CrossRef - Uribe ML, Marrocco I, Yarden Y. EGFR in Cancer: Signaling Mechanisms, Drugs, and Acquired Resistance. Cancers (Basel). 2021;13(11):2748.
CrossRef







