Manuscript accepted on :12-06-2025
Published online on: 24-06-2025
Plagiarism Check: Yes
Reviewed by: Dr. Arif Ansori
Second Review by: Dr. Satya Namani
Final Approval by: Dr. Anton R Keslav
Aisha Mohammed Abdallah 1,4 , Mutaz Fathelrhman Saad1*
, Mutaz Fathelrhman Saad1* , Hind Haidar Ahmed1
, Hind Haidar Ahmed1 , Tagwa Salah Ahmed2,3
, Tagwa Salah Ahmed2,3 Samar Mohammed Saeed1
Samar Mohammed Saeed1 and Wafaa Mohammed Abdalla1
and Wafaa Mohammed Abdalla1
1Department of Microbiology, College of Medical Laboratory Science, Sudan University of Science and Technology, Khartoum, Sudan.
2Department of Clinical Laboratory Sciences, College of Applied Medical Sciences, Jouf University, Saudi Arabia.
3Department of Parasitology, Medical Entomology, College of Medical Laboratory Science, Sudan University of Science and Technology, Khartoum – Sudan.
4Department of Microbiology, College of Medical Laboratory Science, National Ribat University, Khartoum, Sudan.
Corresponding Author E-mail: mutazsaad74@gmail.com
DOI : https://dx.doi.org/10.13005/bpj/3208
Abstract
Carbapenems are the most important β-lactam antibiotics against bacteria especially multidrug resistant isolates like Pseudomonas aeruginosa. This study was conducted to detect carbapenem resistance genes of P. aeruginosa from clinical samples in Khartoum State, Sudan. Fifty (n=50) isolates of p. aeruginosa were included in this descriptive cross-sectional laboratory-based study. P aeruginosa were isolated from urine, blood, wound swab, sputum and Body fluids samples, which were attained from different hospitals in Khartoum State. The samples were collected from both males and ladies with different age using non-self-constructing information list. The isolated organism was stored in 20% Glycerol peptone media and was inoculated on CLED agar. Antibiotic susceptibility tests were carried out using Kirby Bauer technique. DNA was extracted using boiling procedure. The isolates were tested for the presence of carbapenem resistance genes (NDM, IMP) using Multiplex PCR. Out of 50 P. aeruginosa isolates, 19 (38%) were positive for one or both carbapenem resistance genes; 6 (12%) were positive for NDM gene and 13(26%) were positive for IMP gene. The study concluded that, there was a high percentage and multifariousness of carbapenem resistance genes among P. aeruginosa isolates in Khartoum State, Sudan.
Keywords
Carbapenem; IMP; Multiplex PCR; NDM genes; P. aeruginosa; Sudan
Download this article as:| Copy the following to cite this article: Abdallah A. M, Saad M. F, Ahmed H. H, Ahmed T. S, Saeed S. M, Abdalla W. M. Antimicrobial Susceptibility and Molecular Detection of Carbapenem Resistant Genes NDM-1 and IMP-1 of Pseudomonas aeruginosa Isolated from Clinical Samples in Khartoum State-Sudan. Biomed Pharmacol J 2025;18(2). |
| Copy the following to cite this URL: Abdallah A. M, Saad M. F, Ahmed H. H, Ahmed T. S, Saeed S. M, Abdalla W. M. Antimicrobial Susceptibility and Molecular Detection of Carbapenem Resistant Genes NDM-1 and IMP-1 of Pseudomonas aeruginosa Isolated from Clinical Samples in Khartoum State-Sudan. Biomed Pharmacol J 2025;18(2). Available from: https://bit.ly/44jw6gs |
Introduction
P. eruginosa, one of the most common opportunistic pathogen associated with nosocomial infections, including pneumonia, urinary tract infections, and wound infections. Although carbapenems are frequently used as a therapeutic agent for treating infections caused by P. aeruginosa, the high emergence of carbapenem resistance significantly decreases their utility.1 Resistance of Pseudomonas aeruginosa to different classes of antibiotics such as penicillin, cephalosporin, quinolone, aminoglycoside and carbapenem. Metallo beta lactamases (MBL) are enzymes that catalyze the hydrolysis of broad-spectrum beta lactam antibiotics including carbapenems. Resistant to carbapenems especially in Gram negative pathogens is global public health issue because of the spread of transferable carbapenemase encoding genes. The antibiotic resistant elements could be transferred to other Gram-negative bacterial strains and contribute towards spread of antimicrobial resistance rate making the treatment of infected cases complex. 2 Carbapenems are the antibiotics which are utilized for treating multidrug-resistant P.aeruginosa (MDRP) isolates. Carbapenem antibiotics used to be effective agents against MDRP when first presented. However, the growing frequence of carbapenem resistant P. aeruginosa (CRPA) has turned into a severe health problem recently. These strains lead to high mortality rates in cases infected by P. aeruginosa and there are also few effective drugs against them. Resistance to carbapenems can be related to the production of carbapenemase enzymes such as serine carbapenemases (containing KPC and GES enzymes), metallo-β-lactamases (MBLs) such as IMP, VIM, and NDM enzymes, and oxacillinases (such as OXA enzymes). blaIMP and blaVIM are the most constantly acquired MBLs. However, the recently emerged NDM type (New Delhi metallo-β lactamases) is getting the most important carbapenemase. 3
Carbapenems have become the drug of choice for the treatment of severe nosocomial infections caused by Gram-negative bacilli. Carbapenem resistant Enterobacteriales (CRE) is a considerable health problem globally and are associated with increased mortality also There is a significant challenge in controlling the spread of carbapenemases, Surveys of the molecular epidemiology of carbapenemases revealed that the dissemination of carbapenemases, including NDM, VIM, IMP, OXA-48 and KPC producers, are rapid and widespread among healthcare facilities. Molecular techniques remain the gold standard for the precise identification of carbapenemase genes. 4 Most of these techniques are based on PCR and may be followed by a sequencing step if a precise identification of the carbapenemase gene is needed (e.g. VIM type, KPC type, NDM type or OXA-48 type). 4 They are either single or multiplex PCR techniques.4
In Africa, data on the frequence and distribution of carbapenem resistance among the multidrug resistant Gram-negative bacteria is still limited.5 In addition, detection discovery of carbapenem resistance genes producing bacteria may be difficult based on routine antibiotic susceptibility testing.6 Considering the fact that there were no comprehensive studies about prevalence of carbapenem resistance genes in Sudan so detection of those genes in P. aeruginosa from clinical samples will be of a great value. Such data serve an important part in understanding the spread of this pathogen and help in good treatment for it. So this study aimed to detect carbapenem resistance genes of Pseudomonas aeruginosa from clinical samples in Khartoum State, using conventional and PCR based method.
Materials and Methods
Study sample and data collection
A total of fifty non-repetitive P.aeruginosa (n=50), isolated from different clinical specimens
Data included sample type, gender, and age of patients were collected from hospitals’ medical records checklist.
Isolation and identification of bacteria
One colony from each isolate was sub-cultured on Cysteine Lactose Electrolyte Deficient media (CLED) and Nutrient agar and incubated aerobically at 37°C overnight, for further Re-identification and preservation. P.aeruginosa showed smooth blue non-lactose fermenting colonies on CLED, were identified by Gram stain and conventional biochemical tests.
Antimicrobial Susceptibility test
All confirmed isolates were subculture from 15% glycerol on nutrient agar getting overnight and pure growth. Modified Kirby Bauer was performed and interpreted according to the Clinical and Laboratory Standards Institute (CLSI, 2022).
DNA Extraction
The boiling method was used for isolation of bacterial genomic DNA which extracted from fresh overnight incubated isolate that suspended in 200ul of d H2O in 1.5 eppendrof tube, bolied for 10 min at 100°C in thermal block incubator, vortex, then cooling at -20°C following by centrifuged for 10 min at 12000xg. Supernatant were carefully collected and store at -20°C until analyzed. The purity of the extracted DNA was determined by running the DNA sample on 1.5% agarose gel.
Polymerase Chain Reaction Amplification
PCR was done by multiplex PCR, amplification which done using Maxime PCR Premix kit and specific primer which sensitized by Specific primers were used to amplification indicated in table (1). The PCR assay was carried out in a total volume of 25 μL of mixture containing 2 μLMaxime PCR Premix containing 1X PCR buffer, 1.5 mM MgCl2, 200M μ of each dNTP, and 1 U Taq DNA polymerase, 0.5 μL of each of the virulence gene-specific primers (2 μL), 5 μL of template DNA and 13 μL of water for injection. The amplification conditions included three steps: heating at 95°C for 5 min; 34 cycles of denaturation at 95°C for 1 min, annealing at 58°C for 1min for (NDM -1 and IMP -1 genes) and extension at 72°C for 1min; and the final extension at 72°C for 5 min. 7
Table 1: Primers used for amplification of carbapenem resistance genes of P.aeruginosa isolates
| Target gene | Sequence (5′- 3′) | Amplicon size (bp) |
| IMP-F | GGAATAGAGTGGCTTAAYTC |
Bla IMP -1 232 |
| IMP-R | GGTTTAAYAAAACAACCACC | |
| NDM-F | GGTTTGGCGATCTGGTTTTC |
Bla NDM -1621 |
| NDM-R | CGGAATGGCTCATCACGATC |
Gel electrophoresis
The gel casting tray was put into the electrophoresis, tank flooded with 1x TBE buffer just to cover the gel surface, 2μl of PCR products from each 50samples was added to wells of electrophoreses, 5 μl of100-bp DNA ladder, was added to the well in each run. The electrophoresis was carried out at 120Volts for 2 Hours. Then the gel was visualized for DNA bands by U.V transilluminator and photographed.
Statistical analysis
All outcome data were analyzed by chi-square using Statistical Package for Social Sciences (SPSS; Version16).
Results
Out of the 50 isolates, 18 were isolated from urine samples, 11 from wound swabs, 6 from blood, 2 from high vaginal swabs, 8 from sputum, 3 from ear swabs, and 2 from body fluids, after culturing most strains were show pigmentation 32 (74%), the most pigmented strain was found in urine (26%) and wound samples (20%) table (2). Demographic data show out of 50 the male 23 were (46%),where female were 27 (54%) with ratio1 : 0.85. Ages ranged from 4 to 76 years, with an average of 49. Majority of the samples (37%) were found in the age group (40 – 60) year table (3).
Approximately one-third 18 (36%) of isolates were resistant to Aztreonam, 10 (20%) were resistant to Ciprofloxacine and Colistin, 11 (22%) resistant to Gentamicin, Amikacin, and Pipracillin/tazobactam combinations, only one isolate was resistant to Imipenem, from 18 urine samples 3 (19%) isolates were resistant to Norfloxacin table (4).
Out of 50 P. aeruginosa isolates; only 14 (28%) were MDR according to the preliminarily described definition , most isolates were sensitive to all antibiotics 10 (36%), 12 (24%) isolates resist to one antibiotic, 6 (12%) isolates resist to two antibiotics. In MDR isolates we found two strains were resistant to >7 antibiotics, whereas just three isolates were found to be resistant to 6 antibiotics. There was insignificant Association between sex and antimicrobial susceptibility results table (5). The NDM-1gene was detected in 6 isolates (12%) and negative in 44 isolate (88%). They were recovered from urine (8%), wound (4%), ear swab (0%), sputum (0%), blood (0%) high vaginal swabs (0%) and body fluid (0%) table (6). While the IMP-1 gene was detected in 13(26%), and negative in 37(74%) isolate table (6). There was insignificant difference in the percentage of resistant genes among the different infection site. The association between the presence of NDM-1 and IMP-1genes and antibiotics was significant with Pipracillin, Impenem, Aztreonam, Ceftazidime table (7).
Table 2: Frequency of pigmented strains among different clinical samples
| Pigment appearance | |||
| Green Pigment | Non pigmented | ||
|
Sample |
Urine sample | 13 (26%) | 5 (10%) |
| Sputum | 7 (14%) | 1 (2%) | |
| Wound | 10 (20%) | 1 (2%) | |
| Blood | 4 (8%) | 2 (4%) | |
| Ear swab | 1 (2%) | 2 (4%) | |
| High vaginal swab | 2 (4%) | 0 (0%) | |
| Body fluids | 1 (2%) | 1 2%) | |
Table 3: Distribution of positive samples among different gender and age groups
| Gender | ||||
| Male | Female | Total | ||
| Age group | <20 | 1 (2%) | 0 (0%) | 1(2%) |
| 20-40 | 3 (6%) | 10 (20%) | 13 (26%) | |
| 40-60 | 10 (20%) | 10 (20%) | 20 (40%) | |
| >60 | 9 (18%) | 7 (14%) | 16 (32%) | |
Table 4 : Antimicrobial susceptibility pattern of P . aeruginosa isolates
| Sample | Urine(N=18) | Wound swab(n=11) | Blood(N=6) | HVS(N=2) | Sputum(N=8) | Ear swab(N=3) | Body fluid(N=2) | Total(N=50) | |
| Antibiotic | |||||||||
| CIP | Resistant | 3 | 3 | 2 | 0 | 1 | 2 | 0 | 11(22%) |
| Sensitive | 15 | 8 | 4 | 2 | 7 | 1 | 2 | 39(78%) | |
| CN | Resistant | 4 | 5 | 2 | 0 | 2 | 1 | 1 | 15(30%) |
| Sensitive | 14 | 6 | 4 | 2 | 6 | 2 | 1 | 35(70%) | |
| TPZ | Resistant | 5 | 3 | 0 | 0 | 1 | 2 | 0 | 11(22%) |
| Sensitive | 13 | 8 | 6 | 2 | 7 | 1 | 2 | 39(78%) | |
| CAZ | Resistant | 6 | 4 | 3 | 0 | 2 | 1 | 1 | 17(34%) |
| Sensitive | 12 | 7 | 3 | 2 | 6 | 2 | 1 | 33(66%) | |
| ATM | Resistant | 12 | 4 | 3 | 1 | 7 | 1 | 1 | 30(60%) |
| Sensitive | 6 | 7 | 3 | 1 | 1 | 2 | 1 | 20(40%) | |
| AK | Resistant | 2 | 5 | 1 | 0 | 3 | 1 | 1 | 13(26%) |
| Sensitive | 16 | 6 | 5 | 2 | 5 | 2 | 1 | 37(74%) | |
| IMP | Resistant | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 2 (4%) |
| Sensitive | 16 | 11 | 6 | 2 | 8 | 3 | 2 | 48(96%) | |
| CT | Resistant | 5 | 1 | 0 | 1 | 2 | 1 | 0 | 10(20%) |
| Sensitive | 13 | 10 | 6 | 1 | 6 | 2 | 2 | 40(80%) | |
Table 5: Association between sex and Antimicrobial susceptibility results
| Antibiotic | Sex group | |||
| male | Female | |||
| Sensitive | Resistant | Sensitive | Resistant | |
| CAZ | 15 | 8 | 18 | 9 |
| AZT | 11 | 12 | 10 | 17 |
| TPZ | 17 | 6 | 22 | 5 |
| CT | 21 | 2 | 19 | 8 |
| CN | 13 | 10 | 22 | 5 |
Table 6: Association between P . aeruginosa Resistant genes and infection sites
| Resistant genes | Sample sites | |||||
| UrineN=18 | WoundN=11 | Ear swabN=3 | BloodN=6 | Body fluidN=2 | SputumN=8 | |
| NDM-1 | 4 | 2 | 0 | 0 | 0 | 0 |
| IMP-1 | 6 | 4 | 0 | 1 | 1 | 1 |
Table 7: Association between the presence of IMP-1gene and NDM -1 gene WITH antibiotics susceptibility
| Antibiotic | IMP -1 | NDM -1 | |||
| (+ve) | (-ve) | (+ve) | (-ve) | ||
| Ciprofloxacin | R | 5 | 12 | 2 | 15 |
| S | 8 | 25 | 4 | 29 | |
| Colistin | R | 3 | 10 | 3 | 10 |
| S | 10 | 27 | 3 | 34 | |
| Gentamicin | R | 4 | 18 | 2 | 20 |
| S | 9 | 19 | 4 | 24 | |
| Amikacin | R | 1 | 5 | 0 | 6 |
| S | 12 | 32 | 6 | 38 | |
| Pipracillin | R | 7 | 19 | 1 | 25 |
| S | 6 | 18 | 5 | 19 | |
| Impenem | R | 5 | 18 | 2 | 21 |
| S | 8 | 19 | 4 | 23 | |
| Aztreonam | R | 8 | 24 | 3 | 29 |
| S | 5 | 13 | 3 | 15 | |
| Ceftazidime | R | 10 | 26 | 3 | 33 |
| S | 3 | 11 | 3 | 11 | |
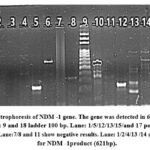 |
Figure 1: Gel electrophoresis of NDM -1 gene. The gene was detected in 6 isolates out of the 50 isolates. Lane: 9 and 18 ladder 100 bp. Lane: 1/5/12/13/15/and 17 positives for IMP -1 product (232bp). Lane:7/8 and 11 show negative results. Lane: 1/2/4/13 /14 and16 show positive for NDM -1product (621bp).Click here to view Figure |
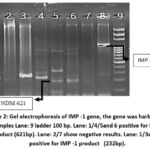 |
Figure 2: Gel electrophoresis of IMP -1 gene, the gene was harbored in13samples Lane: 9 ladder 100 bp. Lane: 1/4/5and 6 positive for NDM -1 product (621bp). Lane: 2/7 show negative results. Lane: 1/3and 8 positive for IMP -1 product (232bp).Click here to view Figure |
Discussion
In the current study, most of isolates harboring carbapenem resistance genes were phenotypically susceptible to imipenem and most of them were positive for IMP gene. This confirm what 8 said that this gene is not strong and relies upon other synergistic mechanisms to mediate resistance against carbapenems. In addition to imipenem other antibiotics were analyzed in this study. Majority of the isolates showed varying degrees of resistance to the other antibiotics (ciprofloxacin, gentamicin and ceftazidime). Resistance to these antibiotics may also be due to the presence of ESBLs and other mechanisms like efflux pumps and porin mutations 9, which were not covered in this study.
In the current study, the most prevalent gene among the 50P. aeruginosa isolates was IMP gene (26 %). NDM gene was also present but on a smaller scale (12%, n =6 ). This is consistent with previous findings by in Turkey10, which detected OXA-48 gene (58%) and NDM gene (2%) 11, detected OXA-48 gene (64.7%) and NDM gene (5.8%) and 12 detected OXA-48 gene (78%) and NDM gene (20%) both from Saudi Arabia, IMP gene were not detected in these studies
These variations in results may be because, they only detect resistant genes in carbapenem-resistant isolates or due to differences in method of detection as the first one used real-time PCR or differences in the sample size of tested organism and that was seen with the other two studies.
Of 50 P. aeruginosa isolates detected of having carbapenem resistance genes, two had multiple genes. This finding agree with 13 and 6 which showed multiplicity of genes in their isolates. The presence of multiple resistance genes in one isolate have not been generally detected in a large number of studies probably due to the limited number of genes studied since most of the studies research on one or two genes. 6 Regarding NDM and IMP gene, 6 (12%) and 13(26%) were positive respectively. Ceftazidime resistant Isolates showed significant association with both NDM and IMP gene as detected 3, 10 isolates respectively. One of the limitations of this study; it was conducted on a limited number of samples so future studies should be carried with large sample size and different clinical isolates to and sequencing technique to document the prevalence of these resistant genes and to help on therapeutic strategies.
Conclusion
In conclusion, the findings of the present study disclosed a high percentage andmultiplicity of carbapenem resistance genes among P. aeruginosa isolates in Khartoum State. This study showed that the NDM-1 and IMP-1 genes were generally circulated among the P. aeruginosa.
Acknowledgment
The authors would like to thank all the colleagues in Microbiology Laboratory, College of Medical Laboratory, Sudan University of sciences and technology who helped in the study, and also our thanks to Dr. Osama Mohamed a leader of EXON lab of molecular biology for his guidance and help in molecular work.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflect of Interest
The author(s) do not have any conflict of interest
Data Availability
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required
Clincal Trial Registeration
This research does not involve any clinical trials.
Permission to Reproduce Material from Other Source
Not Applicable
Authors Contributions
- Aisha Mohammed Abdallah: Methodology, and conducted the study
- Mutaz Fathelrhman Saad: Supervised the study, data analysis conceptualized, visualized, designed, interpretation of the results, and wrote the drafted paper.
- Hind Haidar Ahmed and Samar Mohammed Saeed: Data analysis and check the methodology, editing and approved the manuscript critically for remarkable intellectual contents.
- Tagwa Salah Ahmed and Wafaa Mohammed Abdalla: Data analysis and check the methodology, editing and approved the manuscript critically for remarkable intellectual contents.
References
- Shahin and Ahmadi. Molecular characterization of NDM-1-producing Pseudomonas aeruginosa isolates from hospitalized patients in Iran. Ann Clin Microbiol Antimicrob. 2021; 20:76 https://doi.org/10.1186/s12941-021-00482-3.
CrossRef - Ali A, Ahmad K, Rahat S, Ahmad I. Genetic diversity and molecular analysis of metallo beta lactamases among imipenem resistant clinical isolates of Pseudomonas aeruginosa from Peshawar, Pakistan. Pak J Med Sci. 2021;37(7): 1865-1870.doi: https://doi.org/10.12669/pjms.37.7.4303
CrossRef
- Farajzadeh Sheikh A, Shahin M, Shokoohizadeh L, et al.. Molecular epidemiology of colistin-resistant Pseudomonas aeruginosa producing NDM-1 from hospitalized patients in Iran. Iran J Basic Med Sci. 2019; 22:38-42. doi: 10.22038/ijbms.29264.7096.
- Nordmann, P. and Poirel, L. Strategies for identification of carbapenemase-producing Enterobacteriaceae. Journal of Antimicrobial Chemotherapy. 2012; 68(3), pp.487-489.
CrossRef - Mushi MF, Mshana SE, Imirzalioglu C, Bwanga F. Carbapenemase genes among multidrug resistant gram-negative clinical isolates from a tertiary Hospital in Mwanza, Tanzania. Biomed Res Int. 2014:3–8.
CrossRef
- Satir S., Elkhalifa A, Ali MA., et al. Detection of Carbepenem Resistance Genes among Selected Gram-Negative Bacteria Isolated from Patients in -Khartoum State, Sudan. Clinical Microbiology: Open Access. 2016; 5(6).
CrossRef
- Laurent Poirel, Timothy R. Walsh, Vincent Cuvillier Patrice Nordmann. Multiplex PCR for detection of acquired carbapenemase genes Diagnostic. Microbiology and Infectious Disease. 2010; 70:119–123.
CrossRef - Walsh T., Emerging carbapenemases: a global perspective. International Journal of Antimicrobial Agents. 2010; 36, pp.8-14.
CrossRef - Singh-Moodley, A. and Perovic, O. Antimicrobial susceptibility testing in predicting the presence of carbapenemase genes in Enterobacteriaceae in South Africa. BMC Infectious Diseases. 2016; 16(1).
CrossRef - Candan E, and Aksöz N. Klebsiella pneumoniae: characteristics of carbapenem resistance and virulence factors. Acta Biochimica Polonica. 2015; 62(4), pp.867-874.
CrossRef - AlTamimi M., AlSalamah A., AlKhulaifi M. and AlAjlan H. Comparison of phenotypic and PCR methods for detection of carbapenemases production by Enterobacteriaceae.Saudi Journal of Biological Sciences. 2017; 24(1), pp.155-161.
CrossRef - Shibl A., Al-Agamy M., Memish Z., et al. The emergence of OXA-48- and NDM-1-positive Klebsiella pneumoniae in Riyadh, Saudi Arabia. International Journal of Infectious Diseases. 2013; 17(12), pp.1130-1133.
CrossRef - Ali AHI. and Omer AA. Molecular Characterization of Carbapenemase Producing Klebsiella pneumoniae Dominance of OXA48, KPC, VIM and NDM Producers in Khartoum, Sudan. Journal of Clinical Review & Case Report. 2017; 2(3), pp.1-6.
CrossRef
Abbreviations
CLSI: Clinical and Laboratory Standards Institute,
MDR/PA: Multidrug resistant P. aeruginosa.
CRE: Carbapenem resistant Enterobacteriales.
NDM: New Delhi metallo-β lactamases.
MBLs: metallo-β-lactamases.







