Manuscript accepted on :02-06-2025
Published online on: 14-08-2025
Plagiarism Check: Yes
Reviewed by: Dr. Randa Salah Gomaa Mahmoud
Second Review by: Dr. Niharika Kondepudi
Final Approval by: Dr. Kamal Upreti
Aryan Sinha1 , Rekha Verma2
, Rekha Verma2 , Yogita Ale1*
, Yogita Ale1* , Vikash Jakhmola3
, Vikash Jakhmola3 , Sanjeev Kumar Shah4
, Sanjeev Kumar Shah4 , Arif Nur Muhammad Ansori5,6
, Arif Nur Muhammad Ansori5,6 and Manoj Kumar Mishra7
and Manoj Kumar Mishra7
1Department of Pharmaceutics, Uttaranchal Institute of Pharmaceutical Sciences, Uttaranchal University, Dehradun, Uttarakhand, India.
2Department of Management, DBS Global University, Dehradun, Uttarakhand, India.
3Department of Pharmaceutical Chemistry, Uttaranchal Institute of Pharmaceutical Sciences, Uttaranchal University, Dehradun, Uttarakhand, India.
4Department of Computer Sciences, Uttaranchal Institute of Technology, Uttaranchal University, Dehradun, Uttarakhand, India.
5Department of Virology and Bioinformatics, Postgraduate School, Universities Airlangga University, Surabaya, Indonesia.
6Department of Virology and Bioinformatics, Virtual Research Centre for Bioinformatics and Biotechnology, Surabaya, Indonesia.
7Department of Pharmaceutics, Amity Institute of Pharmacy, Amity University, Gwalior, Madhya Pradesh, India.
Corresponding Author E-mail: yogitaale7@gmail.com
Abstract
Neurogenomics is an interdisciplinary field that focuses on studying the genetic fundamentals of neurodegenerative diseases, seeking to understand how genetic variations contribute to the onset, progression, and severity of diseases such as Alzheimer's and Parkinson's diseases. This review aims to investigate and critically assess the evolving roles of neurogenomics and molecular technologies in the understanding, early identification, and treatment advancement for Alzheimer's and Parkinson's diseases. Neurogenomics aims to find specific genetic mutations, polymorphisms, and epigenetic variables to provide new therapeutic targets and potential biomarkers for early diagnosis. Molecular diagnosis plays a pivotal role in identifying genetic risk factors and distinguishing between different neurodegenerative conditions. This article seeks to emphasize recent progress in gene expression profiling, biomarker identification, and transgenic model research, while addressing the therapeutic significance of tools such as PCR, ISH, and Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR). The study highlights how the integration of genomic techniques with molecular biomarkers can enhance disease prognosis, patient stratification, and individualized therapy approaches for neurological diseases.
Keywords
Alzheimer's disease; Neurodegenerative diseases; Neurogenomics; Molecular Diagnosis; Parkinson's disease
| Copy the following to cite this article: Sinha A, Verma R, Ale Y, Jakhmola V, Shah S. K, Ansori A. N. M, Mishra M. K. Application of Neurogenomics and Molecular Technologies for Alzheimer's and Parkinson’s Diseases. Biomed Pharmacol J 2025;18(October Spl Edition). |
| Copy the following to cite this URL: Sinha A, Verma R, Ale Y, Jakhmola V, Shah S. K, Ansori A. N. M, Mishra M. K. Application of Neurogenomics and Molecular Technologies for Alzheimer's and Parkinson’s Diseases. Biomed Pharmacol J 2025;18(October Spl Edition). Available from: https://bit.ly/3VWYnFS |
Introduction
Neurogenomics is the intersection of two different branches, neuroscience and genomics. This has emerged as a critical discipline in identifying the genetics and molecular framework for numerous neurodegenerative diseases (ND).1 Neurogenomics becomes helpful in the detection of genes related to neurological diseases and studying the transgenic models of neurological diseases. According to recent studies, the study of neuropathology at the molecular level has become much easier due to neurogenomics. In-situ hybridization (ISH) and polymerase chain reaction (PCR) are vital methodologies in the investigation and diagnosis of neurological disorders, including Parkinson’s and Alzheimer’s diseases. ISH enables researchers to identify and localize specific gene mutations or aberrant RNA expression directly inside brain tissue samples, offering insights into disease-associated cellular alterations. For instance, ISH can assist in identifying modified expression of genes associated with neurodegeneration. PCR is employed to amplify and analyze specific DNA or RNA sequences, facilitating the identification of genetic risk factors (like mutations in the APP, PSEN1, or LRRK2 genes) and assisting in the monitoring of molecular alterations associated with disease progression. These techniques are essential for comprehending the molecular origins of neurodegenerative diseases to promote early detection and developing of targeted therapies.
The imprints of neurogenomics are quite significant in molecular neuropharmacology as it assists in encoding different genes related to neurotransmitter receptors and ion channels, leading to a better understanding of the mechanism of action of drugs, that facilitates better drug development. Additional possible applications of neurogenomics are the investigation of genetic anomalies in neurological diseases, the analysis of gene expression within the central nervous system (CNS), and the imaging of gene expression in the CNS.2 Neurogenomics examines the expression and regulation of genes inside the neurological system in both normal and pathological states. Changes in gene expression can elucidate the critical molecular processes that are disturbed in neurodegenerative diseases. Neurogenomics has facilitated a more sophisticated approach to neurodegenerative diseases through enhanced diagnostic instruments. The identification of biomarkers using genetic analysis of patient samples, including cerebrospinal fluid or blood, facilitates precise illness detection. 1,3,4 Molecular diagnostics refers to the clinical use of molecular technology pertaining to DNA and RNA to elucidate, diagnose, and monitor human diseases.
The objective of this study is to investigate and critically assess the evolving roles of neurogenomics and molecular technologies in the understanding, early identification, and treatment advancement for Alzheimer’s and Parkinson’s diseases. This article seeks to emphasize recent progress in gene expression profiling, biomarker identification, and transgenic model research, while addressing the therapeutic significance of tools such as PCR, ISH, and Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR). The study highlights how the integration of genomic techniques with biomarker can enhance disease prognosis, patient stratification, and individualized therapy approaches for neurological diseases.
Neurodegenerative disorder
Neurodegenerative diseases are intricate conditions that cause deterioration of the structure and functioning of the CNS, which leads to diminished quality of life and distressing symptoms. Neurodegenerative diseases lead to irreversible damage to the CNS, resulting in motor impairments. Neurodegenerative illnesses result in detrimental outcomes such as memory impairment, diminished motor control, and decreased communication abilities, all of which substantially diminish the quality of life.1 Neurodegenerative diseases impact millions globally, with aging being the primary contributing factor to the onset of all varieties of these conditions. Although the gene expression linked to neurodegenerative diseases occurs, the timing and severity of neurodegeneration are contingent upon the surrounding environment.6
Among the broad range of neurodegenerative diseases, several are recognized for their distinctive clinical characteristics and significant societal impact. Diseases such as Parkinson’s, Alzheimer’s, amyotrophic lateral sclerosis, and Huntington’s are marked by a progressive degeneration of a significant quantity of neurons, unlike the selective and static neuronal loss caused by metabolic or toxic conditions. Neurodegenerative illnesses are frequently marked by the aggregation of certain proteins and anatomical vulnerability. These illnesses exhibit numerous fundamental mechanisms linked to increasing neuronal malfunction and demise, including proteotoxic stress, oxidative stress-related programmed cell death, and neuroinflammation.7 In these diseases, there is an accumulation of proteins with changed physicochemical properties, referred to as misfolded proteins. Certain illnesses, such as spastic paraplegia and certain variants of spinocerebellar ataxia, do not exhibit identifiable protein inclusions using existing detection methodologies. A novel notion known as prion-like spreading suggests that proteins linked to neurodegenerative illnesses disseminate throughout the brain. Cell culture and animal models have been utilized in neurodegenerative disease research to illustrate that cells internalize pathogenic proteins and subsequently transmit them to adjacent cells.6
The primary obstacle in the development of pharmaceuticals for neurodegenerative illnesses is the blood-brain barrier (BBB). Notwithstanding numerous triumphs achieved by surgical and highly invasive procedures, their clinical adoption remains restricted due to apprehensions regarding their long-term effects, particularly the possible harm to the blood-brain barrier (BBB). Nanotherapeutics are being developed as viable alternatives for reversing neurodegeneration, having the capability to traverse the blood-brain barrier. 6
Alzheimer’s disorder (AD)
Dementia encompasses a wide range of cerebral illnesses marked by a progressive deterioration in cognitive abilities, ultimately resulting in heightened death rates. The predominant kind of dementia includes AD. This disorder results in a gradual decline of cognitive functions, with loss of memory as the most common early sign. Neuropathologically, AD is defined by neuronal degeneration in particular brain parts, namely the medial temporal lobes, and temporoparietal association cortices. The actual cause of AD has not been confirmed yet, but a well-known theory named as “amyloid hypothesis” explains the cause of AD. According to this proposition, the aggregates of amyloid-beta (Aß) plaques play the primary role in the disease mechanism and progression of AD. Due to the ongoing research on this disorder, the focus has been shifted to another potential pathogenesis of AD, but still the “amyloid hypothesis” remains the most renowned concept of AD.8 The other pathogenic characteristic of AD is the intracellular buildup of hyperphosphorylated tau protein, resulting in the formation of neurofibrillary tangles. Tauopathies are a class of neurological diseases characterized by the pathological accumulation of tau protein in the brain. Tauopathy is a key characteristic of AD that leads to synaptic dysfunction and neuronal degeneration. Tauopathies encompass more than AD, including conditions like frontotemporal dementia and progressive supranuclear palsy.9
A patient suspected of having Alzheimer’s disease must go through a series of examinations, including a neurological test, an MRI (magnetic resonance imaging) of the brain, and vitamin B12 laboratory testing.
Alzheimer’s disease biomarkers are divided into two categories: those that indicate brain amyloid pathology (such as PET and CSF amyloid-beta levels) and those that represent neuronal damage. CSF tau proteins are neuronal injury biomarkers, as is fluorodeoxyglucose (FDG)-PET for assessing cerebral metabolism and magnetic resonance imaging (MRI) for detecting brain shrinkage.12
Alzheimer’s disease Neuropathology
Individuals suffering from AD exhibit two types of alterations: positive lesions, marked by accumulation of neurofibrillary tangles, amyloid plaques, and negative lesions, marked by pronounced atrophy due to the loss of neural tissue, neuropil, and synapses. Extracellular aggregates of beta-amyloid protein (Aß) known as senile plaques display various architectural forms, such as neuritic, dispersion, dense-cored, and typical compact plaques. The ß-secretase and γ-secretase proteolytic enzymes promote Aß deposits production from the amyloid precursor protein (APP). Diverse kinds of Aß monomers may coalesce to generate soluble oligomers as well as amyloid plaques that can propagate into the brain. The abnormal filaments, neurofibrillary tangles, at a specific stage, intertwine to create paired helical filaments (PHF) as well as aggregates at neural perikaryal cytoplasm, axon, and dendrites, leading to the destruction of tubulin-associated proteins and cytoskeletal microtubules.12
Biomarkers of AD
Biomarkers play a significant role in the clinical management of Alzheimer’s disease, enabling early and precise diagnosis, tracking disease progression, forecasting therapeutic response, and differentiating Alzheimer’s from other dementias, thereby enhancing personalized and timely interventions. The biomarker of AD is classified into categories such as amyloid pathology, Tau pathology biomarkers and Presenilin () mutation.
Amyloid pathology
The Aß component accumulation is a crucial process in the development of the amyloid plaques. Aß peptides arise from the breakage of APP, the processing of APP is shown in Figure 1. The senile plaques are developed due to Aß fragments accumulation. Extracellular deposits of Aß protein (senile plaques) can take various forms, including diffuse, neuritic, classic, dense-cored, and compact. Proteolytic enzymes like ß-secretase and γ-secretase help produce Aß deposits from APP.12 APP is a type I membrane protein with eight isoforms due to alternative splicing. Neurons produce APP695, while microglia create APP751 and APP770. A soluble extracellular domain with an NH2 terminal and an intracellular COOH terminal segment (AICD) of 83 amino acids (C83) is produced by α-secretase and processed by γ-secretase. Beta-secretase has two cleavage sites, with the main one in the Aβ region, producing an 89-amino-acid AICD (C88). An intracellular segment featuring the Aß region (C99) results from the alternate position. The γ-secretase cleaves the C99 fragment, resulting in the Aß1-40 and Aß1-42 peptide variants, along with additional fragments.13 Biomarkers of cerebrospinal fluid (CSF) for detecting AD include the measurement of Aß42 content in CSF utilizing antibody-based methods such as ELISA enzyme-linked immunosorbent assay and antibody-independent methods like spectrometry. Recent discoveries indicate that patients with AD exhibit reduced quantities of Aß42 in their cerebrospinal fluid. The reduction in Aß42 indicates the peptide aggregation in the brain, resulting in the formation of senile plaques, as demonstrated by postmortem findings and in-vivo amyloid PET imaging.9
Tau pathology
The microtubule protein, Tau located in axons, is involved in axonal movement and microtubule assembly. Typically, each mole of tau protein has 2-3 moles of phosphorus. Tau possesses six isoforms that differ in the number of microtubule linkages in repeats.13 The tau gene is found on the 17thchromosome at band location 17q21 and comprises 16 exons. Tau protein is classified as a microtubule-associated protein (MAP), characterized by its heat resistance and minimal susceptibility to acid treatment. This characteristic of tau protein arises from its low secondary structure composition. The tau protein typically exists in an unfolded conformation. Due to the extreme flexibility and varied conformations of disordered proteins, nuclear structure spectroscopy is the sole viable approach that offers high-resolution imaging. This enables the acquisition of a comprehensive backbone assignment for 441 residues of tau, the longest isoform of tau protein found in the human CNS.14
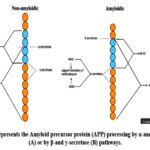 |
Figure 1: Represents the Amyloid precursor protein (APP) processing by α-and γ-secretase (A) or by β-and γ-secretase (B) pathways.Click here to view Figure |
The excessive aggregation of phosphorylated tau in disease-specific inclusions serves as a pathogenic indicator in both primary and secondary tauopathies. NFT tau can be enhanced via detergent along with acid extraction, facilitating the isolation of insoluble, accumulated tau from the AD patient’s brain.15 In Alzheimer’s disease, the usual function of tau protein in maintaining an orderly cytoskeleton within the axonal process is compromised due to its diminished ability to connect to microtubules. Post-translational modifications (PTMs) are regarded as the primary reason for the dysfunction of tau protein, however, the precise explanation remains a subject of contention. The tau phosphorylation modulates its binding capacity to microtubules, however, normal phosphorylation levels are essential for good tau functionality, whereas hyperphosphorylation results in the loss of its biological activity. The longest version of tau protein, consisting of 441 amino acids, contains around 80 possible phosphorylation sites for serine or threonine. In instances of pathological tau protein, the aberrant phosphorylation of tau may occur from tau kinase(s) or the downregulation of tau phosphatase(s), however, these factors are not mutually exclusive. Several enzymes have been assessed, with particular emphasis on kinases that are considered pivotal in the context of protein kinase A (PKA) and calcium/calmodulin-dependent protein kinase II (CaMK-II), among which glycogen synthase kinase 3 beta (GSK-3ß) was regarded as the most significant in phosphorylation processes. Patients with AD exhibit abnormally phosphorylated cytosolic tau (AD P-tau), which is not linked to tubulin or facilitates microtubule assembly, hence losing its biological function. The hyperphosphorylated tau protein intertwines to create paired helical filaments, resulting in the formation of neurofibrillary tangles (NFT).14
Presenilin (PSEN) mutation
Neurogenomics discovered mutations in the PSEN1, PSEN2, and APP genes that are connected to increased sensitivity to Alzheimer’s disease. Homeologous genes PSEN1 and PSEN2 are recognized by vertebrates. To assemble a biologically active γ-secretase (GS) complex, which cleaves amyloid precursor protein (APP), Nicastrin, presenilin enhancer 2 (), and anterior pharynx-defective 1 interact with presenilins (PSEN1 and PSEN2). Recently, approximately 300 distinct pathogenic variants have been found in the PSEN1 gene. The PSEN1 gene mutations are mainly responsible for familial Alzheimer’s disease (FAD). The PSEN1 protein functions as a component of the catalytic subunit of γ-secretase (GS). Aß40 is associated with around 90% of individuals suffering from Alzheimer’s disease. The influence of PSEN1 on Aß40 production has been elucidated through two possibilities. According to the amyloid hypothesis, mutations in PSEN1 initiate disease pathogenesis by increasing Aß42 production. This theory is supported by research indicating that minor clinical PSEN1 mutations result in elevated plasma levels of Aß40, contributing to FAD. Additionally, there is one more theory that is presenilin hypothesis, which states that PSEN1 mutations may lead to the essential presenilin function loss in the brain, hence precipitating dementia and neurodegeneration in FAD. There are 38 known PSEN2 mutations, resulting in the increased activity of γ-secretase (GS). elevated Aß42/40 ratios, an increased Aß42 concentration, and extracellular Aß40 reduction. Excluding the Lys306f and Glu26fs frameshifts, most PSEN2 mutations are nonsynonymous. Mutations in PSEN2 result in an extended disease duration and a delayed age of onset. In addition to FAD, mutations in PSEN2 are also implicated in other neurodegenerative diseases. 16
Parkinson’s disease (PD)
Parkinson’s disease is among the most common movement problems and a frequent neurodegenerative ailment. The defining hallmarks of PD are neuronal mortality at substantia nigra, resulting in striatal dopamine insufficiency and the buildup of α-synuclein in intraneuronal inclusions. Additional pathogens of Parkinson’s disease include oxidative stress, mitochondrial malfunction, cellular calcium dysregulation, neuroinflammation, and abnormalities in many neurotransmitter systems. The primary motor characteristics of PD include slowness of movement (bradykinesia), stiffness, and rest tremor. In addition to these, Parkinson’s disease is linked to a diverse array of non-motor symptoms that augment the total illness burden. The significant non-motor symptom of PD is Cognitive impairment, occurring six times more frequently in affected persons than in the healthy population. Parkinson’s disease patients’ cognitive impairment varies from subjective cognitive deterioration (SCD) to moderate cognitive impairment (PD-MCI) to dementia. SCD signifies the condition wherein individuals sense a deterioration in their cognitive abilities. SCD is not associated with any acute incident. Conversely, PD-MCI is a cognitive impairment identified by an informant or a clinician, linked to cognitive abnormalities evident in formal neuropsychological assessments or a global cognitive abilities scale. PDD is characterized by abnormalities in executive functions, attention, visuospatial skills, and memory.17
Parkinson’s disease is characterized by the accumulation of α-synuclein protein aggregates in the neurons of the substantia nigra, referred to as Lewy bodies and Lewy neurites. The Queen Square Brain Bank clinical diagnostic criteria for PD focus on the levodopa-responsive motor phenotype and neuronal degeneration in the substantia nigra and caudate and putamen dopaminergic denervation. PD can be seen as a genetic disorder in a specific context, where certain patients possess characteristic uncommon variations that result in familial disease, and in a broader context, where many patients exhibit a polygenic predisposition to the disease. Three recognized autosomal dominant genes (SNCA, LRRK2, and VPS35) and three recognized autosomal recessive genes (PRKN, PINK1, and DJ1) are identified as causative factors of Parkinson’s disease.18
Biomarkers of PD
Imaging biomarkers
Various imaging technologies, including Magnetic Resonance Imaging (MRI), Transcranial Sonography, and Single Photo Emission Computed Tomography, are employed to detect structural alterations in the brain. The methods for evaluation include measuring dopamine transporter concentration, vesicular monoamine transporter level, postsynaptic dopamine receptor levels, aromatic amino acid decarboxylase activity, and proteins (α-synuclein, tau) or iron in the brain. A unique substantia nigra dopaminergic neuron aggregation called nigrosome-1, is observable with high-resolution MRI; the characteristic ‘swallow-tail’ morphology of a healthy nigrosome-1 is absent in Parkinson’s disease. Standard routine structural magnetic resonance imaging scans may have limited utility in diagnosing Parkinson’s disease, however, advanced MRI scans are potentially beneficial for supporting diagnosis of the condition.19
Genetic biomarkers
Genome-wide association research (GWAS), linkage analysis, and exome sequencing have identified around 28 chromosomal loci associated with hereditary types of Parkinson’s disease (PD), of which 18 are part of the PARK family, designated as Park 1/4, 2, 3, 5, 6, etc. Mutations in the loci of these genes can result in monogenic variants of familial Parkinson’s disease that have recessive or autosomal dominant inheritance. The LRRK2 (PARK8) mutation and SNCA (PARK1/4) mutation, which generate α-synuclein, are monogenic variants of PD that cause autosomal dominant PD. SNCA mutations are rare, although duplication and triplication of the locus can cause familial Parkinson’s disease, especially in triplication cases.18
Biochemical markers
A diverse array of molecules has been analysed in blood, cerebrospinal fluid, and biological fluids, including dopamine and its metabolites, different amine and amino acid neurotransmitters, oxidative damage markers, neuropeptides, purine metabolites including uric acid, inflammatory markers, , and microRNAs. Brain-derived neurotrophic factor (BDNF) is a neurotrophin, important for neuronal development, survival, and proliferation. The BDNF levels variation in the cerebrospinal fluid and blood among PD patients are intricate and inconsistent, rendering BDNF unsuitable as a possible biomarker. Purine derivative uric acid may be an indicator of risk and diagnostic for Parkinson’s disease. Recent research indicates that diminished levels of uric acid in the blood and cerebrospinal fluid serve as a biomarker for PD. The dopamine and its metabolites reduction in CSF and blood highlight the decline of central dopaminergic activities, which correlates with motor impairment. Conserved short RNAs called miRNAs influence gene expression post-transcriptionally. The deregulation of miRNAs may serve as a crucial indicator for diagnosis. A further possible application of miRNA is the reestablishment of miRNA composition modified in illness. Modulating the levels of altered miRNA can facilitate repair.20
Alpha–synuclein as a biomarker
Alpha-synuclein is a small, inherently disordered protein. Their aggregation results in the creation of Lewy bodies. The detection of α-synuclein is regarded as a marker in the PD diagnosis (19). Alpha-synuclein is primarily located in the presynaptic vesicles. It is related to the synuclein family. Alpha-synuclein constitutes 1 % of the proteins in the neuronal cytoplasm. The mutant alpha-synuclein protein (A30P and A53T disease mutations) associated with familial Parkinson’s disease has been demonstrated to possess structural deficiencies in membrane binding, resulting in modifications to the protein’s binding characteristics.18 Multiple family variants of Parkinson’s disease exhibit elevated expression of alpha-synuclein attributed to SCNA gene duplication, including A30P, E46K, H50Q, A53E, and A53T. Various mutations and undesirable biomolecular conditions cause alpha-synuclein to misfold and transform into amyloid fibrils abundant in beta sheets.21
Cerebrospinal fluid is an optimal biological fluid for identifying biomarkers associated with Parkinson’s disease. Multiple studies indicate that changes in α-synuclein levels in the CSF indicate PD. Individuals suffering from PD show elevated alpha-synuclein oligomer levels relative to age-matched controls. Multiple system atrophy (MSA) can be distinguished from PD through CSF alpha-synuclein, phosphorylated tau, FMS like tyrosine kinase 3 ligand (FTL3). Post-translational changes of alpha-synuclein influence its solubility and increase its susceptibility to accumulation. Phosphorylation at Ser129 is regarded as the most prevalent pathogenic post-translational alteration of alpha-synuclein, as shown by mass spectrometry.19 Alpha synuclein is proposed to modulate the distribution of synaptic vesicles and the dimensions of presynaptic terminals. High alpha-synuclein levels interact with the soluble N-ethylmaleimide-sensitive factor attachment protein receptor () protein complex to modify endoplasmic reticulum-Golgi vesicular trafficking, impede priming, and decrease synaptic connection, reducing dopamine and glutamate neurotransmission. Most of alpha-synuclein’s effects depend on membrane binding. The combination of both order and disorder of alpha-synuclein configurations at the membrane surface may affect protein function and produce aggregation.21
Advances in Genomic Technologies
The improvement in DNA sequencing over the last few decades has fundamentally transformed the concepts of gene sequencing and tagging. Conventional techniques such as Sanger sequencing were laborious and expensive, although they established the groundwork for genetic analysis. The new Next-generation sequencing (NGS) technology has significantly enhanced our comprehension of human genomes. The NGS has instigated a paradigm change in personalized medicine, cancer genomics, and uncommon illness diagnoses.
Next-Generation Sequencing (NGS)
The NGS has revolutionized the understanding of genome structure, functionality, and dynamics. NGS can sequence several million DNA fragments at once, providing genetic variation, genome, and gene expression data. NGS has provided us with accessible and thorough genome sequencing, as well as advancements in metagenomics, epigenomics, and other genomic research. Illumina, Pacific Biosciences, and Oxford Nanopore NGS technology enable parallel reading of millions to billions of DNA fragments, revolutionizing genomics. This possibility opens new pathways for understanding genetic variability, gene expression, epigenetic alteration, and microbiological diversity. There have been three generations of gene sequencing: the first generation (Sanger sequencing), the second generation, and the third generation (the most recent progress in gene sequencing). NGS analyzes DNA using three distinct methodologies: entire genome sequencing, whole exome sequencing, and targeted sequencing, as shown in Figure 2.22
Whole genome sequencing
This method examines an individual’s full DNA sequence through genomic analysis. It describes a patient’s genome, such as regulatory regions, genes, and non-coding elements. Cancer research, uncommon genetic illness, population genetics, and innovative eukaryotic and prokaryotic genome assembly are the principal uses of whole genome sequencing.22
Whole-exome sequencing (WES)
This sequencing method captures and sequences the exome, the protein-coding part of the genome. The exome contains most pathogenic mutations despite being only 1-2% of the genome.22
Targeted sequencing panels
This technique concentrates on a limited selection of certain genes for purposes such as diagnosis, therapy monitoring, and prognosis. Profiling various clinical samples with reduced tumor concentration and DNA quality, such as circulating tumor DNA and formalin-fixed paraffin-embedded specimens, yields superior sequencing depth compared to non-next-generation sequencing approaches like PCR.22
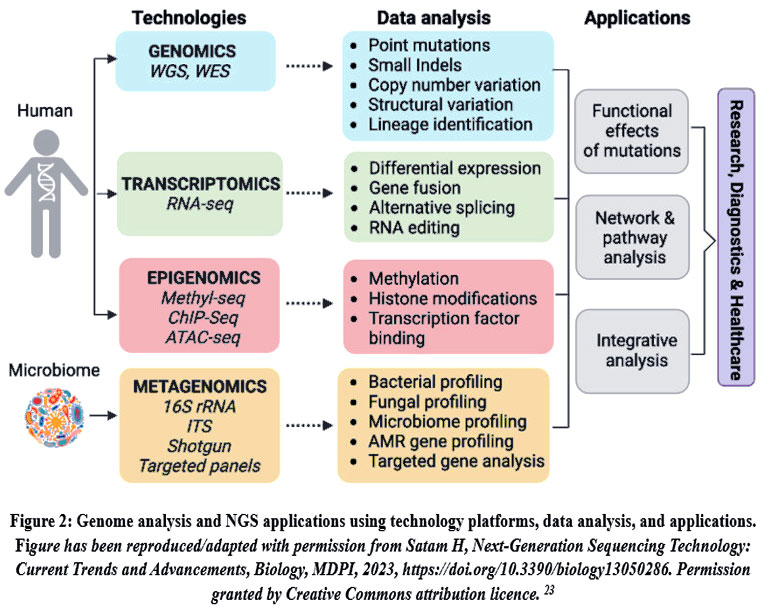 |
Figure 2: Genome analysis and NGS applications using technology platforms, data analysis, and applications. Figure has been reproduced/adapted with permission from Satam H, Next-Generation Sequencing Technology: Current Trends and Advancements, Biology, MDPI, 2023, https://doi.org/10.3390/biology13050286. Permission granted by Creative Commons attribution licence. 23Click here to view Figure |
CRISPR-based tools
Quantitative polymerase chain reaction (qPCR) has been extensively utilized for nucleic acid-based diagnostics. The adaptability and strength of PCR render it an exceptionally useful instrument for identifying DNA and RNA biomarkers, as shown in Figure 3. Nonetheless, the expense of chemicals utilized in PCR is substantial, and the methodology necessitates advanced laboratory facilities and equipment. Consequently, to circumvent the exorbitant costs and complexities associated with existing standards, it became imperative to devise a technique superior to PCR. CRISPR-based diagnostics, utilizing clustered regularly spaced short palindromic repeats, possess the capability to meet all these requirements. CRISPR systems are key to a microbial adaptation process that recognizes foreign nucleic acids by sequence and eliminates them via endonuclease activity related to Cas enzymes. The initial CRISPR-based diagnostic techniques were mostly developed with Cas9 variations to identify double-stranded DNA. Cas9 has three mechanisms: 1) guide-directed restoration of damaged proteins by inactive catalytic Cas9 supporters, 2) Cas9-mediated cleavage of Protospacer Adjacent Motif () containing patterns, and 3) Non-targeted DNA strand unwinding to serve as an aiming site for isothermal amplification. 24 The principal genes identified in the etiology of AD are APP, PSEN1, and PSEN2. Among these, PSEN1 mutations are the principal cause of FAD (dominant autosomal mutation) and lead to the early development of the condition. PSEN1 mutation is associated with high Aß42 level. The CRISPR/Cas9 method possesses the capability to efficiently correct autosomal dominant mutations. A recent study indicated that the CRISPR-Cas9 technique can target the PSEN1M146L allele associated with Alzheimer’s disease, thereby restoring the altered Aß42/40 ratio that contributes to the progression of the diseases.
The α-synuclein expression is intricately associated with the SNCA gene. The missense mutation Ala52Thr (A53T) in SNCA is recognized as a significant contributor to the early PD generation. A recent study revealed that the CRISPR-Cas9 method can eliminate the A53T-SNCA gene, thereby dramatically ameliorating PD conditions, including the overproduction of α-synuclein, neurodegeneration, and motor symptoms associated with the disorder.25
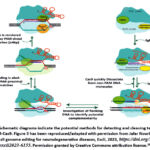 |
Figure 3: Schematic diagrams indicate the potential methods for detecting and cleaving target DNA via CRISPR-Cas9. Figure 3 has been reproduced/adapted with permission from Jafar Nouri Nojadeh, CRISPR/Cas9 genome editing for neurodegenerative diseases, Excli, 2023, https://doi.org/10.17179/excli2023-6155. Permission granted by Creative Commons attribution license.26Click here to view Figure |
Combining Neurogenomics with molecular technologies
The incorporation of neurogenomics into neurological illnesses has markedly enhanced our capacity to comprehend, diagnose, and manage various neurodegenerative conditions. Neurological illnesses include a diverse range of conditions such as epilepsy, autism, AD, PD, and other dementias. Genetic diagnostic approaches have seen increased utilization in recent years owing to technological advancements. These technologies have transformed the area of neurology by enabling more precise diagnosis, treatment, and prognosis. The application of NGS technology, including whole genome sequencing (WGS), has facilitated the identification of genetic variants associated with numerous neurological diseases. The advancement of technology has enhanced diagnosis and propelled the profession of medicine towards personalized treatment. This personalized treatment can be predicated on the individual’s genetic composition. The integration of machine learning (ML) and artificial intelligence (AI), advanced neuroimaging, renders the future of molecular techniques hopeful. The merging of neurogenomics and molecular has enabled extensive genetic analysis and improved diagnoses for neurological illnesses.27
Challenges and Future Perspective
The prospects of NGS possess significant potential for enhanced gene sequencing. The next sequencing platforms will require reduced quantities of input DNA and reagents, hence facilitating gene sequencing. These technologies will become increasingly portable, facilitating enhanced diagnostic applications. Fields such as clinical diagnosis, cancer, and microbial genomics are being transformed by NGS. In the future, it is expected to have a major effect in areas involving single-cell genomics, disease mechanisms, a greater understanding of biological processes, and personalized treatment techniques. Ongoing technology advancements and cost reductions will render NGS technologies more economical and accessible, hence allowing their integration into everyday clinical practices. 23
The domains of synthetic biology, medicines, diagnostics, and metabolic engineering have been transformed by the CRISPR advancements. This groundbreaking technology has surpassed earlier methods, including ZEN (zinc finger nuclease) and TALEN (transcription activator-like effector nuclease) by reducing toxicity and non-target effects. Since the emergence of CRISPR-Cas variations, they have been effectively utilized in prokaryotes and eukaryotes for various applications, including the treatment of human diseases like as cardiovascular disease, cancer, sickle cell anemia, and cystic fibrosis. Challenges persist with advanced technology, including: 1) off-target effects, 2) efficiency of HDR, and 3) delivery of the CRISPR-Cas system. 4) Ethical considerations of gene editing in human embryos. Overall, CRISPR-Cas technology has emerged as an optimal instrument for several diagnostic applications. Advancements in gene editing technologies are unveiling novel pathways for accurate gene modification in many prokaryotic and eukaryotic organisms.28 Overall, progress in gene therapy, stem cell regeneration techniques, and precision medicine, specifically α-synuclein targeting, LRRK2 inhibitors, and CRISPR genome editing, offers potential for modifying disease development and enhancing patient outcomes. Furthermore, the utilization of digital health systems and neuroimaging is improving patient surveillance and individualized treatment approaches.
Conclusion
In conclusion, Neurogenomics is significant in transforming the understanding and treatment of neurodegenerative disorders. The incorporation of next-generation sequencing (NGS) technologies has facilitated the identification of genetic variations responsible for Alzheimer’s Disease (AD) and Parkinson’s Disease (PD). Molecular biomarkers are essential for early and precise diagnosis. In AD, cerebrospinal fluid (CSF) indicators such as reduced Aβ42 and increased phosphorylated tau (p-tau181 and p-tau217) levels signify amyloid pathology and neurofibrillary tangles.
In Parkinson’s disease, the alpha-synuclein seed amplification assay (αSyn-SAA) has surfaced as a promising diagnostic instrument. Research has shown that αSyn-SAA can detect misfolded α-synuclein in cerebrospinal fluid with a sensitivity of 87.7% and a specificity of 91.9%.
The use of CRISPR-Cas9 to excise the A53T mutation in the SNCA gene, linked to early-onset PD, has demonstrated enhancements in diminishing α-synuclein aggregation and alleviating motor symptoms in preclinical animals. The integration of Neurogenomics, biomarker identification, and gene-editing technologies signifies a new epoch in precision medicine. These developments improve diagnostic precision and facilitate early intervention while also establishing a foundation for tailored therapy approaches designed to modify the progression of neurodegenerative disorders.
Acknowledgment
The author expressed gratitude to Mr. Jitender Joshi, President, and Prof. (Dr.) Dharam Buddhi, Vice Chancellor of Uttaranchal University Dehradun, for their assistance in the publication of this review work.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials
Permission to reproduce material from other sources
Not Applicable
Author Contributions
- Aryan Sinha: Writing – Original Draft.
- Rekha Verma: Review and resources.
- Yogita Ale: Conceptualization and Visualization.
- Vikash Jakhmola: Supervision.
- Sanjeev Kumar Shah: Review and resources.
- Arif Nur Muhammad Ansori: Review and resources.
- Manoj Kumar Mishra: Review and resources.
References
- Adeleye OO, Olorunlowu DR, Yusuf JA, Opoola EK, Akanbi ST, Akindokun SS. Neurogenomics contribution to neurodegenerative Diseases. Nepal Journal of Neuroscience. 2024;21(1):3-8. doi:10.3126/NJN.V21I1.58673
CrossRef - Artika IM, Dewi YP, Nainggolan IM, Siregar JE, Antonjaya U. Real-Time Polymerase Chain Reaction: Current Techniques, Applications, and Role in COVID-19 Diagnosis. Genes (Basel). 2022;13(12). doi:10.3390/genes13122387
CrossRef - Ale Y, Verma R, Nainwal N, Shah SK, Jakhmola V, Pant P. IoT in Revolutionizing the Pharmaceutical Sector: Applications and Challenges. Biomedical and Pharmacology Journal. 2025;18(December Spl Edition):1-15. doi:10.13005/bpj/3069
CrossRef - Sautha V, Butola M, Chaudhary M, et al. Overview of 3d Printing Technology with Pharmaceutical Applications, Challenges and Future Aspects. Biomedical and Pharmacology Journal. 2025;18(December Spl Edition):17-32. doi:10.13005/bpj/3070
CrossRef - Jain KK. Applied Neurogenomics. http://www.ashley-pub.com
- Kovacs GG. Concepts and classification of neurodegenerative diseases. In: Handbook of Clinical Neurology. Vol 145. Elsevier B.V.; 2018:301-307. doi:10.1016/B978-0-12-802395-2.00021-3
CrossRef - Dugger BN, Dickson DW. Pathology of neurodegenerative diseases. Cold Spring Harb Perspect Biol. 2017;9(7). doi:10.1101/cshperspect.a028035
CrossRef - Chopade P, Chopade N, Zhao Z, Mitragotri S, Liao R, Chandran Suja V. Alzheimer’s and Parkinson’s disease therapies in the clinic. Bioeng Transl Med. 2023;8(1). doi:10.1002/btm2.10367
CrossRef - Lashley T, Schott JM, Weston P, et al. Molecular biomarkers of Alzheimer’s disease: progress and prospects. Dis Model Mech. 2018;11(5). doi:10.1242/dmm.031781
CrossRef - Singh B, Day CM, Abdella S, Garg S. Alzheimer’s disease current therapies, novel drug delivery systems and future directions for better disease management. Journal of Controlled Release. 2024;367:402-424. doi:10.1016/j.jconrel.2024.01.047
CrossRef - Johnson KA, Fox NC, Sperling RA, Klunk WE. Brain imaging in Alzheimer disease. Cold Spring Harb Perspect Med. 2012;2(4). doi:10.1101/cshperspect.a006213
CrossRef - Breijyeh Z, Karaman R. Comprehensive Review on Alzheimer’s Disease: Causes and Treatment. Molecules. 2020;25(24):5789. doi:10.3390/molecules25245789
CrossRef - Ferrari C, Sorbi S. The complexity of Alzheimer’s disease: an evolving puzzle. Physiol Rev. 2021;101(3):1047-1081. doi:10.1152/physrev.00015.2020
CrossRef - Kolarova M, García-Sierra F, Bartos A, Ricny J, Ripova D. Structure and Pathology of Tau Protein in Alzheimer Disease. Int J Alzheimers Dis. 2012;2012:1-13. doi:10.1155/2012/731526
CrossRef - Wegmann S, Biernat J, Mandelkow E. A current view on Tau protein phosphorylation in Alzheimer’s disease. Curr Opin Neurobiol. 2021;69:131-138. doi:10.1016/j.conb.2021.03.003
CrossRef - Kabir MdT, Uddin MdS, Setu JR, Ashraf GM, Bin-Jumah MN, Abdel-Daim MM. Exploring the Role of PSEN Mutations in the Pathogenesis of Alzheimer’s Disease. Neurotox Res. 2020;38(4):833-849. doi:10.1007/s12640-020-00232-x
CrossRef - Aarsland D, Batzu L, Halliday GM, et al. Parkinson disease-associated cognitive impairment. Nat Rev Dis Primers. 2021;7(1):47. doi:10.1038/s41572-021-00280-3
CrossRef - Morris HR, Spillantini MG, Sue CM, Williams-Gray CH. The pathogenesis of Parkinson’s disease. The Lancet. 2024;403(10423):293-304. doi:10.1016/S0140-6736(23)01478-2
CrossRef - Ganguly U, Singh S, Pal S, et al. Alpha-Synuclein as a Biomarker of Parkinson’s Disease: Good, but Not Good Enough. Front Aging Neurosci. 2021;13. doi:10.3389/fnagi.2021.702639
CrossRef - Surguchov A. Biomarkers in Parkinson’s Disease. In: Neuromethods. Vol 173. Humana Press Inc.; 2022:155-180. doi:10.1007/978-1-0716-1712-0_7
CrossRef - Srinivasan E, Chandrasekhar G, Chandrasekar P, et al. Alpha-Synuclein Aggregation in Parkinson’s Disease. Front Med (Lausanne). 2021;8. doi:10.3389/fmed.2021.736978
CrossRef - Pei XM, Yeung MHY, Wong ANN, et al. Targeted Sequencing Approach and Its Clinical Applications for the Molecular Diagnosis of Human Diseases. Cells. 2023;12(3):493. doi:10.3390/cells12030493
CrossRef - Satam H, Joshi K, Mangrolia U, et al. Next-Generation Sequencing Technology: Current Trends and Advancements. Biology (Basel). 2023;12(7):997. doi:10.3390/biology12070997
CrossRef - Kaminski MM, Abudayyeh OO, Gootenberg JS, Zhang F, Collins JJ. CRISPR-based diagnostics. Nat Biomed Eng. 2021;5(7):643-656. doi:10.1038/s41551-021-00760-7
CrossRef - Nojadeh JN, Eryılmaz NSB, Ergüder Bİ. CRISPR/CAS9 GENOME EDITING FOR NEURODEGENERATIVE DISEASES. EXCLI J. 2023;22:567-582. doi:10.17179/excli2023-6155
- Jafar Nouri Nojadeh NSBEBIE. CRISPR/Cas9 genome editing for neurodegenerative diseases. EXCLI Journal . 2023;3(22):567-582.
- Montgomery RM. Functional Analysis of Genetic Tests in Neurological Diagnostics: Advances and Future Prospects. Published online September 9, 2024. doi:10.20944/preprints202409.0618.v1
CrossRef - Gohil N, Bhattacharjee G, Lam L, Perli SD, Singh V. CRISPR-Cas systems: Challenges and future prospects. doi:10.1016/bs.pmbts.2021.01.00801.008







