Manuscript accepted on :2-02-2026
Published online on: 05-03-2026
Plagiarism Check: Yes
Reviewed by: Dr. Niharika Kondepudi
Second Review by: Dr. Tagreed Altaei
Final Approval by: Dr. Prabhishek Singh
Lazizbek Makhmudov1* , Nigora Tagayalieva1
, Nigora Tagayalieva1 , Izzatullo Abdullaev1
, Izzatullo Abdullaev1 , Ulugbek Gayibov1
, Ulugbek Gayibov1 , Kuzijon Baratov1
, Kuzijon Baratov1 , Sherzod Usmonov1
, Sherzod Usmonov1 , Gulnora Rakhmonova1
, Gulnora Rakhmonova1 , Rano Yaqubova1
, Rano Yaqubova1 , Natalia Vypova1
, Natalia Vypova1 and Shaira Karimova2
and Shaira Karimova2
1Department of Plant Cytoprotectors and Pharmacology, A. S. Sadykov Institute of Bioorganic Chemistry of the Science Academy of Uzbekistan, Tashkent, Uzbekistan.
2Department of Pediatrics and folk medicine, Tashkent State Medical Univiersity, Tashkent, Uzbekistan.
Corresponding Author E-mail:l.maxmudov1990@gmail.com
DOI : https://dx.doi.org/10.13005/bpj/3361
Abstract
Inflammation remains a key pathological process underlying a wide range of chronic diseases, and natural compounds continue to attract attention as safer therapeutic alternatives. In this study, a novel composition (C-4), containing Cysteine, L-Carnitine, and Quercetin, was evaluated through a combination of in vivo, in silico, and pharmacokinetic approaches. The carrageenan-induced paw edema model in rats was employed to assess acute anti-inflammatory activity. C-4 administration (10 mg/kg, oral) significantly reduced paw swelling compared to the control and was comparable to the reference drug Loxidol (7 mg/kg), with the most pronounced effect observed after 3 hours. To further explain these effects, SwissADME pharmacokinetic predictions were performed. Both Cysteine and L-Carnitine exhibited high solubility but poor gastrointestinal absorption and no blood–brain barrier permeability, suggesting limited systemic bioavailability. Neither compound inhibited major cytochrome P450 isoforms nor acted as P-glycoprotein substrates, indicating a favorable safety profile but reduced distribution potential. Molecular docking studies against key pro-inflammatory proteins (COX-2, TNF-α, NF-κB, IL-6, MAPK) revealed that Quercetin possessed the strongest binding affinities, consistent with its marked in vivo anti-exudative activity. L-Carnitine showed moderate interactions, while Cysteine demonstrated weak direct binding, supporting its indirect role in redox regulation rather than direct enzymatic inhibition. The integration of experimental, pharmacokinetic, and computational data highlights Quercetin as the primary contributor to the anti-inflammatory activity of the C-4 composition, with L-Carnitine and Cysteine playing complementary cytoprotective and antioxidant roles. These findings suggest that combined formulations of natural compounds may provide synergistic benefits and represent promising candidates for the development of safe, multi-targeted anti-inflammatory therapies.
Keywords
Carrageenan Model; Cysteine; Inflammation; L-Carnitine; Molecular docking; Pharmacokinetics, Quercetin
Download this article as:| Copy the following to cite this article: Makhmudov L, Tagayalieva N, Abdullaev I, Gayibov U, Baratov K, Usmonov S, Rakhmonova G, Yaqubova R, Vypova N, Karimova S. In silico Pharmacokinetics and In vivo Anti-inflammatory Activity of C-4 Composition. Biomed Pharmacol J 2026;19(1). |
| Copy the following to cite this URL: Makhmudov L, Tagayalieva N, Abdullaev I, Gayibov U, Baratov K, Usmonov S, Rakhmonova G, Yaqubova R, Vypova N, Karimova S. In silico Pharmacokinetics and In vivo Anti-inflammatory Activity of C-4 Composition. Biomed Pharmacol J 2026;19(1). Available from: https://bit.ly/4bukFWX |
Introduction
Inflammation is a fundamental biological response of the immune system against infection, tissue injury, or chemical stimuli; however, excessive or chronic inflammation is closely associated with the progression of various pathological conditions, including cardiovascular diseases, metabolic disorders, and cancer.1 Therefore, the discovery and development of novel anti-inflammatory agents remain an important area of biomedical research. Recent advances in computational pharmacology and molecular docking provide valuable tools for predicting the pharmacokinetic properties and potential molecular targets of bioactive compounds. In parallel, in vivo experimental models, such as carrageenan-induced inflammation, continue to serve as reliable approaches to validate anti-inflammatory activity under laboratory conditions. In this study, we investigated the anti-inflammatory potential of a novel composition, C-4, which combines cysteine, carnitine, and quercetin. Each of these components is known for its distinct biological role: cysteine as a precursor of glutathione with antioxidant properties, carnitine as a regulator of cellular energy metabolism, and quercetin as a flavonoid with well-established antioxidant and anti-inflammatory activities.2 The synergistic combination of these molecules may enhance therapeutic efficacy compared to their individual effects. The C-4 composition was subjected to molecular docking analysis against key inflammation-related proteins to predict its binding affinity and inhibitory potential. In addition, its pharmacokinetic properties were evaluated using the SwissADME platform to assess drug-likeness and bioavailability. Furthermore, the in vivo anti-inflammatory activity of C-4 was validated using the carrageenan-induced inflammation model. Collectively, this study provides both in silico and in vivo insights into the pharmacological potential of the C-4 composition, supporting its possible role as a candidate for anti-inflammatory therapy.3
Materials and Methods
Animal ethics
All preoperative and experimental procedures were carefully reviewed and approved by the Institutional Committee for Animal Use and Care. The animals were housed in a vivarium under standardized conditions, including a relative humidity of 55%–65%, a controlled ambient temperature of 22 ± 2°C, and unrestricted access to water and standard laboratory chow. All aspects of animal care and handling were conducted in full compliance with the European Directive 2010/63/EU, which governs the protection of animals used for scientific research. Ethical clearance for this study was granted by the Animal Ethics Committee of the Institute of Bioorganic Chemistry, Academy of Sciences of the Republic of Uzbekistan (Protocol No. 133/1a/h, dated 4 August 2016).
Evaluation of Anti-Inflammatory Activity Using the Carrageenan-Induced Paw Edema Model
The preliminary evaluation of anti-inflammatory activity was performed using the classical carrageenan-induced paw edema model. The study was conducted on 15 rats (190 ± 20 g, same sex), which were divided into three groups: a control group and two experimental groups, with five animals in each group. Inflammation was induced by subplantar injection of 0.1 ml of 1% carrageenan solution into the right hind paw of the rats.4,5 One hour after carrageenan administration, animals in the experimental groups received the C-4 composition at a dose of 10 mg/kg orally in powder form.6 The control group received distilled water, while the reference drug Loxidol was administered orally at a dose of 7 mg/kg. The inflammatory response was assessed by measuring paw volume at 1, 2, 3, 4, and 5 hours after carrageenan injection. Based on the results obtained at the third hour, the anti-exudative activity was determined.
Molecular Docking “Software and databases”
Molecular Docking “Software and databases” All computational tools applied in this study were freely accessible for academic and educational purposes. Structural data of macromolecules involved in calcium signaling and regulation were retrieved from the Protein Data Bank (PDB), an internationally recognized repository of three-dimensional biomolecular structures.7,8 Reference compounds along with the flavonoid ligands of interest were obtained from the PubChem database, which provides extensive information on pharmacology, molecular targets, chemical structures, and biological pathways. Each PubChem DrugCard contains over 80 fields describing small molecules and their associated protein targets. Molecular structures and docking results were visualized with PyMOL (version 1.2), a Python-based molecular graphics system [http://www.pymol.org]. Docking simulations were carried out using AutoDock 4.2, developed at The Scripps Research Institute (www.scripps.edu). Input preparation and parameter configuration were performed with AutoDock Tools (ADT), a user-friendly graphical interface designed to set up and run docking experiments. AutoDock provides a robust computational framework for predicting ligand–target binding conformations and interactions when three-dimensional structural data are available.
Calculation of Inhibition Constant (Ki) from Binding Energy.
In molecular docking studies, the interaction strength between a ligand and its target protein is commonly expressed as the binding free energy (ΔG), measured in kilocalories per mole (kcal/mol).9 This value can be further applied to estimate the inhibition constant (Ki), which reflects the ligand’s binding affinity, using a simple thermodynamic relationship.
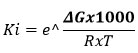
Where:
Ki is the inhibition constant (in mol/L)
ΔG is the binding free energy (in kcal/mol)
R is the universal gas constant = 1.987 cal/(mol·K)
T is the temperature in Kelvin (usually 298.15 K)
The factor 1000 converts kcal to cal.
Synthesis of the Complex
A total of 4 mmol (3.36 g) of GAMS was completely dissolved in 100 mL of 50% ethanol. While stirring intensively, the remaining components — quercetin, carnitine, and cysteine — in a total amount of 1.5 mmol were added gradually to the solution. The reaction mixture was continuously stirred at room temperature (≈25 °C) for 5–6 hours. After completion of the reaction, ethanol was removed under reduced pressure using a rotary evaporator, and the remaining aqueous fraction was lyophilized (freeze-dried). Yield: 3.15 g (93%). Decomposition temperature: 180 ± 1 °C. UV spectrum (λmax, nm (log ε)): 258 (5.38), 366 (5.08). IR spectrum (ν, δ, cm⁻¹): ν(OH) = 3209; ν(CH, CH₂, CH₃) = 2926, 2862; ν(C=O) = 1716; ν(C=O, C=C) = 1651; ν(COO⁻) = 1593; δ(CH₂, CH₃) = 1454; δ(CH) = 1387, 1365, 1261, 1211, 1165; δ(C–O–C, C–OH) = 1032; δ(=CH) = 980, 920, 879.
Statistical Analysis
All values were expressed as mean ± standard error mean (SEM) calculated using the Origin 9.0 (OriginLab, USA). Differences between groups were assessed using the t-test (Two Populations) system. Differences were regarded as significant if p< 0.05 was adopted.
Results
Anti-Exudative Activity of the C-4 Composition in the Carrageenan-Induced Paw Edema Model
The anti-inflammatory potential of the C-4 composition was assessed using the classical carrageenan-induced paw edema model in rats. In the control group, paw swelling developed progressively after carrageenan injection, reaching its maximum at the 3rd hour (63.3 ± 6.2%) compared to baseline values (Figure 1). Despite a slight decline during the following hours, the edema remained significantly elevated (47.2 ± 4.7%) at the 5th hour, confirming the persistence of inflammatory response in untreated animals.10-11
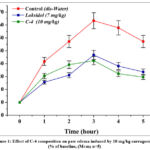 |
Figure 1: Effect of C-4 composition on paw edema induced by 10 mg/kg carrageenan (% of baseline, (M±m; n=5). |
In contrast, both Loxidol (7 mg/kg) and the C-4 composition (10 mg/kg, powder form) demonstrated substantial suppression of paw edema formation. At the peak inflammatory period (3 hours), paw swelling was significantly reduced to 36.7 ± 3.56% and 32.3 ± 3.2% in the Loxidol and C-4 groups, respectively. The calculated anti-exudative activity at this time point revealed that C-4 inhibited edema by 49.0%, whereas Loxidol achieved a 42.0% reduction relative to the control.12,13
A comparative analysis of the edema dynamics further supports the superior effect of the C-4 composition. At the 1st and 2nd hours, all groups showed measurable increases in paw volume; however, in the treated groups, the progression of swelling was significantly attenuated compared with the control. The difference became most evident at the 3rd hour, where the peak edema was effectively controlled by both treatments. Importantly, by the 4th and 5th hours, animals receiving C-4 showed a pronounced decline in paw swelling, with values approaching near-normal levels, while in the control group edema remained persistent. The recovery trend in the Loxidol group was present but less prominent compared to the C-4 group.14,15
Taken together, the results indicate that the C-4 composition exhibits potent anti-inflammatory and anti-exudative properties. Its inhibitory effect was not only stronger at the peak inflammatory stage but also sustained throughout the later observation period, leading to faster resolution of inflammation compared with both the untreated control and the reference drug. These findings suggest that C-4 may represent a promising candidate for further pharmacological evaluation as an anti-inflammatory agent.16,17
Molecular Docking studies with Inflammatory targets
Inflammation is a complex biological event that is important for the host’s protection against pathogens and injury. However, with persistent or inappropriate inflammation, inflammation contributes to the pathogenesis of numerous chronic diseases, including cardiovascular dysfunction, neurodegenerative diseases, diabetes, and cancer. At a molecular level, inflammation is regulated by numerous mediators. Cyclooxygenase-2 (COX-2) is an enzyme that is an important catalyst in the synthesis of pro-inflammatory prostaglandins, where its role is crucial to pain, fever, and tissue injury. Interleukin-6 (IL-6) is a multifunctional cytokine with roles in both acute and chronic inflammation, as well as cardiovascular and metabolic complications. The mitogen-activated protein kinase (MAPK) signaling cascade is responsible for modulating cellular responses to stimuli and integrating responses to stress and inflammation. Nuclear factor kappa system B (NF-κB) acts as the primary transcriptional regulator that effects changes in protein expression associated with immunity and inflammation. Tumor necrosis factor-alpha (TNF-α) functions as a potent pro-inflammatory cytokine that promotes systemic inflammation and suggestive of a wide array of pathological condition. Due to the pivotal role of these molecular targets in the development of inflammation, there is much enthusiasm regarding the identification of compounds that can influence their function. In particular, natural bioactive molecules have considerable appeal, since they offer safety, availability and the potential to elicit pleiotropic effects. Examples of bioactive compounds that have begun to attract interest include carnitine, cysteine, and quercetin. Carnitine is the focus of attention due to its important role in the transport of fatty acids into mitochondria and metabolism. Carnitine also has cytoprotective and anti-inflammatory activity. Cysteine, a precursor to the antioxidant glutathione, has a key role in maintaining redox state while providing cellular protection from oxidative stress. Quercetin, a flavonoid found in many fruits and vegetables, has recently gained attention for its strong antioxidant, anti-inflammatory, and cardioprotective properties.18,19
COX2
In our in silico experiment, the interaction between the COX-2 enzyme and the carnitine molecule resulted in a binding energy of –5.3 kcal/mol. According to thermodynamic principles, the more negative the binding energy, the more stable the interaction. Thus, the interaction between carnitine and COX-2 indicates a moderately stable binding affinity. Based on this energy value, the calculated inhibition constant (Ki) was approximately 100 μM, suggesting that carnitine may act as a moderate inhibitor of COX-2. COX-2 is an inducible enzyme whose overactivation leads to enhanced inflammatory processes, pain, and edema. Prostaglandins produced via COX-2 not only promote inflammation but also stimulate tumor cell proliferation, angiogenesis, and metastasis. Carnitine, on the other hand, is a naturally occurring molecule essential for energy metabolism, particularly in shuttling fatty acids into the mitochondria, and is also known for its antioxidant and cytoprotective properties. Therefore, the ability of carnitine to bind to and inhibit COX-2 suggests a potential dual role: exerting anti-inflammatory effects and possibly slowing cancer progression. Our results demonstrate that carnitine establishes a stable interaction with COX-2 and inhibits it at a moderate level, supporting the notion that carnitine could be further explored as a natural molecule with therapeutic potential against inflammation and tumor development.20,21
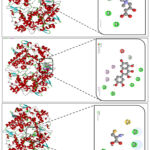 |
Figure 2: Molecular effects of Carnitine (A), Quercetin (B), and Cysteine (C) on the COX2 enzyme |
Molecular docking results demonstrated that carnitine interacts with several key amino acid residues located in the active site of the COX-2 enzyme (Figure 2). Specifically, carnitine formed conventional hydrogen bonds with ASN A:43 and ARG A:44, which help stabilize the ligand within the catalytic pocket. An electrostatic (attractive) interaction was also observed with GLU A:465, where the negatively charged carboxyl group of glutamate engaged with the corresponding moiety of carnitine, further strengthening the overall stability of the complex (Figure 4). In addition, carbon–hydrogen bonds were identified with GLY A:45 and CYS A:41, providing favorable positioning of the ligand and contributing additional stabilization within the binding site. The involvement of these residues highlights their structural and functional significance in COX-2 activity. Polar residues such as ASN and ARG stabilize the ligand orientation via hydrogen bonds, while negatively charged GLU enhances binding through electrostatic attraction. Flexible residues such as GLY and CYS facilitate proper accommodation of the ligand within the active site and contribute to hydrophobic stabilization. Together, these interactions suggest that carnitine forms a moderately stable complex with COX-2 and possesses a modest inhibitory potential. Furthermore, the interaction of COX-2 with the flavonoid quercetin was also investigated. Docking analysis revealed a binding energy of –9.4 kcal/mol (Figure 1B), which corresponds to an estimated inhibition constant (Ki) of approximately 0.13 μM, indicating a highly potent inhibitory effect. Molecular interaction analysis showed that quercetin formed conventional hydrogen bonds with SER B:49, ASN B:34, CYS B:36, GLN B:461, and PRO B:154. In addition, π–alkyl interactions were observed with MET B:48, PRO B:153, PRO B:156, and CYS B:36, while π–donor hydrogen bonding was identified with GLN A:327. An unfavorable acceptor–acceptor interaction was also detected with SER A:548 (Figure 1B). The combination of strong hydrogen bonding, π–alkyl, and π–donor interactions demonstrates that quercetin forms a highly stable complex within the COX-2 active site. These results confirm that quercetin, as a natural flavonoid, acts as a strong COX-2 inhibitor and is of considerable interest due to its anti-inflammatory and potential antitumor properties. In the next experiment, we examined the interaction of cysteine with COX-2. Cysteine was specifically chosen due to its –SH (thiol) group, which is highly reactive and capable of forming critical interactions within or near the active site of enzymes. Additionally, cysteine residues are sensitive to redox modifications and may play a regulatory role in COX-2 function during inflammatory processes. Docking results revealed a binding energy of –4.0 kcal/mol, which corresponds to a calculated inhibition constant (Ki) of approximately 1180 μM, indicating weak inhibitory activity. Interaction analysis showed that cysteine formed conventional hydrogen bonds with GLU B:236, ARG B:333, GLN B:241, and GLU A:140, along with a π–sulfur interaction involving TRP A:139 (Figure 4C). These results suggest that cysteine interacts with COX-2 in a weak and less stable manner, implying only a limited role in modulating COX-2–mediated inflammatory mechanisms.22,23
IL-6
Interleukin-6 (IL-6) belongs to the cytokine family and plays an important role in regulating inflammation, immune responses, and metabolic processes in the body. It is produced by macrophages, T-lymphocytes, fibroblasts, and endothelial cells. IL-6 levels increase under various external stimuli, including infection, tissue injury, or stress, thereby activating protective mechanisms. IL-6 stimulates the immune system by promoting the differentiation of B-lymphocytes into plasma cells and enhancing antibody production. In hepatocytes, it induces the synthesis of acute-phase proteins such as C-reactive protein. In addition, IL-6 contributes to hematopoiesis by supporting leukocyte maturation and also influences lipid and glucose metabolism. Clinically, elevated IL-6 levels are observed in autoimmune diseases, chronic inflammatory conditions, and malignancies. During COVID-19 infection, a sharp increase in IL-6 is associated with the “cytokine storm” syndrome. Therefore, IL-6 is considered not only an important diagnostic marker but also a therapeutic target, with drugs such as tocilizumab already applied in clinical practice. In our in silico study, we performed molecular docking between IL-6 and the carnitine enzyme. The binding free energy was calculated as –4.2 kcal/mol. The corresponding inhibition potential was determined as Ki = 840 μM, which indicates a weak inhibitory effect. The interaction was mediated through amino acid residues in carnitine forming specific contacts with IL-6, including LEU B:92 and ASN B:144 via conventional hydrogen bonds, and GLU B: through unfavorable acceptor–acceptor interactions (Figure 3).
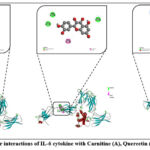 |
Figure 3: Molecular interactions of IL-6 cytokine with Carnitine (A), Quercetin (B), and Cysteine (C). |
In the next stage of our study, we performed molecular docking of IL-6 with quercetin. The binding free energy obtained from this interaction was calculated as –7.8 kcal/mol. Based on this value, the inhibition constant (Ki) was determined to be approximately 1.9 μM, which indicates a considerably stronger inhibitory potential compared to carnitine. Detailed interaction analysis revealed that quercetin established several stabilizing contacts with amino acid residues of IL-6. Specifically, conventional hydrogen bonds were formed with ALA A:152, ASN A:135, and GLU A:129. Additionally, PHE A:136 was involved in a π–π stacking interaction, while ALA A:152 formed a π–alkyl bond. Furthermore, GLU A:129 contributed to a π–anion interaction (Figure 2B). These multiple non-covalent interactions suggest that quercetin exhibits a more stable and energetically favorable binding mode with IL-6, highlighting its potential as a stronger inhibitory candidate in comparison to carnitine. In the subsequent experiment, we carried out docking analysis of IL-6 with cysteine. The calculated binding free energy for this complex was –4.5 kcal/mol, corresponding to an estimated inhibition constant (Ki) of approximately 540 μM. This value reflects a relatively weak binding affinity when compared to carnitine and quercetin. Detailed interaction analysis revealed that cysteine formed multiple conventional hydrogen bonds with TRP C:264, THR A:263, THR C:247, and THR C:245 residues of IL-6. Interestingly, TRP C:264 was also engaged in an unfavorable donor–donor interaction in addition to a π–alkyl contact, suggesting partial instability in the binding conformation. Moreover, π–sigma interactions were detected with HIS C:269 and TYR C:238, further contributing to the molecular recognition between cysteine and IL-6 (Figure 2C). Although cysteine demonstrated the ability to establish hydrogen bonding and aromatic interactions with IL-6, the relatively low binding energy and higher Ki value indicate that these interactions are not sufficiently stable to exert a strong inhibitory effect. In comparison to quercetin, which displayed a much higher binding affinity (–7.8 kcal/mol), and carnitine (–4.2 kcal/mol), cysteine appears to be the least effective ligand in terms of inhibitory potential. Nevertheless, its interactions provide valuable insight into the role of small sulfur-containing amino acids in modulating cytokine activity, which may be further explored in combination with other bioactive compounds to enhance therapeutic potential.24,25
MAPK
MAPK (Mitogen-Activated Protein Kinase) is one of the most important components of intracellular signaling pathways. It is activated by various external stimuli, including growth factors, cytokines, stress, and pathogens, and regulates a wide range of cellular responses. The MAPK cascade consists of a three-tiered kinase module in which MAPKKK (MAPK kinase kinase) phosphorylates MAPKK (MAPK kinase), which in turn activates MAPK. Activated MAPK translocates into the nucleus and modulates transcription factors, thereby influencing processes such as cell proliferation, differentiation, stress response, and apoptosis. The MAPK pathways are mainly divided into three groups: ERK (extracellular signal-regulated kinase), JNK (c-Jun N-terminal kinase), and p38 kinases. ERK is primarily associated with cell growth and development, whereas JNK and p38 play crucial roles in inflammation and stress response.26,27 From a clinical perspective, dysregulation of MAPK signaling is observed in cancer, inflammatory diseases, and neurodegenerative disorders. Therefore, MAPK pathways are considered promising therapeutic targets in drug development. In our docking study with MAPK, carnitine exhibited a binding energy of –4.4 kcal/mol, with an estimated inhibition constant (Ki) of approximately 630 μM, indicating weak inhibitory potential. The interaction was stabilized through several types of bonds: a conventional hydrogen bond with ALA A:157, a carbon hydrogen bond with LEU A:108, and a salt bridge with LYS A:165. In contrast, an unfavorable negative–negative interaction was observed with GLU A:163, which reduced the overall stability of the complex (Figure 4).
The reason for these specific interactions can be attributed to the physicochemical properties of the residues involved. ALA A:157, being a small nonpolar residue, easily forms hydrogen bonds with polar groups of carnitine, providing localized stabilization. LEU A:108, with its hydrophobic side chain, contributes to weaker but supportive carbon–hydrogen bonding. LYS A:165, a positively charged residue, readily interacts with the negatively charged carboxyl group of carnitine, forming a salt bridge that reinforces electrostatic attraction. Conversely, GLU A:163 is negatively charged, and when positioned near the acidic groups of carnitine, it creates electrostatic repulsion, which explains the unfavorable interaction observed. Taken together, these results highlight that the binding of carnitine to MAPK is determined by a balance of stabilizing hydrogen bonds and electrostatic interactions, counteracted by unfavorable charge repulsion. This explains why the overall binding affinity remains relatively weak despite multiple contacts within the binding pocket.
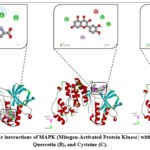 |
Figure 4: Molecular interactions of MAPK (Mitogen-Activated Protein Kinase) with Carnitine (A), Quercetin (B), and Cysteine (C). |
In our docking study with MAPK, quercetin demonstrated a binding energy of –7.8 kcal/mol, with an estimated inhibition constant (Ki) of approximately 2 μM, indicating a strong inhibitory potential compared to carnitine. The interaction profile revealed multiple stabilizing contacts. Quercetin formed conventional hydrogen bonds and π–cation interactions with LYS A:165, HIS A:107, and ARG A:49, which significantly contributed to the stability of the complex through both hydrogen bonding and electrostatic attraction. In addition, quercetin established a π–alkyl interaction with MET A:109, further reinforcing hydrophobic stabilization within the binding pocket (Figure 3B). However, an unfavorable acceptor–acceptor interaction was observed with LEU A:108, which slightly reduced the binding efficiency. Despite this, the overall interaction pattern was dominated by strong hydrogen bonds, cation–π contacts, and hydrophobic interactions, which together account for the notably higher binding affinity. The specificity of these interactions can be explained by the structural complementarity between quercetin and MAPK residues. The hydroxyl groups of quercetin readily donate and accept hydrogen bonds, allowing stable contacts with polar and charged residues such as LYS, HIS, and ARG. The aromatic rings of quercetin enable π–cation interactions with positively charged side chains, which provide additional electrostatic stabilization. Furthermore, its planar aromatic system favors π–alkyl interactions with nonpolar residues like MET A:109, enhancing van der Waals contacts. In our docking experiment with MAPK, cysteine exhibited a binding energy of –3.8 kcal/mol, with an estimated inhibition constant (Ki) of approximately 1.5 mM, indicating a very weak inhibitory potential compared to quercetin and carnitine. The interaction analysis revealed that cysteine engaged in both polar and non-polar contacts within the MAPK binding pocket. Specifically, π–sulfur interactions were observed with HIS A:64 and TYR A:35, while ARG A:67 formed alkyl interactions that provided weak hydrophobic stabilization. In addition, cysteine established conventional hydrogen bonds with TYR A:35 and SER A:56, contributing to localized binding stability. The nature of these interactions can be explained by the presence of the thiol (–SH) group in cysteine, which is highly reactive and capable of forming π–sulfur interactions with aromatic residues such as HIS and TYR. Meanwhile, the polar backbone groups of cysteine facilitated hydrogen bonding with residues like TYR and SER, although the strength of these bonds was not sufficient to produce a strong overall binding affinity. The alkyl interaction with ARG A:67 provided minimal hydrophobic stabilization but did not significantly improve the overall inhibitory potential.
NF-kb
Nuclear factor kappa B (NF-κB) is a transcription factor complex that plays a central role in regulating immune and inflammatory responses, cell survival, and proliferation. It is normally sequestered in the cytoplasm in an inactive form by inhibitory proteins (IκBs). Upon stimulation by cytokines, stress signals, or pathogens, IκBs are degraded, allowing NF-κB to translocate into the nucleus, where it binds DNA and regulates the expression of genes involved in inflammation, immunity, and apoptosis. Dysregulation of NF-κB signaling is closely associated with chronic inflammation, autoimmune disorders, and cancer, making it an attractive therapeutic target. In our docking experiment with NF-κB, carnitine exhibited a binding energy of –4.9 kcal/mol, with an estimated inhibition constant (Ki) of approximately 270 μM, suggesting a weak-to-moderate inhibitory potential. The interaction profile revealed both stabilizing and destabilizing contacts. Carnitine formed conventional hydrogen bonds with DG D:722 and DG D:720, as well as carbon hydrogen bonds with DC C:715, DT C:714, DT C:716, and DA D:721. These hydrogen-bonding interactions provided localized stabilization within the binding pocket (Figure 4A). However, unfavorable electrostatic interactions were also observed, specifically positive–positive repulsion with LYS B:575 and negative–negative repulsion with DG D:722. Such unfavorable interactions likely reduced the overall stability of the complex, partially counteracting the favorable hydrogen-bond contributions. The observed binding pattern can be explained by the zwitterionic nature of carnitine, which contains both positively charged (trimethylammonium) and negatively charged (carboxylate) groups. While these groups facilitate hydrogen bonding with DNA bases and amino acid residues, they also increase the risk of charge–charge repulsion when positioned unfavorably, as seen with LYS and DG. In our docking study, quercetin exhibited a binding energy of –9.4 kcal/mol against NF-κB, with an estimated inhibition constant (Ki) of approximately 140 nM, indicating a strong inhibitory potential. The interaction analysis revealed that quercetin established multiple stabilizing contacts. Conventional hydrogen bonds were formed with DA D:724, DG
D:722, DG D:720, and DC C:715, providing significant stabilization through polar interactions. Additionally, a π–cation interaction was observed with LYS B:575, further strengthening the binding via electrostatic attraction between the positively charged lysine side chain and the aromatic ring system of quercetin (Figure 5). However, an unfavorable donor–donor interaction occurred with DA D:721, which slightly weakened the overall binding efficiency. Despite this, the interaction pattern was dominated by strong hydrogen bonds and π–cation contacts, which contributed to the notably high binding affinity. These results highlight quercetin as a potent inhibitor of NF-κB, with the ability to form diverse and stable interactions within the binding pocket. Its strong binding affinity suggests that quercetin may play a significant role in modulating NF-κB–mediated pathways, potentially offering therapeutic value in conditions associated with NF-κB overactivation.28,29
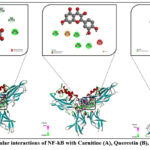 |
Figure 5: Molecular interactions of NF-kB with Carnitine (A), Quercetin (B), and Cysteine (C). |
In our docking study with cysteine, the interaction with NF-κB yielded a binding energy of –4.0 kcal/mol, corresponding to an estimated inhibition constant (Ki) in the millimolar range (~1.1 mM). This relatively weak binding affinity indicates that cysteine has a limited inhibitory effect on NF-κB compared to other tested ligands such as quercetin. Interaction analysis revealed that cysteine formed conventional hydrogen bonds with DC C:715 and DT C:714, which contribute to stabilizing the complex through polar interactions with nucleobase residues. However, an unfavorable donor–donor interaction was observed with DG D:722, introducing steric and electrostatic strain that reduces the overall stability of the complex (Figure 4C)
TNF alpha
In our docking experiment, the interaction between TNF-α and carnitine was analyzed. The docking results demonstrated a binding energy of –5.0 kcal/mol, corresponding to an estimated inhibition constant (Ki) of approximately 216 µM, suggesting a moderate binding affinity. Interaction mapping revealed that carnitine formed a conventional hydrogen bond with LEU A:120, which plays a role in stabilizing the ligand–protein complex through polar interactions. Additionally, a π–anion interaction was observed with TYR C:119, indicating electrostatic attraction between the negatively charged carboxyl group of carnitine and the aromatic ring system of tyrosine (Figure 5A). These interactions suggest that carnitine has the potential to moderately influence TNF-α activity, though the relatively higher Ki value compared to strong inhibitors indicates that its binding efficiency is limited. Nevertheless, the presence of stabilizing hydrogen bond and π–anion contacts highlights a possible role for carnitine in modulating TNF-α–mediated inflammatory responses.
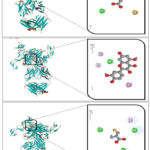 |
Figure 6: Molecular interactions of TNF alpha with Carnitine (A), Quercetin (B), and Cysteine (C). |
In our docking study, quercetin exhibited a binding energy of –9.7 kcal/mol against TNF-α, corresponding to an estimated inhibition constant (Ki) of approximately 85 nM, indicating a strong inhibitory potential. Interaction analysis revealed several stabilizing contacts. π–alkyl interactions were observed with LEU A:57 and LEU B:57, where the hydrophobic side chains of leucine provided favorable nonpolar contacts with the aromatic system of quercetin, thereby enhancing van der Waals stability. Additionally, TYR B:119 formed a conventional hydrogen bond, contributing polar stabilization through interaction between the hydroxyl group of quercetin and the polar side chain of tyrosine. Furthermore, π–π stacking was established with TYR C:119, reflecting aromatic ring-to-ring interactions 4.3 kcal/mol, corresponding to an estimated inhibition constant (Ki) of approximately 678 µM. This value indicates that cysteine acts as a weak inhibitor of TNF-α due to its relatively low binding affinity. Interaction analysis revealed that cysteine formed conventional hydrogen bonds with TYR A:151, TYR B:119, and GLY B:121, suggesting stabilization through polar contacts between the amino acid residues of TNF-α and the thiol or amino groups of cysteine. Additionally, a π–alkyl interaction was observed with TYR C:119, reflecting hydrophobic contacts between the aromatic ring of tyrosine and the sulfur-containing side chain of cysteine. Although the hydrogen bonds and π–alkyl contacts contributed to binding, the relatively high Ki value indicates that cysteine does not strongly inhibit TNF-α activity. Nonetheless, these interactions demonstrate that cysteine can still weakly interact with TNF-α and may play a role in modulating its function under certain conditions.30
Pharmacokinetic Profile by SwissADME
Quercetin
The pharmacokinetic and drug-likeness characteristics of quercetin were predicted using the SwissADME web tool. The results demonstrated that quercetin complies with Lipinski’s rule of five without any violations, confirming its overall suitability as a drug-like molecule. According to absorption and distribution predictions, quercetin exhibited high gastrointestinal absorption, which indicates good potential for oral administration. However, the compound was predicted not to cross the blood–brain barrier, suggesting that its pharmacological activity is likely restricted to peripheral tissues rather than the central nervous system. Additionally, quercetin was identified as non-substrate for P-glycoprotein (P-gp), implying that its transport and elimination are unlikely to be limited by efflux mechanisms. In terms of solubility, quercetin was classified as soluble in water according to multiple predictive models, although the values (Log S between –3.1 and –3.9) still suggest moderate solubility, which could partly explain its limited oral bioavailability reported in experimental studies (Figure 7). The lipophilicity profile (consensus Log P<sub>o/w</sub> = 1.23) indicated a moderately lipophilic nature, favorable for membrane permeability. Metabolic predictions indicated that quercetin is a potential inhibitor of several cytochrome P450 isoenzymes, specifically CYP1A2, CYP2D6, and CYP3A4, while showing no inhibitory effect on CYP2C19 and CYP2C9. This implies that quercetin may interfere with the metabolism of drugs that are substrates of these enzymes, highlighting the possibility of drug–drug interactions. The predicted bioavailability score was 0.55, indicating moderate oral bioavailability. In terms of medicinal chemistry filters, quercetin presented one PAINS alert (catechol_A) and one Brenk alert (catechol group), both of which are commonly associated with structural motifs that may produce non-specific biological activity. Despite these alerts, the compound’s synthetic accessibility score (3.23) suggests that it is feasible to synthesize.31,32
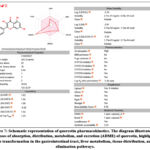 |
Figure 7: Schematic representation of quercetin pharmacokinetics. The diagram illustrates the processes of absorption, distribution, metabolism, and excretion (ADME) of quercetin, highlighting its transformation in the gastrointestinal tract, liver metabolism, tissue distribution, and elimination pathways. |
Pharmacokinetics of Cysteine and L-Carnitine
The pharmacokinetic evaluation of Cysteine and L-Carnitine revealed close similarities in solubility, safety, and metabolic stability, although both compounds showed limited gastrointestinal absorption and restricted systemic distribution. Cysteine, a sulfur-containing amino acid, demonstrated very high water solubility (Log S = 1.11), which can be attributed to its small molecular weight (121.16 g/mol), high polarity, and the presence of multiple hydrogen-bond donors and acceptors. Despite its solubility, Cysteine exhibited poor gastrointestinal absorption and no ability to permeate the blood–brain barrier (BBB), suggesting that oral delivery results in low systemic bioavailability. Importantly, Cysteine was neither a substrate for P-glycoprotein (P-gp) nor an inhibitor of key cytochrome P450 (CYP) isoforms (CYP1A2, CYP2C19, CYP2C9, CYP2D6, CYP3A4), indicating a low potential for drug–drug interactions. Its low skin permeability (Log Kp = –8.81 cm/s) reflects restricted passive diffusion across lipid membranes, consistent with its hydrophilic character. Collectively, these properties emphasize Cysteine’s safety and metabolic stability, although they limit its pharmacokinetic performance as a drug candidate unless formulated with delivery-enhancing systems. Similarly, L-Carnitine, a quaternary ammonium compound involved in fatty acid transport, also displayed high water solubility (Log S = –0.05, classified as highly soluble). However, comparable to Cysteine, L-Carnitine demonstrated low gastrointestinal absorption and an inability to cross the BBB, restricting its central nervous system availability. Like Cysteine, it was not recognized as a P-gp substrate and did not inhibit major CYP450 isoforms, supporting its favorable safety profile and minimal risk of metabolic interference. Its skin permeability (Log Kp = –7.88 cm/s) remained low, which is expected for highly hydrophilic molecules. These findings confirm that while L-Carnitine possesses excellent safety and solubility, its oral bioavailability is inherently poor, highlighting the need for alternative strategies such as transporter-mediated uptake, encapsulation, or chemical modification to enhance delivery (Figure 8).
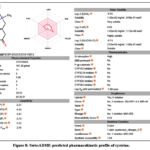 |
Figure 8: SwissADME-predicted pharmacokinetic profile of cysteine. |
 |
Figure 9: SwissADME pharmacokinetic profiles of L-Carnitine. |
Taken together, both Cysteine and L-Carnitine are safe, non-toxic molecules with no PAINS alerts and minimal risk of metabolic interference. Their main pharmacokinetic limitations are related to poor oral absorption and restricted distribution, which may limit systemic bioavailability unless improved through specialized delivery systems or formulation strategies (Figure 9).
Discussion
Inflammation is a multifactorial biological process regulated by complex interactions between immune mediators, metabolic status, and redox homeostasis. To capture this complexity, the present study integrated in vivo pharmacological evaluation, in silico molecular docking, and pharmacokinetic profiling to assess the anti-inflammatory potential of Cysteine, L-Carnitine, and Quercetin. Rather than relying on a single experimental level, this combined approach enables a more nuanced interpretation of efficacy, mechanism, and translational limitations.
In the carrageenan-induced paw edema model—an established model of acute inflammation characterized by sequential mediator release—the three compounds exhibited clearly distinct efficacy profiles. Quercetin produced the most pronounced inhibition of paw swelling, consistent with its ability to interfere with multiple inflammatory pathways. In contrast, L-Carnitine elicited only moderate edema reduction, while Cysteine showed minimal activity. Importantly, these differences cannot be attributed solely to intrinsic molecular potency. Whole-animal efficacy is strongly influenced by pharmacokinetic behavior, tissue exposure, and indirect biological effects, which likely explain the discrepancy between molecular-level predictions and in vivo outcomes.
Molecular docking provided mechanistic insights but must be interpreted with appropriate caution. Quercetin exhibited strong binding affinities toward multiple inflammatory targets, including COX-2, NF-κB, MAPK, IL-6, and TNF-α, supporting its role as a multi-target anti-inflammatory agent. This polypharmacological profile aligns well with its robust in vivo efficacy. However, docking studies inherently represent static, simplified models that do not account for protein dynamics, solvent effects, competitive endogenous ligands, or intracellular concentration constraints. Therefore, while docking supports the plausibility of direct target engagement, it does not confirm functional inhibition under physiological conditions.
L-Carnitine displayed moderate docking affinities toward selected targets such as COX-2 and TNF-α, which may partially explain its limited but measurable anti-inflammatory effect in vivo. Nevertheless, its primary biological role is metabolic rather than enzymatic inhibition, and its anti-inflammatory activity may arise indirectly through improved mitochondrial function, reduced oxidative stress, or modulation of immune cell energetics—mechanisms not captured by docking simulations. Similarly, the weak docking performance of Cysteine should not be interpreted as a lack of anti-inflammatory relevance. Instead, its contribution likely operates through glutathione synthesis and redox regulation, reinforcing the concept that not all anti-inflammatory effects are mediated by direct protein–ligand interactions.
Conclusion
The present study indicates that Cysteine, L-Carnitine, and Quercetin exert distinct and complementary effects in an acute inflammation model. Quercetin showed the most pronounced anti-inflammatory activity, supported by both in vivo outcomes and predicted target interactions, whereas L-Carnitine displayed moderate effects likely related to metabolic support rather than direct enzyme inhibition. Cysteine contributed mainly through indirect antioxidant mechanisms and showed limited efficacy in the acute model. As the study did not include direct enzymatic assays or chronic inflammation models, these findings should be interpreted as preliminary. Further investigations using extended disease models and mechanistic validation are required to clarify the translational relevance of these compounds.
Acknowledgement
We would like to thank Academy Sciences of the Republic of Uzbekistan
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
The protocols were specifically authorized by the Animal Ethics Committee of the Institute of Bioorganic Chemistry, AS RUz (Protocol Number: 133/1a/h, dated August 4, 2014).
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not applicable
Author Contributions
- Lazizbek Makhmudov conceived the study, designed the experiments, and supervised the overall project.
- Nigora Tagayalieva contributed to methodology development, data acquisition, and laboratory analysis.
- Izzatullo Abdullaev performed experiments, conducted data analysis, and contributed to the interpretation of results. Ulugbek Gayibov assisted in experimental work, validation, and preparation of figures and tables.
- Kuzijon Baratov contributed to data curation, formal analysis, and statistical processing.
- Sherzod Usmonov supported experimental procedures, resource management, and technical troubleshooting.
- Gulnora Rakhmonova participated in literature review, data interpretation, and manuscript editing.
- Rano Yaqubova contributed to writing, proofreading, and improving the scientific content of the manuscript.
- Natalia Vypova provided critical review of the manuscript, supervised analytical procedures, and ensured scientific accuracy.
References
- Al-Khayri JM, Sahana GR, Nagella P, Joseph BV, Alessa FM, Al-Mssallem MQ. Flavonoids as Potential Anti-Inflammatory Molecules: A Review. Molecules. 2022 May 2;27(9):2901.
CrossRef - Dilek, N.M., Gümrükçüoğlu, A., Durmaz, A. et al. Quality Preservation Potential of Tanacetum macrophyllum Extract in Refrigerated Meat Products: Antioxidant and Molecular Investigation. Food Biophysics 21, 3 (2026).
CrossRef - Okon IA, Aytar EC, Orie NN, Oka VO, Owu DU, Amadi NG, Bassey EK, Onuoha OG, Jacob AA, Ujong GO, Nwachukwu DC, Udoeyop FA, Uko BR, Amadi JE. Swimming exercise improves vascular function and mitigates cardiac apoptotic response via Bcl-2 and Caspase-3 pathway in streptozotocin-induced diabetic rats: in vivo and in silico molecular docking study. Eur J Med Res. 2025 Dec 17.
CrossRef - Abdugafurova DG, Oripova MZ, Amanlikova DA, et al. Immunomodulatory effects of polysaccharides isolated from Brassica rapa seeds. Pharmaceutical Chemistry Journal. 2024;57(10):1552–1556.
CrossRef - Saydullayevna IA, Gulyamovna AD, Hushvaqovich IA, et al. Biologically active effects of Rubia tinctorum L. in experimental nephrolithiasis in rats. Biomedical and Pharmacology Journal. 2024;17(3):2035–2042.
CrossRef - Saidova K, Askarov I, Islomov A, et al. Development and HPLC characterization of a water-soluble supramolecular complex of propolis and glycyrrhizic acid monoammonium salt. Biomedical and Pharmacology Journal. 2025;18(1):1017–1029.
CrossRef - Sodiqova S, Kadirova S, Zaynabiddinov A, et al. Channelopathy activity of A-41 (propyl ester of gallic acid): Experimental and computational study of antihypertensive activity. Trends in Sciences. 2025;22(9):10496.
CrossRef - Aripov TF, Gayibov UG. Antiradical and antioxidant activity of the preparation “Rutan” from Rhus coriaria L. Journal of Theoretical and Clinical Medicine. 2023;4:164–170.
- Sayidaliyeva R, Kadirova S, Zaynabiddinov A, et al. A-51 as a natural calcium channel blocker: An integrative study targeting hypertension. Trends in Sciences. 2025;22(11):10760.
CrossRef - Zaripova M, Abdullaev I, Bogbekov A, et al. In vitro and in silico evaluation of Gnaphalium extract targeting α-amylase and α-glucosidase. Trends in Sciences. 2025;22(8):10098.
CrossRef - Zoirovich OS, Ugli AIZ, Raxmatillayevich ID, et al. Effects of Ajuga turkestanica on rat aortic smooth muscle ion channels. Biomedical and Pharmacology Journal. 2024;17(2):1213–1222.
CrossRef - Mahmudov AV, Abduraimov OS, Erdonov SB, et al. Bioecological features of Nigella sativa L. under different conditions of Uzbekistan. Plant Science Today. 2022;9(2):421–426.
CrossRef - Gayibov UG, Zoirovich OA, Fotima AS, et al. Plant-derived and synthetic antihypoxic agents in cardiovascular diseases: Mechanisms and therapeutic potential. Plant Science Today. Early Access.
- Inomjonov D, Abdullaev I, Omonturdiev S, et al. In vitro and in vivo evaluation of Crataegus and Inula helenium extracts on rat blood pressure. Trends in Sciences. 2025;22(3):9158.
CrossRef - Abdullaev I, Gayibov U, Omonturdiev S, et al. Molecular pathways in cardiovascular disease under hypoxia. Journal of Biomedical Research. 2025;39(3):254–269.
CrossRef - Abdullaev A, Abdullaev I, Bogbekov A, et al. Antioxidant potential of Rhodiola heterodonta via Nrf2 activation. Trends in Sciences. 2025;22(5):9521.
CrossRef - Zaripova MR, yibova SN, Makhmudov RR, et al. Phytochemical characterization and antihyperglycemic activity of Rhodiola heterodonta. Preventive Nutrition and Food Science. 2024;29(2):135–145.
CrossRef - Mahmudov AV, Abduraimov OS, Erdonov SB, et al. Seed productivity of Linum usitatissimum L. in Uzbekistan. Plant Science Today. 2022;9(4):1090–1101.
CrossRef - Gayibov UG, Gayibova SN, Pozilov MK, et al. Influence of quercetin and dihydroquercetin on rat liver mitochondria. Journal of Microbiology, Biotechnology and Food Sciences. 2021;11(1):e2924.
CrossRef - Gayibov UG. Influence of a new polyphenolic compound from Euphorbia on mitochondrial function. Journal of Microbiology, Biotechnology and Food Sciences. 2019;8(4):1021–1025.
CrossRef - da Silva MLF, Aytar EC, Gasparotto Junior A. The Role of the NO/cGMP Pathway and SKCa and IKCa Channels in the Vasodilatory Effect of Apigenin 7-Glucoside. Molecules. 2025 Oct 31;30(21):4265.
CrossRef - Vakhobjonovna AG, Jurayevich KE, Ogli AIZ, et al. Tannins as modulators of mitochondrial dysfunction. Trends in Sciences. 2025;22(8):10436.
CrossRef - Abdurazakova I, Zaynabiddinov A, Abdullaev I, et al. Pharmacological evaluation of F-45 using in vitro, in vivo, and molecular docking approaches. Trends in Sciences. 2025;22(12):10924.
CrossRef - Mamajanov M, Abdullaev I, Sotimov G, et al. Mitochondrial and pharmacokinetic insights into 3,5,7,2′,6′-pentahydroxyflavanone. Trends in Sciences. 2025;22(12):10984.
CrossRef - Cherbal A, Aytar EC, Aydoğmuş Z, Fenghour M, Gheddar K. Comprehensive evaluation of the anti-inflammatory effects of Lavandula stoechas L.: in vivo, in vitro, and in silico studies. J Ethnopharmacol. 2026 Feb 28;357:120887.
CrossRef - Mertoglu K, Fischer A, Zargarchi S, Durul MS, Köpsel M, Aytar EC, Bulduk I, Kaki B, Esatbeyoglu T. Comparative analysis of pomological and phytochemical characteristics in white- and red-fleshed pitaya (Hylocereus spp.), with molecular docking insights into key bioactive compounds. J Sci Food Agric. 2025 Nov 19.
- Gaibullayeva O, Islomov A, Abdugafurova D, et al. Immunity-enhancing effects of Inula helenium root extract in sunflower oil. Biomedical and Pharmacology Journal. 2024;17(4):2729–2737.
CrossRef - Umidakhon Y, Erkin B, Ulugbek G, et al. Correction of mitochondrial dysfunction by haplogenin-7-glucoside under hypoxia and ischemia. Trends in Sciences. 2022;19(21):6260.
CrossRef - Aytar EC, Torunoğlu EI, Gümrükçüoğlu A, Al-Farraj S, Durmaz A, Sillanpää M. Assessing the chemical profile and biological potentials of Tamarix smyrnensis flower extracts using different solvents by in vitro, in silico, and network methodologies. PLoS One. 2025 Dec 1;20(12)
CrossRef - Li M, Zhang M, Aripov TF, et al. Green-synthesized Zn²⁺-polyphenol networks for multifunctional food preservation. Food Measurement. 2025.
CrossRef - Ramos, R.C., Magalhães, L.G., Veneziani, R.C.S. et al. In vitro and in silico schistosomicidal activity of hexane fraction from Copaifera oblongifolia leaves. In Silico Pharmacol. 13, 205 (2025).
CrossRef - Azimova AQQ, Islomov AX, Maulyanov SA, et al. Determination of vitamins and pharmacological properties of Vitis vinifera syrup-honey. Biomedical and Pharmacology Journal. 2024;17(4):2779–2786.
CrossRef







