Manuscript accepted on :29-08-2025
Published online on: 19-09-2025
Plagiarism Check: Yes
Reviewed by: Dr. Abeer Gatea
Second Review by: Dr. Anjaneyulu Konuri
Final Approval by: Dr. Kapil Joshi
Rekha Sharmily Rajadhurai* , Karthik Balaguru
, Karthik Balaguru and Vijayan Thiruvengadam
and Vijayan Thiruvengadam
Department of ECE, Bharath Institute of Higher Education and research, Chennai, Tamil Nadu, India
Corresponding Author E-mail:rekhasharmily@gmail.com
Abstract
Brain tumors are malignant neoplasms characterized by one of the lowest survival rates. Deep learning, an established and vigorous machine learning technology, has been extensively employed in numerous applications to address complicated challenges demanding exceptional accuracy in the medical domain. Classifying a brain tumor is an essential step after its identification to formulate an effective treatment strategy. The primary goal of this research is to build a framework for the automatic classification of brain tumor with less computation time. Three classifiers namely, Inception V3, Xception, and EfficientNetB3 were utilized, incorporating a callback function along with L1 and L2 regularization to optimize the models, maintain sparsity, and preserve key features. The three models exhibited impressive outcomes, however the Xception network displayed the higher accuracy of 98.78%, precision, recall and F1 measure of 98.33%, 99% and 98.67%. The outcomes are compared with various previous studies and state-of-the-art techniques conducted using the same dataset applied in our work. The metrics reveal that, the suggested research provides valuable insights toward automatically classifying brain tumors with fewer computational units and time. In future this model can be interpreted with unknown dataset.
Keywords
Brain Tumor; Deep learning; Efficientnet B3; Inception V3; MRI; Xception
| Copy the following to cite this article: Rajadhurai R. S, Balaguru K, Thiruvengadam V. An Enhanced Framework Leveraging Pre-Trained CNN Models for Brain Tumor Classification in MRI Scans. Biomed Pharmacol J 2025;18(October Spl Edition). |
| Copy the following to cite this URL: Rajadhurai R. S, Balaguru K, Thiruvengadam V. An Enhanced Framework Leveraging Pre-Trained CNN Models for Brain Tumor Classification in MRI Scans. Biomed Pharmacol J 2025;18(October Spl Edition). Available from: https://bit.ly/4gstfHs |
Introduction
Brain tumors are lumps created when brain cells proliferate abnormally and the brain’s systems of regulation malfunction.1 Within the realm of radiology, early brain tumor prediction and categorization is a crucial area of research that aids in choosing the most practical course of therapy to preserve the lives of patients.2 A biopsy is used to classify brain tumors, which is typically done after the decisive brain surgery. The National Academy of Medicine (2015) estimated that approximately 12 million patients in the U.S. experience diagnostic errors annually, with delayed or missed cancer diagnoses accounting for a significant percentage. Depending on the type and appearance of the tumor, delays in diagnosis for brain tumors in particular might last anywhere from 30 to 90 days. This is frequently caused by modest symptoms and radiological discrepancies (Brain Tumor Foundation, 2022). Delays may result in more intrusive operations or worse prognoses.
Radiologists can diagnose tumors utilizing deep learning based technological advancements without using invasive procedures.3 The noninvasive, painless medical imaging technique known as magnetic resonance imaging generates scans of the internal organs of the human body. Since it generates high-resolution scans of the brain tumor, it is frequently utilized and regarded as the most reliable and efficient method for detecting and classifying malignancy. Over the past few years, medical image classification has attracted much focus. The most commonly utilized neural network model for this task is the Convolutional Neural Network.4 Deep neural networks, can classify tumors by analyzing their type, size, and unique characteristics, enabling a more personalized treatment approach.5.6 Few models can even assess tumor aggressiveness, helping to plan treatment intensity accordingly. Deep learning models can integrate image analysis with additional patient data, such as genetic information and medical history, to offer a more comprehensive assessment of the tumor. This comprehensive perspective allows for more effective guidance in determining the treatment type and course. Deep learning models can be implemented on a large scale, enabling high-quality diagnostic services for broad populations, including those in remote areas with limited specialist access. So that prognosis of the type of tumor can be analyzed within a few minutes with less cost.
Three fine-tuned optimized deep learning networks namely Inception V3, Xception and the EfficientnetB3 are employed for the grouping of brain tumor in to three classes.7 The significance of our work is given as follows
Three classifiers namely Inception V3, Xception and the EfficientnetB3 are employed for sorting brain tumors in to glioma, meningioma and pituitary classes
The callback function is deployed for optimizing the model
L1 and L2 regularization are used for establishing the sparse model and to retain the features
The Adamax optimizer technique that can efficiently accommodate sparse gradients is implemented.
The Relu and Softmax activation functions are employed in the hidden and output layer for implementing nonlinearity and to sort images into classes
The manuscript is organized into five sections. The I, II, III, IV and V sections comprises of the introduction, related works, methodology, results and discussions and conclusion respectively.
Related Works
The related study was based on models run on the benchmark Kaggle dataset, which is employed in our work. A study was performed to predict and classify brain tumors using a hybrid model employing ViT and GRU. The spatial features and the temporal features extricated using ViT and GRU. When combined together, ViT extracts rich spatial features, and GRU captures temporal or ordered relationships, resulting in a robust hybrid system. This approach is especially powerful in brain tumor classification where both fine-grained structural details and sequential patterns across slices or modalities are critical. The model is tested with two datasets. The model generated high outcomes. The model’s efficiency is compared with 8 DL models and three optimizers.8 Accordingly, a convolutional neural network (CNN) with three multi-scale channels was developed to extract small, medium, and large features. These multi-scale features were then fused to enhance classification performance, effectively capturing tumors of varying sizes and textures.9 The model is tested for various metrics. Badža MM and Barjaktarović MČ.10 proposed a CNN model with K fold cross validation for the 3-class classification of brain tumor. A deep learning network was designed to group the 3 classes of the brain tumor. Thus, two models such as Efficientnet B0 and InceptionResnetV2 were used with a sparse autoencoder. Feature fusion is performed using the Quantum Theory-based Marine Predator Algorithm (QTbMPA) to enhance classification accuracy. The model is further optimized with Bayesian optimization to fine-tune parameters. The system effectively classifies tumors into glioma, meningioma, and pituitary types, achieving high outcomes.11 An enhanced deep learning method for classifying brain cancer from MRI images was developed. The approach utilizes residual networks (ResNet) to overcome vanishing gradient issues and allow deeper feature learning. The model is trained to classify brain tumors into glioma, meningioma, and pituitary tumor categories. It demonstrates high accuracy and robustness in tumor classification. The study confirms the effectiveness of ResNet in extracting complex features from medical imaging data.12 A brain tumor categorization model is established by based on CNN technique and high outcomes are achieved.13 The multi-scale feature scaling was implemented through MultiFeNet, and a three-class grouping of images were performed.14 Two DL techniques namely Inception V3 and Xception models were employed for 3class classification, the finetuned DL models were further combined with ML models and it was observed that Inception V3 gave higher outcomes. After identifying the presence of tumor, it is mandatory to find the type of tumor, so we have compared the performance of three classifiers.
Materials and Methods
The necessary libraries are imported from the TensorFlow. keras library. This research was carried out in 13th Gen Intel(R) Core (TM) i5-13420H 2.10 GHz, RAM: 16GB, NVIDIA RTX 2050 machine. The dataset is taken from the online source, freely accessible web-based platform the Kaggle.15 The dataset contains 3064 MRI scans belonging to meningioma, pituitary and glioma categories, it comprises tumor images from all the axis such as sagittal, axial, and coronal. The dataset consists of 708 MRI images of meningioma, 1426 MRI images of glioma and 930 MRI images of pituitary tumor. Then the dataset is broken in to 3 sets of data, out of which 80% of the data is separated for training, 10% of the data is allotted for validating and 10% of the data for testing. The train set data undergoes augmentation. The augmentation of image dataset is done with Rotation of ±15°, Zoom of ±10%, Width/height shift of ±5% and Horizontal flip.
The input image is rescaled to 224 x 224 pixels. The callback function is implemented for optimizing training, through reducing the learning rate and to stop the training when the accuracy stops increasing. Three classification algorithms such as Inception V3, Xception, EfficientnetB3 are employed for classification of the input images as pituitary, meningioma and glioma classes.16 Subsequently, the outcomes of the classifiers are analysed through various assessment metrics and their performances are contrasted. The proposed architecture is shown in figure1, the L1 and L2 regularizers were employed for implementing the sparse model and to retain all the features. The L1 and L2 values are 0.00001and 0.001respectively. The maxpooling layers reduces the feature maps proportions. The batch normalization is applied to normalize and thus stabilize the outcomes of the previous layer. A dropout layer is implemented to eradicate overfitting. The Adamax optimizer is implemented for enhancing the model and Relu function is used to implement nonlinearity and for computing the losses categorical cross entropy function is deployed. The dense layer with softmax function is implemented for classification.
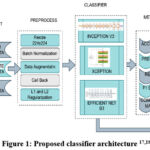 |
Figure 1: Proposed classifier architecture 17,18,19 |
The processed images are fed in to the Inception V3 network.20 The Inception Network is a pretrained convolutional network. It is trained with millions of images from Imagenet datasets and they can classify up to thousand output classes. The Inception modules consist of various filter sizes in the convolutional layers. By processing numerous filter sizes simultaneously, enables the network to assist the model in learning features at various scales. It lowers the number of parameters and processing by using factorized convolutions, which divide convolutions into smaller operations. Training Inception V3 can be accelerated and stabilized by employing strategies like batch normalization.
The research was further performed with Xception net and EfficientnetB3 classifiers. The Xception net model, exists as a light weight Inception model. EfficientNet’s parameter efficiency is a compelling advantage, for mobile or resource-constrained environments, our primary goal was to develop a high-performance model for specialized clinical applications In the Inception model filters of various sizes are used and their outputs are concatenated whereas in the Xception network a stack of depth wise separable convolutions are performed. As a result, the model is light weight and efficient. The residual connections prevent the model from the vanishing gradient problem and thus capable of training very deep networks.
The EfficientnetB3 classifiers architecture is based on inverted mobile bottleneck convolutional blocks. It undergoes compound scaling to uniformly scale the dimensions and the resolution. Due to its transfer learning capabilities, and the squeeze excitation blocks it extracts the features vividly. They can be applied for detection, segmentation and classification tasks. They generate benchmark results even with fewer parameters.
Results
The training, testing and validation are performed through iterating the available dataset for 40 iterations with 40 samples per iteration. The callback technique is used to set the parameters. The duration of training period of all the three classifiers are also analyzed for inventing the optimal model. The outcomes of the three classifiers are evaluated by the accuracy, loss, precision, recall and F1 score of the approach.21 Moreover, the accuracy and loss plot of the classifiers are drawn and the best epoch was determined. The confusion matrix of all the three classifiers are generated. The efficiency of the classifier is assessed by accuracy, precision, recall, f1-measure and confusion matrix. 22,23 They can be computed by,

The Inception V3 classifier has provided an accuracy score of 0.9979 for training, 0.9640 for validation and 0.9642 for testing. The computation time of the model is found to be 25 min 38.37 sec. The accuracy and loss plot for training and validation are given in the figure 3. It can be observed that the accuracy elevates above the fifth epoch and thusly, the fifth epoch is found to be the best epoch. Similarly, the loss drops after the tenth epoch. It can be noted that only few MRI modalities are misclassified in the confusion matrix.
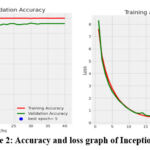 |
Figure 2: Accuracy and loss graph of Inception V3 |
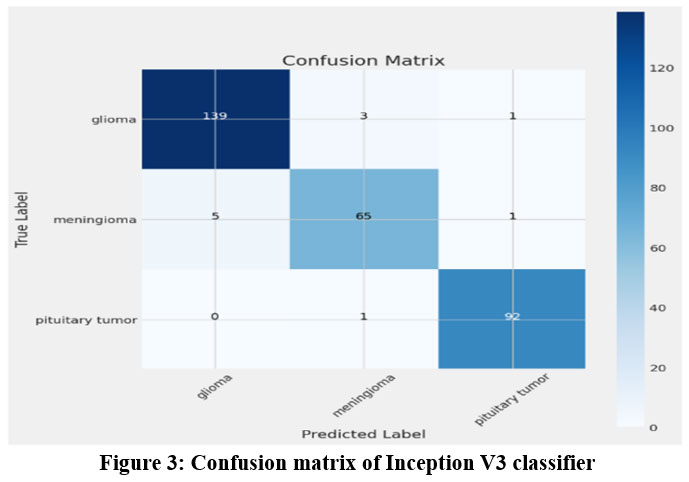 |
Figure 3: Confusion matrix of Inception V3 classifier |
The classification report generated by the InceptionV3 classifier is as follows in the table 1. The pituitary class achieved higher scores for all the measures.
Table 1: Classification report of the InceptionV3 classifier
| InceptionV3 classifier | |||
| Class of Tumor | PrecisionScore | Recallscore | F1 score |
| Glioma | 0.97 | 0.97 | 0.97 |
| Meningioma | 0.94 | 0.92 | 0.93 |
| Pituitary | 0.98 | 0.99 | 0.98 |
The Xception network attained the training, validating and testing accuracies as 1.0, 0.9837 and 0.9878 respectively. The duration of computation is 23 minutes and 25.17 seconds. Figure 5 displays the accuracy, loss, and training and validation plots for the Xception classifier. The accuracy increases above the seventh epoch and the seventeenth epoch is found to be the best epoch. Similarly, the loss drops after the tenth epoch and the 40th epoch is found to be the best epoch in the loss plot. It can be noted that only few MRI modalities are misclassified. The figure 6 shows the confusion matrix of the classifier and it is observed that almost all the images are classified correctly except a few images.
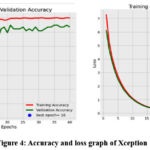 |
Figure 4: Accuracy and loss graph of Xception |
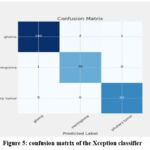 |
Figure 5: confusion matrix of the Xception classifier |
The results of the Xception classifier is given in the table 2. The pituitary class achieved the highest scores.
Table 2: Classification report of the Xception classifier
| Xception classifier | |||
| Class of Tumor | Precision Score | Recall Score | F1 score |
| Glioma | 0.99 | 0.98 | 0.99 |
| Meningioma | 0.97 | 0.99 | 0.98 |
| Pituitary | 0.99 | 1.0 | 0.99 |
The accuracy values obtained by the Efficientnet B3 classifier are 0.9996, 0.9771 and 0.9804 for training, validating and testing respectively. The training time taken is 24 min 43.23 sec. The accuracy, loss plot and the confusion matrix are given in figure 7 and 8. The accuracy plot shows that the tenth epoch is the best, the loss reduces rapidly after the 10th epoch and the 38th epoch is found to be the best epoch. Moreover, in the confusion matrix only very few images are misclassified.
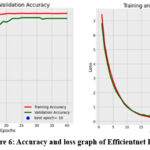 |
Figure 6: Accuracy and loss graph of Efficientnet B3 |
Table 3: Classification report of the Efficientnet B3 classifier
| Efficientnet B3 classifier | |||
| Class of Tumor | Precision Score | Recall score | F1 score |
| Glioma | 0.99 | 0.98 | 0.98 |
| Meningioma | 0.95 | 0.97 | 0.96 |
| Pituitary | 1.0 | 0.99 | 0.99 |
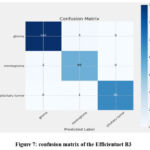 |
Figure 7: confusion matrix of the Efficientnet B3 |
Discussion
The table 4 and 5 compares the performance of the three classifiers. Likely, the Xception model exhibited the highest training, testing and validating accuracy. Moreover, it elapsed the least training time of 23 min 25.17 sec. The quantity of parameters in total are less in the Efficientnet approach, however the other two approaches have nearly twice the number of parameters. On comparing the efficiency of all the three approaches, the Xception networks achieved the highest accuracy with least duration for computation. The Xception network is an extreme inception network which uses the depth wise separable convolutions and pointwise convolutions for classification.
Table 4: Comparison of loss and accuracy of the classifier
| CLASSI FIER | ACCURACY | LOSS | ||||||
| TRAI N | VALI DATE | TEST | TRAIN | VALI DATE | TEST | |||
| Inception V3 | 0.9979 | 0.9640 | 0.9641 | 0.2663 | 0.3605 | 0.3588 | ||
| Xception | 1.0 | 0.9837 | 0.9878 | 0.1266 | 0.1636 | 0.1756 | ||
| Efficient netB3 | 0.9996 | 0.9771 | 0.9804 | 0.1270 | 0.1927 | 0.1944 | ||
Table 5: Comparison of Performance of the classifier
| CLASSIFIER | TEST ACCUR ACY |
COMPUTATION TIME | TOTAL PARAMETERS | TRAINABLEPARAMETERS |
| InceptionV3 | 0.9641 | 25 min 38.37 sec | 22,336,291 | 22,297,763 |
| Xception | 0.9878 | 23 min 25.17 sec | 21,394,987 | 21,336,363 |
| Efficient net B3 | 0.9804 | 24 min 43.23 sec | 11,183,922 | 11,093,547 |
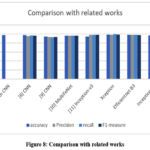 |
Figure 8: Comparison with related works |
Table 6: Comparison of Performance of the classifier with related work
| Refere nce | No. of images |
Class | Model | Accur acy | Precis ion | Rec all | F1 measure |
| [5]DíazPernas | 3064 | 3 | Multi path CNN | 0.973 | NA | NA | NA |
| [6] Badža | 3064 | 3 | CNN | 97.39 | 95.44 | 96.94 | 96.11 |
| [9] S. Das | 3064 | 3 | CNN | 94.39 | 93.33 | 93 | 93.33 |
| [10] Agrawal | 3064 | 3 | Multi FeNet | 96.4 | 96.4 | 96.4 | 96.4 |
| [11]N.Noreen | 3064 | 3 | Inception-v3 | 94.34 | 98 | 98 | 98 |
| Proposed work | 3064 | 3 | XceptionEfficientnetB3InceptionV3 | 98.7898.0496.41 | 98.339896.33 | 999896 | 98.6797.6796 |
The ability of the suggested models in categorizing brain tumor images were examined and contrasted with the models that already exists.
The comparison is performed on approaches that have utilized the dataset which was employed in our proposed work. When our models’ accuracy is compared to that of other relevant models that use the same dataset, it is found that our model produced the best results. The average precision, recall and F1 score are compared and the proposed Xception network scored the highest outcomes. The Xception network generalize better by learning complex spatial relationships in tumor images. It also reduces the redundancies of features, thus improving performance on unknown new datasets.
Xception’s exceptional performance stems from its remarkable efficiency in capturing spatial hierarchies and channel-wise relationships. The Xception model replaces standard convolutions with depthwise separable convolutions, which split the convolution process into two distinct steps, a depthwise convolution that filters each input channel separately, and a pointwise convolution that combines the outputs. This design significantly reduces computational cost and overfitting while maintaining the ability to learn complex patterns. By separating spatial and cross-channel feature learning, Xception minimizes unnecessary feature mixing and noise, leading to more efficient and accurate feature extraction especially useful in medical imaging tasks like brain tumor classification.
Conclusion
In this research Inception V3, Xception and EfficientnetB3 were employed for classifying the MRI modalities into glioma, meningioma and pituitary classes. The pretrained Xception, EfficientnetB3, InceptionV3 models are optimized by using callback function to reduce learning rate, and L1 and L2 regularization is used for sparse model and to retain all the features. The Xception classifier attained the highest evaluation scores such as an accuracy of 98.78%, average precision, recall and F1 score of 98.33%, 99% and 98.67% respectively. Furthermore, the computation time was found to be 23 min 25.17 sec which was a short period. Being able to scale seamlessly across layers allows for robust learning of intricate tumor patterns. Furthermore, its lightweight architecture allows for the processing of larger amounts of input per iteration, increasing both convergence speed and accuracy. It reduces the human errors in diagonizing the presence of tumors, aids in the early-stage prognosis of brain tumors, regardless of their complexity in shape, size, or location, by effectively capturing both fine-grained and global features through its efficient convolutional design.
This model can be deployed for the automatic classification of brain tumor and it can aid the clinicians for the efficient analysis within a short duration. The number of trainable parameters are less thus requires less computation cost. So we can conclude that the depthwise separable convolutions are more efficient than other types of convolutions. Furthermore, this model holds potential for future adaptation to classify other types of cancers that are diagnosed using medical imaging techniques, provided it is retrained on relevant data and validated on larger, diverse datasets.
Acknowledgement
The authors gratefully acknowledge the support provided by Bharath Institute of Higher Education and Research in carrying out this research
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials
Permission to reproduce material from other sources
Not Applicable
Authors’ Contribution
- Rekha Sharmily Rajadhurai contributed to the conceptualization, methodology design, data collection, and drafting of the original manuscript.
- Karthik Balaguru oversaw the research through supervision, performed data analysis, and contributed to reviewing and editing the manuscript.
- Vijayan Thiruvengadam was responsible for data visualization and involved in the final review and approval of the manuscript.
References
- Solanki S, Singh U, Chouhan S, Jain S. Brain tumor detection and classification using intelligence techniques: An overview. IEEE Access. 2023;PP(0):1-1. doi:10.1109/ACCESS.2023.3242666
CrossRef - Nazir M, Shakil S, Khurshid K. Role of deep learning in brain tumor detection and classification (2015 to 2020): A review. Comput Med Imaging Graph. 2021;91:101940. doi:10.1016/j.compmedimag.2021.101940
CrossRef - Saeedi S, Rezayi S, Keshavarz H, et al. MRI-based brain tumor detection using convolutional deep learning methods and chosen machine learning techniques. BMC Med Inform Decis Mak. 2023;23:16. doi:10.1186/s12911-023-02114-6
CrossRef - Sharmily R, Rekha B, Karthik B, Vijayan T. Brain tumour detection and classification using deep learning and transfer learning techniques. 2023 Intelligent Computing and Control for Engineering and Business Systems (ICCEBS). IEEE; 2023.
CrossRef - Brain tumor detection and classification using deep feature fusion and stacking concepts. J Popul Ther Clin Pharmacol. 2024;31(1):1339-1356. doi:10.53555/jptcp.v31i1.4179
CrossRef - Sarkar A, Maniruzzaman M, Alahe MA, Ahmad M. An effective and novel approach for brain tumor classification using AlexNet CNN feature extractor and multiple eminent machine learning classifiers in MRIs. J Sensors. 2023;2023:1224619. doi:10.1155/2023/1224619
CrossRef - Ahmmed S, Podder P, Mondal MRH, Rahman SMA, Kannan S, Hasan MJ, et al. Enhancing brain tumor classification with transfer learning across multiple classes: An in-depth analysis. 2023;3:1124-1144. doi:10.3390/biomedinformatics3040068
CrossRef - Ahmed MM, Hossain MM, Islam MR, et al. Brain tumor detection and classification in MRI using hybrid ViT and GRU model with explainable AI in Southern Bangladesh. Sci Rep. 2024;14:22797. doi:10.1038/s41598-024-71893-3
CrossRef - Díaz-Pernas F, Martínez Zarzuela M, Antón-Rodríguez M, González-Ortega D. A deep learning approach for brain tumor classification and segmentation using a multiscale convolutional neural network. 2021;9:153. doi:10.3390/healthcare9020153
CrossRef - Badža MM, Barjaktarović MČ. Classification of brain tumors from MRI images using a convolutional neural network. Appl Sci. 2020;10:1999. doi:10.3390/app10061999
CrossRef - Eves K, Valasek J, Ullah MS, et al. Brain tumor classification from MRI scans: A framework of hybrid deep learning model with Bayesian optimization and quantum theory-based marine predator algorithm. Front Oncol. 2024;14:1335740. doi:10.3389/fonc.2024.1335740
CrossRef - Abdelaziz Ismael SA, Mohammed A, Hefny H. An enhanced deep learning approach for brain cancer MRI images classification using residual networks. Artif Intell Med. 2020;102:101779. doi:10.1016/j.artmed.2019.101779
CrossRef - Das S, Aranya OFMRR, Labiba NN. Brain tumor classification using convolutional neural network. In: Proceedings of the 2019 1st International Conference on Advances in Science, Engineering and Robotics Technology (ICASERT); Dhaka, Bangladesh; 2019:1-5. doi:10.1109/ICASERT.2019.8934603
CrossRef - Agrawal T, Choudhary P, Shankar A, Singh P, Diwakar M. MultiFeN et: Multi-scale feature scaling in deep neural network for the brain tumor classification in MRI images. Int J Imaging Syst Technol. 2024;34(1):e22956. doi:10.1002/ima.22956
CrossRef - Cheng J. Brain tumor dataset. 2017. doi:10.6084/m9.figshare.1512427.v5
- Azzahra TS, Jesslyn Cerelia J, Azhar Lutfi Nugraha F, Apriliyanti Pravitasari A. MRI-based brain tumor classification using Inception ResNet V2. Enthusiastic Int J Appl Stat Data Sci. 2023;3(2):163-175. doi:10.20885/enthusiastic.vol3.iss2.art4
CrossRef - Ramaneswaran S, Srinivasan K, Vincent PM Durai Raj, Chang CY. Hybrid Inception v3 XGBoost model for acute lymphoblastic leukemia classification. Comput Math Methods Med. 2021;2021:2577375. doi:10.1155/2021/2577375
CrossRef - Ibrahem H, Salem A, Kang HS. DTSNet: Depth-to-space networks for fast and accurate semantic object segmentation. 2022;22:337. doi:10.3390/s22010337
CrossRef - Matarneh S, Elghaish F, Edwards D, et al. Automatic crack classification on asphalt pavement surfaces using convolutional neural networks and transfer learning. J Inf Technol Constr. 2024;29:1239-1256. doi:10.36680/j.itcon.2024.055
CrossRef - Mahmoud A, Awad NA, Alsubaie N, et al. Advanced deep learning approaches for accurate brain tumor classification in medical imaging. 2023;15:571. doi:10.3390/sym15030571
CrossRef - Srinivasan S, Francis D, Mathivanan SK, et al. A hybrid deep CNN model for brain tumor image multi-classification. BMC Med Imaging. 2024;24:21. doi:10.1186/s12880-024-01195-7
CrossRef - Nassar SE, Yasser I, Amer HM, et al. A robust MRI-based brain tumor classification via a hybrid deep learning technique. J Supercomput. 2024;80:2403-2427. doi:10.1007/s11227023-05549-w
CrossRef - Gupta M, Sharma SK, Sampada GC. Classification of brain tumor images using CNN. Comput Intell Neurosci. 2023;2023:2002855. doi:10.1155/2023/2002855
CrossRef







