Manuscript accepted on :28-02-2025
Published online on: 06-11-2025
Plagiarism Check: Yes
Reviewed by: Dr. Digamber Singh and Dr. Tolmas Hamroyev
Second Review by: Dr. Grigorios Kyriakopoulos
Final Approval by: Dr. Eman Refaat Youness
Himanshu Pant1* , Garima Joshi2
, Garima Joshi2 and Bhupesh Rawat1
and Bhupesh Rawat1
1Department of Computer Science, Graphic Era Hill University, Bhimtal Campus, India.
2Department of Zoology, Kumaun University, Nainital, India.
Corresponding author Email: himpant7@gmail.com
DOI : https://dx.doi.org/10.13005/bpj/3303
Abstract
Chest X-ray (CXR) imaging is a fundamental diagnostic tool for detecting various thoracic diseases. However, manual interpretation is prone to errors due to subtle variations in disease patterns. This study explores the use of deep learning-based computer-aided detection (CAD) systems to enhance diagnostic accuracy and efficiency in CXR analysis. We assess the efficacy of five deep learning architectures—VGG-16, VGG-19, ResNet, InceptionNet, and CapsNet—on publically accessible CXR datasets that include various disorders. The dataset comprises 3,043 CXR images categorized into four classes: pneumonia (1,345), healthy (1,341), tuberculosis (138), and COVID-19 (217). To address potential dataset challenges, we introduce a novel approach and compare model performance against existing methods. Our results indicate that CapsNet achieves the highest accuracy of 98.48%, surpassing other models based on confusion matrix analysis and key performance metrics. The improved performance of CapsNet is due to its willingness to preserve spatial hierarchies and being resilient to alterations in picture orientation, a frequent drawback of conventional convolutional networks. These findings highlight the potential of CapsNet for improving automated chest disease detection in CXR imaging, demonstrating its viability for clinical applications and further advancements in medical AI research.
Keywords
CapsNet; Deep Learning; Precision Medicine; Thoracic Disorders; Transfer Learning
Download this article as:| Copy the following to cite this article: Pant H, Joshi G, Rawat B. Precision Medicine for the Lungs: Deep Learning Applications in Thoracic Imaging. Biomed Pharmacol J 2025;18(4). |
| Copy the following to cite this URL: Pant H, Joshi G, Rawat B. Precision Medicine for the Lungs: Deep Learning Applications in Thoracic Imaging. Biomed Pharmacol J 2025;18(4). Available from: https://bit.ly/4nGB7Y8 |
Introduction
The thorax, a vital anatomical region of the human body, houses essential organs responsible for respiration and circulation, including the lungs, heart, and associated structures.1 Given its crucial role in sustaining life, diseases affecting this region can pose severe health risks and contribute significantly to global morbidity and mortality rates. Thoracic diseases, which encompass conditions such as pneumonia, tuberculosis, cardiomegaly, and COVID-19, are estimated to impact approximately 450 million individuals annually, 2 leading to nearly four million deaths worldwide. These disorders arise from numerous pathological processes influencing the pulmonary and cardiovascular systems, necessitating prompt and precise diagnosis for efficient treatment and improved patient outcomes.3
Among the various diagnostic modalities available, chest X-ray (CXR) imaging remains one of the most widely utilized techniques due to its accessibility, cost-effectiveness, and ability to provide rapid diagnostic insights.4 CXRs are commonly used for detecting lung infections, cardiovascular abnormalities, and other thoracic conditions. However, traditional interpretation of these images relies heavily on the expertise of radiologists, making the diagnostic process susceptible to human error, particularly in high-workload settings,5 where fatigue and time constraints may impact accuracy. Additionally, subtle differences in disease manifestations can make accurate diagnosis challenging, necessitating the development of more advanced diagnostic tools to enhance efficiency and reliability.
In recent years, technological advancements in artificial intelligence (AI) and deep learning have transformed various aspects of medical imaging, including automated disease detection and classification.6 Deep learning, a subset of machine learning (ML), has demonstrated significant promise in the field of medical image analysis, particularly in identifying patterns and anomalies in CXR images. These AI-driven approaches have the potential to augment radiologists’ capabilities, reduce diagnostic errors, and improve the overall efficiency of healthcare systems.7
Chest X-ray imaging has unique characteristics that make it an essential diagnostic tool for thoracic diseases. The technique involves directing X-ray radiation through the chest, where different anatomical structures absorb varying amounts of radiation. This differential absorption produces a grayscale image in which bones, being highly radio dense, appear white, soft tissues appear in shades of gray, and air-filled structures like the lungs appear black.8 These distinctive visual characteristics allow medical professionals to identify abnormalities such as lung opacities, pleural effusions, and cardiac enlargements. Despite its advantages, the interpretation of CXR images remains complex, requiring substantial expertise and experience to differentiate between normal and pathological findings accurately.9
Over the past decade, deep learning techniques have gained significant traction in medical image analysis, offering superior performance in various diagnostic tasks, including disease classification, segmentation, and anomaly detection. Convolutional Neural Networks (CNNs) have been widely adopted in this field due to their ability to automatically extract hierarchical features from medical images.10 Numerous studies have demonstrated the effectiveness of CNN-based models in classifying conditions such as pneumonia, tuberculosis, pulmonary edema, and COVID-19. However, despite their success, conventional CNN architectures face limitations in capturing intricate spatial hierarchies and preserving crucial contextual information within images, which may hinder accurate diagnosis in complex medical scenarios.11
To address these challenges, this research proposes a novel deep learning-based approach utilizing Capsule Networks (CapsNet) for thoracic disease detection in CXR images. CapsNet, an advanced neural network architecture, introduces the concept of dynamic routing between capsules, enabling it to capture spatial hierarchies and maintain robust feature representations. Unlike traditional CNNs, which primarily rely on max-pooling operations that may discard essential spatial information, CapsNet preserves the hierarchical relationships between image features, 12 making it well-suited for medical imaging applications where structural integrity is critical.
Capsule Networks offer several advantages over conventional CNN models. One of the key strengths of CapsNet is its ability to handle variations in image orientation and perspective, a common challenge in medical imaging. Traditional CNNs often struggle with viewpoint variations, requiring extensive data augmentation techniques to compensate for these discrepancies. CapsNet, on the other hand, effectively encodes spatial relationships within images, reducing the reliance on excessive data augmentation and improving overall model generalization. This capability is particularly beneficial for CXR analysis,13 where abnormalities may present with subtle differences across patients.
The objective of this research is to compare CapsNet’s performance with that of well-known deep learning models, such as VGG-16, VGG-19, ResNet, and InceptionNet, in order to evaluate the extent to which it detects thoracic disorders. Using a widely available CXR dataset with 3,043 pictures split into four separate classes—pneumonia, TB, COVID-19, and healthy people—we train and evaluate these algorithms. We examine each model’s diagnostic performance using established assessment criteria, including accuracy, precision, recall, and F1-score,14 in order to identify the best effective method for detecting thoracic diseases.
Furthermore, this research addresses potential challenges associated with dataset imbalances and variations in image quality, proposing strategies to enhance model robustness and generalization. The integration of advanced deep learning techniques in CXR analysis holds significant potential for improving diagnostic workflows, reducing human error, and facilitating early disease detection. By leveraging the strengths of CapsNet, we aim to contribute to the ongoing efforts in developing AI-driven solutions for automated medical imaging analysis.
The remainder of this paper is organized as follows: Section 1 introduces the fundamental concepts and objectives of the proposed study. Section 2 provides an extensive review of related work and existing literature on deep learning applications in medical imaging. Section 3 outlines the materials, datasets, and methodologies employed in our research. Section 4 presents the experimental setup, performance evaluation, and comparative analysis of the deep learning models used in this study. Through this structured approach, we aim to provide a comprehensive understanding of the potential of CapsNet in enhancing thoracic disease diagnosis using CXR images, paving the way for future advancements in AI-driven healthcare solutions.
Related Work
Image classification encompasses the specialized domain of medical image categorization, leveraging various algorithms for effective analysis. These algorithms may utilize image enhancement techniques to improve the discriminability of features, enhancing classification accuracy. Existing studies on chest X-ray analysis primarily rely on artificial neural networks and traditional supervised/unsupervised machine learning approaches. This work delves into the effectiveness of alternative deep learning architectures for identifying and classifying abnormalities and clutter. Motivated by its demonstrably high performance in diverse image classification tasks, particularly within medical imaging, the convolutional neural network (CNN) is chosen as the core model for this investigation. Given its proven record of accomplishment of achieving high accuracy with minimal training data, the CNN is expected to offer exceptional performance in this application.
Ramalingam et al15 concentrated in analyzing COPD using deep convolutional neural networks (CNN) with an emphasis on image processing and machine learning. The study provides insights into COPD with COVID-19 symptoms and offers a comprehensive overview with articles achieving over 80% accuracy. The study also provides an overview of COPD with COVID-19 symptoms.
Appavu et al 16 has created a computer-aided diagnosis (CAD) system to analyze chest x-ray pictures, addressing issues related to the availability and expertise of radiologists. The system uses pre-trained CNN models and a Support Vector Machine classifier for TB detection. Data augmentation techniques enhance performance, with the proposed method achieving a 98.9% accuracy and 1.00 AUC on augmented images.
Constantinou et al17 use capsule networks (CapsNet) to automate the identification of coronary artery disease (CAD) from ECG data. The one-dimensional version of Capsule Network was used to identify Coronary Artery Disease (CAD) in electrocardiogram (ECG) segments spanning two and five seconds from a sample of 40 perfectly healthy people and 7 CAD patients. The model, named 1D-CADCapsNet, achieved a 5-fold diagnosis accuracy of 99.44% and 98.62%, respectively, with the highest performance using a two-second ECG segment.
Chen W et al18 used Artificial Intelligence (AI) technology, specifically convolutional neural networks (CNNs), for automatic segmentation and categorization of heart sounds. CNNs encounter difficulties while categorizing the mel-scale frequency cepstral coefficient (MFCC) spectrum. A Capsule Neural Network (CapsNet) was introduced to address these constraints, obtaining validation accuracies of 90.29% and 91.67% on a clinical auscultation dataset.
Thanoon et al19 introduces a novel method for screening early-stage lung cancer by using a capsule neural network and support vector machine. The method involves using image-processing methods such as lung field segmentation, feature extraction, and classification. Capsule networks overcome limitations of traditional methods, capturing spatial relationships between features. The approach achieved an accuracy of 96% on two distinct information sets, suggesting it has potential for improving early-stage lung cancer detection in regions lacking advanced screening methods.
Nasser et al 20 developed an algorithm capable of diagnosing pneumonia from frontal-view CXR images with higher accuracy than radiologists. Their system was trained using ChestX-ray14 dataset on a 121-layer Dense Convolutional Network (DenseNet).
An independent investigation employs multiple CNN models to classify chest radio- graphs as TB-positive or TB-negative, as described by Nafisah et al. 21 In assessing the efficacy of the system, two publicly accessible datasets are utilized: the Montgomery County chest X-ray set and the Shenzhen chest X-ray set. The proposed computerized demonstration framework for tuberculosis screening attains an accuracy rate exceeding 80%, a level of precision comparable to that of radiologists.
Prior studies show that deep learning may diagnose thoracic diseases on radiological pictures, relieving radiologists. This study compares deep learning algorithms for diagnosing thoracic illnesses to overcome uncertainty. We optimize VGG-16, VG-19, GoogLeNet, ResNet-50 and Capsule Neural Network for thoracic illness classification using their thoracic disease detection success.
A method for distinguishing distinct thoracic disorders using x-ray images is presented in this study. The literature study suggests that while there is a limited quantity of data available for medical imaging, CNNs need a significant number of data. Not taking into consideration the input image’s spatial orientation has a detrimental effect on CNN performance. Our focus for categorizing thoracic illness is on the capsule network (CapsNets).
Materials and Methods
Dataset Descriptions
Due to the scarcity of publicly available chest X-ray datasets, this study utilized a combination of sources to improve both model training and evaluation. Specifically, we used the NIH ChestX-ray (CHXR) dataset and a specialized COVID-19 CHXR dataset. The NIH dataset, developed by the NIH Clinical Center, includes 14,863 annotated X-ray images that cover a wide range of thoracic conditions, such as pneumonia, tuberculosis, and lung nodules. Each image is labeled to indicate the presence or absence of 14 distinct abnormalities, making it well-suited for training deep learning models under supervised learning conditions. In contrast, the COVID-19 CHXR dataset contains X-ray images from confirmed or suspected COVID-19 cases and is particularly valuable for identifying COVID-related lung issues.22
To construct a balanced and representative dataset for multi-class classification, we selected a carefully filtered subset of 3,043 images from the combined datasets. Although the NIH collection is extensive, many images are weakly annotated, contain overlapping diagnoses, or suffer from label uncertainty. To address these issues, we focused on selecting images with clear, single-disease labels and ensured approximately equal representation across four key categories: COVID-19, pneumonia, tuberculosis, and normal (healthy) cases. This selection strategy helped minimize class imbalance, reduce the impact of noisy labels, and support the development of a robust classification model.
The final dataset was divided into training and testing subsets, with 2,291 images (80%) used for training and 752 images (20%) reserved for testing. Despite their usefulness, publicly available chest X-ray datasets have several limitations, including label noise due to automated or inconsistent annotations, class imbalance and overlapping disease categories, demographic and institutional bias from limited geographic or equipment diversity, and variations in image quality and acquisition settings, all of which can affect model generalization and real-world diagnostic reliability.
For classification and segmentation tasks, we implemented a hybrid deep learning architecture that combines Capsule Neural Networks (CapsNet) with an attention-based U-Net.23 This model architecture enhances spatial feature learning, improves resilience to positional variations, and is well-suited to medical imaging applications. It accurately identifies, segments, and classifies four thoracic conditions, thereby improving diagnostic effectiveness. Additionally, the model is capable of generating synthetic chest X-ray images and segmentation masks, as illustrated in Figure 1. Through this integrated deep learning approach, the study aims to support reliable and automated diagnosis of thoracic diseases from chest X-rays.
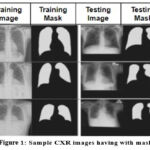 |
Figure 1: Sample CXR images having with mask |
Projected Framework
This study proposes a robust and explainable deep learning framework for the detection and classification of thoracic diseases in chest X-ray images. The model is structured in three major stages: segmentation, feature extraction, and classification, each leveraging state-of-the-art architectures to maximize diagnostic accuracy. The pipeline begins with preprocessing, where all input X-ray images are resized to 224 × 224 pixels, normalized, and standardized to eliminate variations caused by imaging equipment and acquisition protocols. These steps ensure uniformity in input data and help the model focus on learning clinically relevant patterns rather than technical artifacts.
To localize the region of interest, particularly the thoracic cavity, an Attention U-Net is employed as the segmentation module. This architecture enhances the conventional U-Net by integrating attention gates, which dynamically suppress irrelevant background structures and highlight lung regions during the segmentation process. By focusing exclusively on the most informative anatomical areas, the model improves its ability to detect subtle disease markers and reduces the chances of false positives.24 This segmented output is then passed to the feature extraction module.
The next stage uses DenseNet-121, a densely connected convolutional neural network known for its efficient feature reuse and gradient flow. The dense connections between layers help the model learn complex radiographic features while minimizing redundancy. A spatial dropout layer with a dropout rate of 0.3 is applied after each dense block to reduce overfitting and enhance generalization, particularly when working with a moderately sized dataset. The rich feature maps generated by DenseNet capture fine-grained texture and shape variations within the thoracic region.
These extracted features are then fed into a Capsule Network (CapsNet), which excels at preserving the spatial relationships between features—an essential aspect in medical image analysis. Unlike traditional CNNs, CapsNet models spatial hierarchies and is robust to changes in orientation, scale, and viewpoint. This property is particularly valuable in chest X-rays, where variations in patient positioning and anatomy are common. The capsule layers capture pose information of the extracted features, and dynamic routing between capsules ensures that relevant features are grouped and classified effectively. The final output capsule predicts the probability distribution across four diagnostic categories: COVID-19, pneumonia, tuberculosis, and normal.
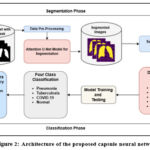 |
Figure 2: Architecture of the proposed capsule neural network |
By integrating segmentation, DenseNet-based feature extraction, and Capsule Networks for classification, this framework presents a novel and effective approach to automated thoracic disease detection. The combination of these advanced techniques offers a promising pathway for improving radiographic analysis, ultimately aiding in early diagnosis and better patient outcomes.25 Future research can further refine this framework by optimizing its architecture and expanding its applicability to a broader spectrum of thoracic diseases.
Data Pre-processing
The datasets utilized in this study contained chest X-ray images with varying resolutions, which posed a challenge for deep learning model consistency. Since the Capsule Network (CapsNet) models used in this work require images of a fixed dimension (224×224 pixels), all images were resized to meet this standard. This resizing step ensured uniformity across the dataset, 26 addressing inconsistencies in resolution while maintaining essential image details for effective feature extraction.
Despite resizing, a significant challenge remained the limited volume of available data. To address this issue and enhance the model’s ability to generalize effectively, data augmentation techniques were employed. Data augmentation artificially increases the dataset size by introducing slight variations to existing images, generating additional training samples without requiring new actual images.27 These variations help improve model robustness and prevent overfitting by exposing the network to diverse representations of the same underlying features.
Variations in image resolution, size, and quality arise due to differences in scanning equipment and acquisition methodologies. Such inconsistencies can affect network performance, potentially leading to biased model learning. To mitigate the impact of data heterogeneity, this study implemented several preprocessing techniques tailored to chest X-ray images. The preprocessing pipeline begins with converting the original chest X-ray images to grayscale, which simplifies the image by removing redundant color information. Following grayscale conversion, histogram equalization is applied to enhance local contrast, revealing crucial features that may otherwise remain indistinct. This step improves the visibility of abnormalities, making disease patterns more discernible for both human and machine interpretation.
Once the images undergo contrast enhancement, data augmentation techniques are applied to further diversify the training dataset. To maximize the unpredictability of augmentation and strengthen the model’s ability to generalize, 28 several augmentation strategies are employed. These include random rotation, vertical flipping, and random cropping. Random rotation slightly alters the image orientation, preventing the model from becoming overly reliant on specific directional patterns. Vertical flipping introduces additional variability by mirroring the images, simulating different viewing angles that might naturally occur in real-world clinical scenarios. Random cropping selectively removes small portions of the image, encouraging the network to learn robust features without relying on specific image regions.
Additionally, standardization is performed to expedite model convergence and enhance generalization. Standardization involves normalizing pixel intensity values across images, ensuring that the model learns patterns based on meaningful variations rather than being influenced by differences in absolute intensity values. By applying these preprocessing and augmentation techniques, the dataset becomes more diverse and representative, ultimately improving the model’s ability to detect thoracic diseases accurately.
Incorporating these steps into the data preparation pipeline ensures that the deep learning model is trained on high-quality, diverse, and representative samples, thereby enhancing diagnostic performance. The combination of resizing, grayscale conversion, contrast enhancement, augmentation, and standardization plays a crucial role in improving the network’s ability to identify thoracic abnormalities in chest X-ray images.
Deep learning approaches
A popular deep learning method called transfer learning uses pre-trained weights from enormous imagine datasets to categorize new photos, doing away with the need to create models from scratch. In transfer learning, data extraction and fine-tuning are the two main strategies. Feature extraction entails removing the top levels of classification from the new dataset and using previously trained models to extract pertinent features.29 A new classifier is then trained using the extracted features, allowing the pre-trained model to contribute useful representations without including its original classifier.
Fine-tuning, on the other hand, takes advantage of the pre-trained model’s weights by updating and modifying them during training. Unlike feature extraction, fine-tuning adjusts the model’s parameters to align with the new dataset, 30 enabling it to generate dataset-specific features rather than generic feature maps. This process ensures that general learned features are adapted to the specific classification task, rather than replacing them entirely.
In this study, the final layers of five state-of-the-art deep learning models—VGG-16, VGG-19, GoogLeNet, ResNet-50, and CapsNet—were optimized by employing transfer learning techniques. The pre-trained models were used as feature extractors, and their original classification layers were replaced with a new structure designed for multiclass classification. Instead of fully connected layers, a flattening layer was introduced to transform data into a one-dimensional tensor,31 followed by a softmax activation function for classification.
To enhance the performance and generalization of these models, a dropout rate of 0.5 was implemented to regulate the learning process and reduce overfitting. Additionally, a dense layer was incorporated to activate previous layers, ensuring improved feature representation. By employing transfer learning, the models benefited from the pre-trained knowledge while being fine-tuned to effectively classify thoracic diseases in chest X-ray images.
GoogLeNet was included for its efficient multi-scale feature extraction and low computational cost, making it suitable for limited medical datasets. Alongside VGG and ResNet, it offers architectural diversity. Although ensemble methods were considered for boosting performance, they were not used due to increased complexity and inference time in clinical settings.
This hybrid approach of utilizing both feature extraction and fine-tuning allowed the models to retain crucial pre-learned features while adapting to the specific task of thoracic disease classification. By integrating transfer learning techniques with deep learning architectures, this study successfully enhanced model accuracy, making them more robust for medical image analysis.
Attention U-net Model for Segmentation
The Attention U-Net model is an enhanced version of the conventional U-Net architecture, specifically designed for semantic segmentation tasks. This model improves segmentation accuracy by integrating an attention mechanism that enables the network to focus on the most significant regions of an image, enhancing its overall precision.
Similar to the standard U-Net, the Attention U-Net comprises an encoder and a decoder. The encoder is responsible for extracting essential features from the input image through multiple convolutional layers, effectively capturing hierarchical spatial information. The decoder, consisting of several deconvolutional layers, 32 reconstructs the segmentation map by progressively up-sampling the extracted feature maps.
The primary distinction of the Attention U-Net lies in the attention mechanism embedded within the decoder. This mechanism selectively highlights critical areas in the feature maps generated by the encoder, allowing the model to prioritize relevant regions while suppressing less important background details. By dynamically adjusting the focus based on contextual information, the attention module enhances the model’s ability to distinguish between key structures in an image, leading to improved segmentation outcomes.
This advanced architecture is particularly beneficial in medical image analysis, where precise localization of anatomical structures and abnormalities is crucial. By integrating attention mechanisms, the Attention U-Net provides a more robust and accurate segmentation framework, making it an effective tool for various medical imaging applications.
VGG-16 Model
VGG-16 is a deep convolutional neural network (CNN) architecture designed for image classification. It consists of 16 weight layers: 13 convolutional layers with 3×3 filters, followed by ReLU activations, and 3 fully connected (FC) layers. The network includes five max-pooling layers (2×2) after convolutional blocks33 to reduce spatial dimensions while preserving key features. The final FC layers output 1,000 class probabilities (for ImageNet). VGG-16 is known for its uniform architecture, deep feature extraction capability, and high computational cost. Despite being parameter-heavy, it serves as a strong backbone for tasks like object detection, segmentation, and medical image analysis.
VGG-19 Model
One deep convolutional neural network (CNN) that is well-known for its consistent design and potent feature extraction powers is VGG-19. Sixteen convolutional layers with 3×3 filters, five max-pooling layers (2×2) for downsampling, and three fully connected (FC) layers at the end make up its 19 layers. To improve non-linearity, 34 ReLU activation comes after each convolutional layer. The final output layer uses softmax for classification. Designed for large-scale image recognition (e.g., ImageNet), VGG-19 captures complex patterns but is computationally expensive. It is widely used in transfer learning, medical imaging, and object detection due to its deep hierarchical feature representation.
GoogLeNet(Inception) Model
Google devised the deep convolutional neural network architecture known as GoogLeNet, or the Inception network. The Inception module, which effectively grabs attributes at various spatial scales, is its main invention. An input layer, stem network, Inception modules, auxiliary classifiers, global average pooling, and softmax output constitute the architecture. 35 The Inception module employs parallel convolutional operations of different filter sizes and dimensionality reduction techniques to improve computational efficiency. To generate the output, the outcomes of these procedures are assembled along the depth dimension.
ResNet Model
ResNet is a complex convolutional neural network design created by Kaiming and his team 36 at Microsoft Research in 2015. The problem of vanishing gradients in training deep neural networks is addressed by using skip connections or shortcut connections. ResNet uses residual blocks to learn residual functions, making it easier to optimize deeper networks and preventing degradation. The architecture consists of an input layer, convolutional layers, and residual blocks, skip connections, stacking blocks, global average pooling, and a fully connected layer and softmax output.37 The main novelty of ResNet is the use of skip connections, which permit the immediate integration of one layer’s output to a deeper layer’s output.
Capsule Neural Network
A Capsule Neural Network (CapsNet) is a deep learning architecture that enhances traditional CNNs by preserving spatial hierarchies through capsules—groups of neurons that encode both feature presence and orientation. The architecture consists of a convolutional layer for feature extraction, a Primary Capsule layer that converts features into capsules, and a Digit Capsule layer that uses dynamic routing to determine spatial relationships.38 Instead of max pooling, CapsNet uses squashing functions to retain detailed spatial information. The margin loss function ensures robust classification. CapsNet improves viewpoint invariance, making it highly effective for medical imaging, object recognition, and small dataset learning.
Results
Using a 16-GB NVIDIA GeForce GTX GPU, the deep learning algorithms VGG-16, VGG-19, InceptionNet, ResNet50, and CapsNet were trained for optimum computational efficiency. In advance of training, all images were resized to an uniform 224 × 224 pixel size 39 to guarantee homogeneity across the dataset.
For model training, the categorical cross-entropy loss function was employed to compare the predicted outputs with the ground truth labels. To optimize model performance and minimize the loss function, the Adam optimizer was utilized with a learning rate of 0.001. Additionally, early stopping based on validation performance was implemented to prevent both overfitting and underfitting.40 If no improvement in validation loss was observed after 40 consecutive epochs, the training process was halted.
Throughout the training phase, multiple models were developed, and their respective accuracies and losses were recorded and visualized using graphical representations.41 After training, the test accuracy of each model was assessed and compared against previously established studies on CNN-based lung disease diagnosis. This comparative analysis provided insights into the effectiveness of the trained models and their potential for improving lung disease detection in medical imaging applications.
Table 1 presents the accuracy and loss values for training and validation, while Figures 3-7 illustrate these metrics for each optimized model. Among the trained models, GoogLeNet and ResNet demonstrated the fastest convergence, achieving over 98% validation accuracy within 40 epochs. DenseNet and CapsNet exhibited superior training accuracy with minimal loss, indicating their robustness in feature extraction and classification.42 While CapsNet achieved the lowest validation loss, AlexNet, VGGNet, and DenseNet displayed higher validation accuracy.
Table 1: Accuracy and loss in training and validation
| Model | Training Loss | Training Accuracy | Validation Loss | Validation Accuracy |
| VGG-16 | 0.0110 | 0.9980 | 0.0000e+00 | 1.0000 |
| VGG-19 | 0.0121 | 0.9992 | 0.0000e+00 | 1.000 |
| Inception Net | 0.0023 | 0.9992 | 0.0012 | 1.0000 |
| ResNet-50 | 1.1179e04 | 1.0000 | 0.0629 | 0.9977 |
| CapsNet | 6.1362e−09 | 1.0000 | 7.2102e−07 | 1.0000 |
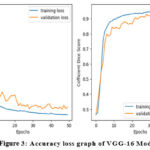 |
Figure 3: Accuracy loss graph of VGG-16 Model |
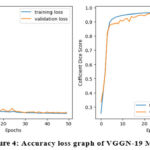 |
Figure 4: Accuracy loss graph of VGGN-19 Model |
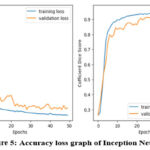 |
Figure 5: Accuracy loss graph of Inception Net Model |
Following the completion of training, the models underwent testing on a separate test set to determine their level of accuracy. F1-score, recall (sensitivity), precision, and classification accuracy 43 served as indicators of the models’ performance. A confusion matrix, along with Scikit-Learn’s metrics function, 44was employed to generate a classification report by assessing the number of correctly identified images.
This comprehensive evaluation provided insights into model performance across various disease categories as shown in Table 2.
Table 2: Confusion Matrix.
| Model | True Positive | True Negative | False Positive | False Negative |
| VGG-16 | 624 | 82 | 26 | 18 |
| VGG-19 | 620 | 80 | 30 | 20 |
| Inception Net | 620 | 82 | 27 | 21 |
| ResNet 50 | 620 | 81 | 28 | 21 |
| CapsNet | 625 | 83 | 24 | 18 |
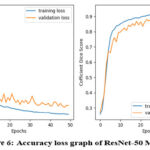 |
Figure 6: Accuracy loss graph of ResNet-50 Model |
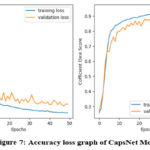 |
Figure 7: Accuracy loss graph of CapsNet Model |
For training the segmentation model, input images were paired with corresponding ground truth segmentation masks, forming labeled training data for the Fully Convolutional Network (FCN) model. The model’s weights were updated using Adam optimization, employing pixel-wise loss functions such as binary cross-entropy or dice loss to enhance segmentation accuracy.
The Receiver Operating Characteristic (ROC) curve, a widely used statistical tool in radiology for evaluating detection limits and screening performance, is presented in Figures 8-11. This curve provides a visual representation of the model’s diagnostic ability at various threshold settings.
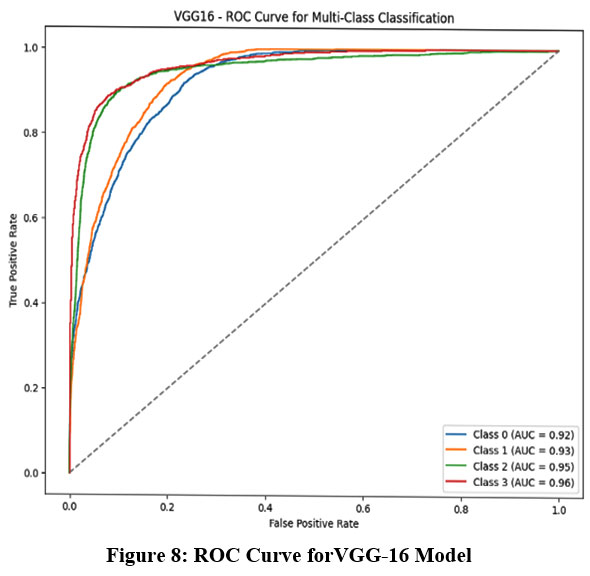 |
Figure 8: ROC Curve forVGG-16 Model |
 |
Figure 9: ROC Curve forVGG-19 Model |
Discussion
Based on these findings, it can be concluded that among the seven evaluated models, CapsNet exhibited the most consistent and superior performance during both training and validation, making it a promising approach for chest X-ray image classification.45
The study evaluated the performance of pre-trained deep learning models based on key metrics such as accuracy, precision, recall, and F1 score.46 The results, summarized in Table 3, indicate that CapsNet achieved the highest accuracy at 98.48%, outperforming DenseNet, which attained an accuracy of 98.16%.
Table 3 also provides a comparison of computational time for training and testing each model. Among the models tested, DenseNet demonstrated the shortest training duration, completing the process in 11 minutes and 50 seconds. However, it had the longest testing time, requiring 16 minutes and 14 seconds to complete. In contrast, CapsNet, while taking a longer time to train (20 minutes), demonstrated superior efficiency during testing, completing the task in 15 minutes and 36 seconds.
Table 3: Performance Matrix
| Model | Accuracy | Precision | Recall | F1Score | Training Time (S) | Testing Time (S) |
| VGG-16 | 97.18 | 97.34 | 97.18 | 97.22 | 1344 | 1946 |
| VGG-19 | 97.21 | 97.79 | 97.21 | 97.38 | 899 | 1083 |
| Inception Net | 98.16 | 98.57 | 98.96 | 98.02 | 710 | 974 |
| ResNet 50 | 97.32 | 97.42 | 97.32 | 97.35 | 783 | 1056 |
| CapsNet | 98.48 | 98.54 | 98.48 | 98.49 | 1200 | 936 |
 |
Figure 10: ROC Curve for InceptionV3 Model |
 |
Figure 11: ROC Curve for Resnet-50 Model |
Based on these results, CapsNet is recommended for the classification of various thoracic diseases in chest X-ray images. It achieved the highest accuracy (98.48%) and demonstrated robust performance across other evaluation metrics; including precision (98.54%), recall (98.48%), and F1 score (98.49%). These findings highlight the model’s reliability and effectiveness in thoracic disease diagnosis.
CapsNet outperforms others by preserving spatial hierarchies, making it effective in detecting subtle and positionally varied thoracic abnormalities—crucial in clinical settings. Computationally, it offers efficient inference. However, real-world deployment may face challenges like data heterogeneity, patient movement, and imaging artifacts, requiring robust preprocessing and domain-specific model adaptation.
Conclusion
The hybrid deep learning model described in this work uses chest X-ray pictures to diagnose thoracic disorders by integrating a Convolutional Neural Network (CNN), namely VGG-16, with a Capsule Network (CapsNet). To conquer the difficulties caused by imbalanced and small datasets, data augmentation methods were used, which enhanced the model’s capacity for generalization. With a 98.48% accuracy rate and a 98.49% F1-score, the improved CapsNet architecture surpassed the other models in the evaluation. These results highlight the model’s effectiveness in identifying and classifying thoracic diseases with greater precision compared to other deep learning models. Statistical evaluations confirm that the proposed CapsNet-based model surpasses previous classification techniques, offering a more robust and reliable solution for thoracic disease detection. This advancement paves the way for improved diagnostic tools in clinical settings.
Further research will focus on utilizing chest X-ray images of infected patients to develop more precise diagnostic tools for detecting multiple thoracic conditions. Additionally, to make disease detection more accessible, a mobile-based application will be developed in the future. This application will allow individuals to perform preliminary disease screening using their smartphones or web-based platforms, providing a convenient and user-friendly diagnostic tool. By leveraging artificial intelligence and deep learning models, this innovation aims to assist both medical professionals and the public in early disease detection and self-assessment. The integration of such technology into mobile healthcare applications has the potential to revolutionize disease diagnosis, making it more efficient and widely accessible.
Acknowledgement
The author would like to thank Graphic Era Hill University for providing the necessary resources, facilities, and a conducive environment for completing the research work.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability
This statement does not apply to this article.
Ethics statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable
Authors’ Contribution
- Himanshu Pant: Conceived and designed the research study, developed the methodology, Writing the original Draft;
- Garima Joshi: Data Collection, conducted data preprocessing, assisted with visualization; Data Preprocessing, Results interpretation; Data Analysis, Domain Expert;
- Bhupesh Rawat: Worked on deep learning and machine learning models and assisted in fine-tuning the algorithms.
References
- Al-qaness MAA, Zhu J, AL-Alimi D, et al. Chest X-ray Images for Lung Disease Detection Using Deep Learning Techniques: A Comprehensive Survey. Archives of Computational Methods in Engineering. 2024;31(6):3267-3301. doi:10.1007/s11831-024-10081-y
CrossRef - Agrawal T, Choudhary P. ALCNN: Attention based lightweight convolutional neural network for pneumothorax detection in chest X-rays. Biomed Signal Process Control. 2023;79:104126. doi:10.1016/j.bspc.2022.104126
CrossRef - Sultana S, Pramanik A, Rahman MdS. Lung Disease Classification Using Deep Learning Models from Chest X-ray Images. In: 2023 3rd International Conference on Intelligent Communication and Computational Techniques (ICCT). IEEE; 2023:1-7. doi:10.1109/ICCT56969.2023.10075968
CrossRef - Hameed RA. The Role of artificial intelligence in early detection of lung cancer using chest X-rays. International Journal of Radiology and Diagnostic Imaging. 2024;7(4):33-38. doi:10.33545/26644436.2024.v7.i4a.421
CrossRef - Stamate E, Piraianu AI, Ciobotaru OR, et al. Revolutionizing Cardiology through Artificial Intelligence—Big Data from Proactive Prevention to Precise Diagnostics and Cutting-Edge Treatment—A Comprehensive Review of the Past 5 Years. Diagnostics. 2024;14(11):1103. doi:10.3390/diagnostics14111103
CrossRef - Sufian MA, Hamzi W, Sharifi T, et al. AI-Driven Thoracic X-ray Diagnostics: Transformative Transfer Learning for Clinical Validation in Pulmonary Radiography. J Pers Med. 2024;14(8):856. doi:10.3390/jpm14080856
CrossRef - Alaufi R, Kalkatawi M, Abukhodair F. Challenges of deep learning diagnosis for COVID-19 from chest imaging. Multimed Tools Appl. 2023;83(5):14337-14361. doi:10.1007/s11042-023-16017-1
CrossRef - Choudhry IA, Iqbal S, Alhussein M, Qureshi AN, Aurangzeb K, Naqvi RA. Transforming Lung Disease Diagnosis With Transfer Learning Using Chest X‐Ray Images on Cloud Computing. Expert Syst. 2025;42(2). doi:10.1111/exsy.13750
CrossRef - Nazir A, Hussain A, Singh M, Assad A. Deep learning in medicine: advancing healthcare with intelligent solutions and the future of holography imaging in early diagnosis. Multimed Tools Appl. 2024;84(17):17677-17740. doi:10.1007/s11042-024-19694-8
CrossRef - Asif S, Wenhui Y, ur-Rehman S, et al. Advancements and Prospects of Machine Learning in Medical Diagnostics: Unveiling the Future of Diagnostic Precision. Archives of Computational Methods in Engineering. 2025;32(2):853-883. doi:10.1007/s11831-024-10148-w
CrossRef - Chen G, Li C, Wei W, et al. Fully Convolutional Neural Network with Augmented Atrous Spatial Pyramid Pool and Fully Connected Fusion Path for High Resolution Remote Sensing Image Segmentation. Applied Sciences. 2019;9(9):1816. doi:10.3390/app9091816
CrossRef - Kansal K, Chandra TB, Singh A. Advancing differential diagnosis: a comprehensive review of deep learning approaches for differentiating tuberculosis, pneumonia, and COVID-19. Multimed Tools Appl. 2024;84(13):11871-11906. doi:10.1007/s11042-024-19350-1
CrossRef - Exarchos KP, Gkrepi G, Kostikas K, Gogali A. Recent Advances of Artificial Intelligence Applications in Interstitial Lung Diseases. Diagnostics. 2023;13(13):2303. doi:10.3390/diagnostics13132303
CrossRef - Albahli S, Rauf HT, Arif M, Nafis MT, Algosaibi A. Identification of Thoracic Diseases by Exploiting Deep Neural Networks. Computers, Materials & Continua. 2021;66(3):3139-3149. doi:10.32604/cmc.2021.014134
CrossRef - Ramalingam R, Chinnaiyan V. Intelligent optimization-based pulmonary emphysema detection with adaptive multi-scale dilation assisted residual network with Bi-LSTM layer. Biomed Signal Process Control. 2024;88:105643. doi:10.1016/j.bspc.2023.105643
CrossRef - Appavu NK, C NKB, Kadry S. COVID-19 classification in X-ray/CT images using pretrained deep learning schemes. Multimed Tools Appl. 2024;83(35):83157-83177. doi:10.1007/s11042-024-18721-y
CrossRef - Constantinou M, Exarchos T, Vrahatis AG, Vlamos P. COVID-19 Classification on Chest X-ray Images Using Deep Learning Methods. Int J Environ Res Public Health. 2023;20(3):2035. doi:10.3390/ijerph20032035
CrossRef - Chen W, Zhou Z, Bao J, et al. Classifying Heart-Sound Signals Based on CNN Trained on MelSpectrum and Log-MelSpectrum Features. Bioengineering. 2023;10(6):645. doi:10.3390/bioengineering10060645
CrossRef - Thanoon MA, Zulkifley MA, Zainuri MAAM, Abdani SR. A Review of Deep Learning Techniques for Lung Cancer Screening and Diagnosis Based on CT Images. Diagnostics. 2023;13(16):2617. doi:10.3390/diagnostics13162617
CrossRef - Nasser AA, Akhloufi MA. A Review of Recent Advances in Deep Learning Models for Chest Disease Detection Using Radiography. Diagnostics. 2023;13(1):159. doi:10.3390/diagnostics13010159
CrossRef - Nafisah SI, Muhammad G. Tuberculosis detection in chest radiograph using convolutional neural network architecture and explainable artificial intelligence. Neural Comput Appl. 2024;36(1):111-131. doi:10.1007/s00521-022-07258-6
CrossRef - Wang X, Peng Y, Lu L, Lu Z, Bagheri M, Summers RM. ChestX-Ray8: Hospital-Scale Chest X-Ray Database and Benchmarks on Weakly-Supervised Classification and Localization of Common Thorax Diseases. In: 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). IEEE; 2017:3462-3471. doi:10.1109/CVPR.2017.369
CrossRef - Pant H, Lohani MC, Bhatt AK. Attention U-Net Model for Accurate Thoracic Disease Detection, Segmentation and Localization. In: 2024 Fourth International Conference on Advances in Electrical, Computing, Communication and Sustainable Technologies (ICAECT). IEEE; 2024:1-7. doi:10.1109/ICAECT60202.2024.10469141
CrossRef - Jalloul M, Alkhulaifat D, Miranda-Schaeubinger M, Benedetti LDL, Otero HJ, Dako F. Artificial Intelligence in Chest Radiology: Advancements and Applications for Improved Global Health Outcomes. Curr Pulmonol Rep. 2024;13(1):1-9. doi:10.1007/s13665-023-00334-9
CrossRef - Kamal U, Zunaed M, Nizam NB, Hasan T. Anatomy-XNet: An Anatomy Aware Convolutional Neural Network for Thoracic Disease Classification in Chest X-Rays. IEEE J Biomed Health Inform. 2022;26(11):5518-5528. doi:10.1109/JBHI.2022.3199594
CrossRef - Kaur T, Gandhi TK. Automated Brain Image Classification Based on VGG-16 and Transfer Learning. In: 2019 International Conference on Information Technology (ICIT). IEEE; 2019:94-98. doi:10.1109/ICIT48102.2019.00023
CrossRef - Khan RU, Zhang X, Kumar R, Aboagye EO. Evaluating the Performance of ResNet Model Based on Image Recognition. In: Proceedings of the 2018 International Conference on Computing and Artificial Intelligence. ACM; 2018:86-90. doi:10.1145/3194452.3194461
CrossRef - Klangbunrueang R, Pookduang P, Chansanam W, Lunrasri T. AI-Powered Lung Cancer Detection: Assessing VGG16 and CNN Architectures for CT Scan Image Classification. Informatics. 2025;12(1):18. doi:10.3390/informatics12010018
CrossRef - Kumar S, Singh J, Ravi V, et al. Deep Learning and MRI Biomarkers for Precise Lung Cancer Cell Detection and Diagnosis. Open Bioinforma J. 2024;17(1). doi:10.2174/0118750362335415240909061539
CrossRef - Kordnoori S, Sabeti M, Mostafaei H, Banihashemi SSA. Front Cover: Advances in medical image analysis: A comprehensive survey of lung infection detection. IET Image Process. 2024;18(13). doi:10.1049/ipr2.13281
CrossRef - Lococo F, Ghaly G, Chiappetta M, et al. Implementation of Artificial Intelligence in Personalized Prognostic Assessment of Lung Cancer: A Narrative Review. Cancers (Basel). 2024;16(10):1832. doi:10.3390/cancers16101832
CrossRef - Pant H, Lohani MC, Bhatt AK. X-rays imaging analysis for early diagnosis of thoracic disorders using capsule neural network: a deep learning approach. International Journal of Advanced Technology and Engineering Exploration. 2023;10(104). doi:10.19101/IJATEE.2022.10100468
CrossRef - Pallumeera M, Giang JC, Singh R, Pracha NS, Makary MS. Evolving and Novel Applications of Artificial Intelligence in Cancer Imaging. Cancers (Basel). 2025;17(9):1510. doi:10.3390/cancers17091510
CrossRef - Praveena M, Ravi A, Srikanth T, Praveen BH, Krishna BS, Mallik AS. Lung Cancer Detection using Deep Learning Approach CNN. In: 2022 7th International Conference on Communication and Electronics Systems (ICCES). IEEE; 2022:1418-1423. doi:10.1109/ICCES54183.2022.9835794
CrossRef - Preuss K, Thach N, Liang X, et al. Using Quantitative Imaging for Personalized Medicine in Pancreatic Cancer: A Review of Radiomics and Deep Learning Applications. Cancers (Basel). 2022;14(7):1654. doi:10.3390/cancers14071654
CrossRef - Rehman A, Butt MA, Zaman M. A Survey of Medical Image Analysis Using Deep Learning Approaches. In: 2021 5th International Conference on Computing Methodologies and Communication (ICCMC). IEEE; 2021:1334-1342. doi:10.1109/ICCMC51019.2021.9418385
CrossRef - Saravanaprasad P, Karuppusamy SA. Advanced lung tumor diagnosis using a 3D deep neural network based CAD system. Biomed Signal Process Control. 2024;89:105650. doi:10.1016/j.bspc.2023.105650
CrossRef - Sharma S, Guleria K. A Deep Learning based model for the Detection of Pneumonia from Chest X-Ray Images using VGG-16 and Neural Networks. Procedia Comput Sci. 2023;218:357-366. doi:10.1016/j.procs.2023.01.018
CrossRef - Swarup C, Singh KU, Kumar A, Pandey SK, varshney N, Singh T. Brain tumor detection using CNN, AlexNet & GoogLeNet ensembling learning approaches. Electronic Research Archive. 2023;31(5):2900-2924. doi:10.3934/era.2023146
CrossRef - Sakthi U, Vaishnave K V, M AKA. A Comprehensive Multimodal Analysis for Detecting Pulmonary Infiltrates in Chest X-Ray Image. In: 2024 Third International Conference on Distributed Computing and Electrical Circuits and Electronics (ICDCECE). IEEE; 2024:1-5. doi:10.1109/ICDCECE60827.2024.10549653
CrossRef - Wu Z, Shen C, van den Hengel A. Wider or Deeper: Revisiting the ResNet Model for Visual Recognition. Pattern Recognit. 2019;90:119-133. doi:10.1016/j.patcog.2019.01.006
CrossRef - Yazar S. A Comparative study on classification performance of Emphysema with transfer learning methods in deep convolutional neural networks. Computer Science Journal of Moldova. 2022;30(2 (89)):259-278. doi:10.56415/csjm.v30.15
CrossRef - Zouch W, Sagga D, Echtioui A, et al. Detection of COVID-19 from CT and Chest X-ray Images Using Deep Learning Models. Ann Biomed Eng. 2022;50(7):825-835. doi:10.1007/s10439-022-02958-5
CrossRef - Nayak SR, Nayak DR, Sinha U, Arora V, Pachori RB. Application of deep learning techniques for detection of COVID-19 cases using chest X-ray images: A comprehensive study. Biomed Signal Process Control. 2021;64:102365. doi:10.1016/j.bspc.2020.102365
CrossRef - Kaur P, Harnal S, Tiwari R, et al. A Hybrid Convolutional Neural Network Model for Diagnosis of COVID-19 Using Chest X-ray Images. Int J Environ Res Public Health. 2021;18(22):12191. doi:10.3390/ijerph182212191
CrossRef - Hasan MdR, Ullah SMA, Islam SMdR. Recent advancement of deep learning techniques for pneumonia prediction from chest X-ray image. Medical Reports. 2024;7:100106. doi:10.1016/j.hmedic.2024.100106
CrossRef







