Manuscript accepted on :08-10-2025
Published online on: 27-10-2025
Plagiarism Check: Yes
Reviewed by: Dr. Alaa Saadi Abbood
Second Review by: Dr. Fatma Çavuş Yonar
Final Approval by: Dr. Kamal Upreti
Shaik Aminabee* , Mittapalli Meghana Sri Naga Chaitanya
, Mittapalli Meghana Sri Naga Chaitanya , Mohammad Ayesha Begum
, Mohammad Ayesha Begum , Sree Kavya Jonnalagadda
, Sree Kavya Jonnalagadda , Mudunuri Syamala Devi
, Mudunuri Syamala Devi , Polukonda Thanmai
, Polukonda Thanmai , Moka Bhagya Naga Deepika
, Moka Bhagya Naga Deepika and Saladhi Veera Mounika Lakshmi
and Saladhi Veera Mounika Lakshmi
Department of Pharmacology, V. V. Institute of Pharmaceutical Sciences, Gudlavalleru, Krishna District, A.P., India.
Corresponding Author E-mail:aminaammi786@gmail.com
Abstract
Alzheimer’s disease, Parkinson’s disease, Huntington’s disease, and amyotrophic lateral sclerosis are complex neurological disorders driven by intricate genetic and molecular mechanisms. Current therapies are mainly symptomatic and fail to correct underlying genetic defects. The advent of CRISPR-Cas9 gene-editing technology offers unprecedented opportunities to address these disorders at the genomic level through precise DNA editing, gene regulation, and epigenetic modulation. Preclinical studies using neuronal models, brain organoids, and transgenic animals have demonstrated the feasibility of correcting gene defects with CRISPR. Advances such as improved delivery systems, high-fidelity Cas9 variants, and base editing further strengthen its clinical potential. However, key challenges remain, including safe delivery across the blood–brain barrier, minimizing off-target effects, and addressing ethical concerns. Overall, CRISPR-Cas9 holds promise for integration into precision medicine to revolutionize treatment of neurodegenerative and neurodevelopmental diseases.
Keywords
CRISPR-Cas9; Gene Editing; Neurological Disorders; Neurotherapeutics; Precision Medicine
| Copy the following to cite this article: Aminabee S, Chaitanya M. M. S. N, Begum M. A, Jonnalagadda S. K, Devi M. S, Thanmai P, Deepika M. B. N, Lakshmi S. V. M. Gene Editing in Neurological Disorders: Role of CRISPR-Cas9 in Targeted Therapies. Biomed Pharmacol J 2025;18(October Spl Edition). |
| Copy the following to cite this URL: Aminabee S, Chaitanya M. M. S. N, Begum M. A, Jonnalagadda S. K, Devi M. S, Thanmai P, Deepika M. B. N, Lakshmi S. V. M. Gene Editing in Neurological Disorders: Role of CRISPR-Cas9 in Targeted Therapies. Biomed Pharmacol J 2025;18(October Spl Edition). Available from: https://bit.ly/3Le4fIt |
Introduction
One of the major causes of disability and premature death across the world is because of neurological disorders. The Global burden of disease study indicates that terms like Alzheimer disease, Parkinson disease, multiple sclerosis, Huntingtons disease, amyotrophic lateral sclerosis (ALS), and other neurodevelopmental disorders are among the most prevalent causes of the global burden of diseases.1 These disorders are characterized by progressive loss of neurons or impaired neural development, leading to cognitive decline, motor dysfunction, and profound behavioral and psychological disturbances. This burden is most pronounced with ageing populations where the prevalence of neurodegenerative diseases exhibits an ever increasing steep slope. Also, as far as economic implications are concerned, it costs billions of dollars on a yearly basis to provide direct medical care, long-term institutionalization, and providing informal care to the affected individuals, so the necessity of disease-modifying therapeutics has never been as urgent as it is right now.2
Despite advances in diagnostics and symptomatic management, traditional pharmacological approaches remain largely inadequate for treating neurological disorders. The available drugs show only temporary effects or postpone the emergence of symptoms rather than curing them and act at the genetic and molecular levels of etiologies. In addition, the complexity of the brain architecture, existence of blood-brain barrier and the multifactorial aspects of these diseases make therapeutic intervention a very difficult problem.3
In this context, the field of genome editing has emerged as a transformative approach in biomedical science. Early genome-editing tools such as zinc finger nucleases (ZFNs) and transcription activator-like effector nucleases (TALENs) provided targeted but technically complex methods for gene manipulation.4 The CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats)-Cas9 system later introduced and advanced has transformed the field of genetic engineering because it is easy to use, editable and can be programmed to make a specific set of germline or somatic genetic modifications with high levels of accuracy.5
CRISPR-Cas9 has rapidly evolved from a bacterial defense mechanism to a versatile platform for targeted gene knockout, correction of pathogenic mutations, transcriptional regulation, epigenetic modification, and even RNA editing. Such abilities have created a new era of treatment of monogenic and complex neurological disorders.6 In vitro and in vivo models of CNS disorders, preclinical studies have shown its potential to reverse the genetic mutations or turn off toxic genes, or put a disrupted gene back to normal functioning.
The objective of this article is to critically evaluate the current landscape of CRISPR-Cas9 applications in neurology. Our goal is to give an in-depth look at how it is being applied to disease modeling, therapeutic gene correction, and delivery methods across the blood-brain barrier, and new technologies like base editing and CRISPR interference/activation. As well, we look into the issues of off-target effects, immunogenicity, ethics, and regulatory aspects.7 By synthesizing these insights, this review outlines the prospects of integrating CRISPR-Cas9 into precision medicine strategies for the treatment of neurological and neurodevelopmental disorders.
Molecular Pathogenesis of Neurological Disorders
Cognitive and motor mechanisms of neurological disorders are complex and multifold on the molecular level. At the heart of many of these disorders lie specific gene disease correlations. The mutations could be inherited either in autosomal dominant or recessive fashion, and others could be acquired in de novo fashion.8 In this respect, such disease as Huntington is an autosomal dominant mutation that leads to the toxic expansion of polyglutamine through HTT gene. Likewise, most forms of familial Parkinson disease have been associated with the mutation of dominant genes, including SNCA and LRRK2. Conversely, conditions such as Friedreich ataxia are caused by autosomal recessive deformations, e. g. deformations of FXN gene and are associated with loss of activity. De novo mutations – those not inherited but appearing spontaneously are also common in severe neurodevelopmental disorders like Rett syndrome or certain forms of autism and epilepsy, involving genes such as MECP2, SCN2A, or SYNGAP1.9
However, in addition to DNA sequence alterations, epigenetic alterations are also essential in the pathogenesis of neurological diseases. Such modifications as aberrant DNA methylation, alterations to histone complexes and unbalanced non-coding RNAs can affect gene expression without affecting the genetic code. In Alzheimer’s disease, for instance, global hypomethylation and histone acetylation anomalies have been implicated in neurodegeneration and abnormal amyloid processing.10 In a similar manner, the impairment of RNA metabolism by alteration of the non-coding RNA compositions in ALS leads to progressive death of motor neurons. Due to reversibility, epigenetic modulations provide new therapeutic modulation steps including epigenome editing methods using CRISPR.11
Every neurodegenerative disorder is more or less coupled with their own typical proteinopathies which tends to interfere with the normal cellular functions. The pathogenesis of Alzheimer disease is characterized by the accumulation of the amyloid-beta peptides derived by aberrant cleavage of amyloid precursor protein (APP) and neurofibrillary tangle formation as the result of hyperphosphorylation of tau proteins.12 Huntington’s disease is marked by the expansion of CAG repeats in the HTT gene, resulting in mutant huntingtin protein that misfolds and aggregates, disrupting neuronal function and causing cell death. The main characteristic of Parkinson disease is dysfunction and aggregation of α-synuclein to form Lewy bodies. The mutations of the associated genes alter the mitochondrial activity and neurons survival leading to the progressive death of dopaminergic neurons.13 In ALS, SOD1, TARDBP, FUS and C9orf72 mutations cause unwarranted amyloids, embarrassed RNA processing and neuronal death (Table 1).
Table 1: Genetic and Pathological Features of Major Neurological Disorders.
| Disorder | Key Mutated Genes | Pathological Hallmarks | Inheritance Pattern |
| Alzheimer’s Disease | APP, PSEN1, PSEN2, APOE | Amyloid-beta plaques, Tau tangles | Mostly sporadic, some AD |
| Huntington’s Disease | HTT (CAG repeat expansion) | Polyglutamine aggregation | Autosomal dominant |
| Parkinson’s Disease | SNCA, LRRK2, PARK7 | α-synuclein aggregates | Sporadic and familial |
| ALS | SOD1, C9orf72, TDP-43 | Motor neuron degeneration | Mixed |
A combination of these genetic and epigenetic processes highlights the complexity of neurological diseases. Knowledge of these pathways is informative in linking disease genesis together with presenting a feasible way to new interventions that would be precisely directed. With its ability to directly modify genes and even influence epigenetic states, CRISPR-Cas9 technology holds great promise in addressing the root causes of these debilitating conditions.14
Mechanistic Insights into Crispr-Cas9 Technology
This CRISPR-Cas9 system which was originally borrowed as an adaptive immune system in Streptococcus pyogenes has become an adaptable and potent tool of genome-editing. It works on the basis of the interaction of the Cas9 endonuclease with a single-guide RNA (sgRNA), that guide Cas9 to a particular location in the genome by Watson-Crick base pairing with these target DNA.15 The structural biology of this system reveals that Cas9 undergoes significant conformational changes upon binding to the sgRNA, which then facilitates target DNA recognition adjacent to a protospacer adjacent motif (PAM). When the Cas9 target sequences are bound, it causes the cell to repair with a break (as a pair of strands), resulting in non-homologous end joining (NHEJ) or homology-directed repair (HDR) to accomplish gene destruction or repair.16
To enhance the precision and functionality of CRISPR, several advanced editing tools have been developed. Base editing is capable of converting the single base of DNA (e.g. C G to T A or A T to G C) without causing DSBs by fusing catalytically incompetent Cas9 to a deaminase enzyme. Prime editing is a further development of the idea which allows controlled insertions, deletions, or base substitutions in a specified region by: Nicking a target sequence in DNA with a Cas9 nickase as part of a chimeric fusion with a reverse transcriptase, and using it to insert a complementary sequence specified by a prime editing guide RNA (pegRNA).17 These tools significantly reduce off-target effects and broaden the therapeutic potential of genome editing, especially in post-mitotic neurons where DSB repair is limited.
Besides DNA rearrangement, CRISPR has been modified to regulate specific expression of genes. In CRISPR interference (CRISPRi), catalytically inactive Cas9 (dCas9) is engineered together with a transcriptional repressor e.g. KRAB allowing stable and reversible gene-silencing.18 Conversely, CRISPR activation (CRISPRa) employs dCas9 fused to activators like VP64 or p300 to upregulate target genes. The methods are especially useful in the understanding of gene activity and controlling networks in the neuronal systems, where extensive and irreversible DNA alteration might not be preferable.19
Another horizon of CRISPR applications is epigenome editing. Under epigenetic writing effect, the epigenetic landscape of particular genes can be rewritten with the help of dCas9 complemented with repurposed epigenetic modifiers (histone acetyltransferases, methyltransferases, or DNA demethylases) in a way that does not involve a change in the underlying DNA.20 In neurons, where epigenetic plasticity governs key processes such as synaptic remodeling and memory formation, these tools offer novel means of dissecting gene regulation and correcting disease-associated epigenetic aberrations.
Other CRISPR-related proteins besides Cas9 have widened the toolbox available to genome editing. Alternative targeting flexibility are conferred by Cas12a (Cpf1) with its unique PAM specificity and pattern of staggered incision.21 Cas13 is used to perform programmable RNA editing, which produces a transient and reversible up or down regulation of gene expression and this is especially desirable where using a system of permanent DNA editing creates safety concerns. Recent innovations such as CasMINI, a compact and engineered variant of Cas12f, promise to enhance delivery efficiency, especially for gene therapy applications requiring viral vectors with limited cargo capacity (Table 2).
Table 2: Comparison of CRISPR Tools and Their Functionalities
| Tool/Variant | Function | Editing Type | Application Example |
| Cas9 | DNA double-strand breaks | Knockout/In-frame | Gene disruption |
| Base Editors | Single base substitution | Precise base change | Point mutation correction |
| Prime Editors | Template-based repair | Small insertions/deletions | Genetic correction |
| dCas9-fusions | Gene activation/repression | Epigenetic editing | Neuronal gene regulation |
| Cas12/Cas13 | DNA (Cas12) / RNA (Cas13) | Transcriptome editing | RNA disorders (e.g., ALS) |
Application in Neurological Disease Models
The CRISPR-Cas9 editing tool has disrupted the process of generating disease models of neurological diseases as it allows specific and effective editing of genes in a number of biological systems. In vitro systems, especially neural cultures of 2D and 3D brain spheroids, are now of immeasurable value in modeling complex in vivo neurodevelopmental and neurodegenerative disorders.22 Brain organoids, derived from pluripotent stem cells, recapitulate key aspects of cortical architecture and allow researchers to study early disease onset, regional vulnerability, and cell-type-specific gene perturbations in a physiologically relevant context. They have successfully used such models to model genetic mutations observed in Alzheimer, Parkinson and Huntington diseases and can further observe the phenotypic outcome in near real-time upon gene editing.23
In vivo, CRISPR has been employed to generate transgenic animal models that closely mimic human neurological disease pathology. In this area mouse models have been traditionally overwhelming and it is possible to study the pathogenesis of Alzheimer, Parkinson disease by conditional or tissue-specific knockouts of a target gene such as APP, PSEN1, or LRRK2.24 More recent developments have already opened the CRISPR door to non-human primates which provide better neuroanatomical and behavioral models of human disorders. These models are crucial not only for elucidating disease mechanisms but also for testing the safety and efficacy of gene therapy candidates.
iPSC (Induced pluripotent stem cell) neurons provide an intermediate model between animal models and clinical trials and can be generated by reprogramming of iPSCs derived from patients with disease-causing mutations; therefore, it is possible to perform personalized modeling studies of disease and correction of disease-causing mutations using the CRISPR technology.25 This strategy has been employed to correct the HTT gene mutation in Huntington’s disease, restore normal neuronal morphology, and reduce cytotoxic protein aggregates in vitro. In equal measure, knock out of LRRK2 in Parkerson disease-derived neurons through iPS cells has revealed promising results in reversal of mitochondrial malfunction and alpha-synuclein pathology.26
Several compelling case studies demonstrate the therapeutic potential of CRISPR in specific neurological disorders. With Huntington disease, CRISPR-based excision of expanded CAG repeats in HTT has been demonstrated to reduce neurodegeneration in cell line- and mouse models. Targeted interference of mutant LRRK2 in Parkinson disease rescues survival and restore the functions of dopaminergic neurons. To model and modify familial mutations in APP and PSEN1, CRISPR has already been utilized in reducing the production of amyloid-beta production in vitro in Alzheimer disease.27 Regarding amyotrophic lateral sclerosis (ALS): one of the most attractive strategies is the CRISPR-mediated inhibition of hexanucleotide repeat expansions in C9orf72 that is the most common genetic cause of familial ALS cases.28 This intervention not only prevents the formation of toxic RNA foci but also reduces aberrant dipeptide repeat protein production, offering hope for a disease currently lacking effective treatments (Table 3).
Table 3: CRISPR Applications in Preclinical Neurological Models
| Model Type | Disease Studied | Gene Targeted | Outcome |
| iPSC-derived neurons | Huntington’s disease | HTT | Reduced aggregation |
| Transgenic mice | Parkinson’s disease | LRRK2 | Improved motor function |
| 3D brain organoids | Alzheimer’s disease | APP, PSEN1 | Amyloid pathology modeling |
| Zebrafish models | ALS | SOD1 | Motor neuron protection |
Delivery Strategies for Brain-Specific Gene Editing
An effective, safe, and targeted delivery of the components of gene editing targeting the brain regions is one of the primary challenges of the CRISPR-Cas9 application to neurological disorders. Brain is anatomically complicated with the existence of blood brain barrier (BBB) which is a highly selective interface that limits most macromolecules. Several delivery routes have been developed to enhance CNS targeting, including intracerebroventricular (ICV), intrathecal, and intranasal administration.29 The intrathecal and ICV provide vectors that directly inject gene-editing-tools into cerebrospinal fluid, as this route is widely distributed in the brain. The intranasal route provides a route with low invasion that can use either olfactory or trigeminal nerve routes of entry into the brain albeit with varying effectiveness.30
Viral vectors remain the most commonly used vehicles for in vivo gene editing due to their high transduction efficiency and sustained expression. The serotype 9, adeno-associated virus (AAV9) has been found to be particularly neurotrophic and has a BBB-crossing capacity, thus, making it suitable in targeting both glial cells and neurons. Lentiviral vectors are an efficient but imperfect approach with a restricted permeation of the BBB and tend to use as ex vivo or localized injectable systems.31 Despite their utility, viral vectors raise concerns regarding immunogenicity, insertional mutagenesis, and limited cargo capacity, prompting the development of alternative strategies.
Non-viral delivery systems have emerged as the more advantageous and secure alternatives to the existing and more elastic system of CNS gene delivery. mRNA-based vaccines delivered by lipid nanoparticles (LNPs) have been especially influenced by advances in other systems; LNPs are being repurposed to assist CRISPR delivery across the BBB, in particular combined with use of targeting ligands or targeted changes on the surface.32 The naturally occurring extracellular vesicles, the so-called exosomes are a promising vehicle since they have an intrinsic biocompatibility and can penetrate the blood-brain-barrier (BBB) and carry Cas9 mRNA, protein or ribonucleoprotein complexes.33 Other nanomaterials such as gold nanoparticles and dendrimers provide structural versatility and tunable surface chemistry for CNS-targeted delivery, although they remain largely in the preclinical stage (Figure 1).
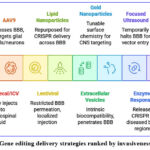 |
Figure 1: Gene editing delivery strategies ranked by invasiveness and targetingClick here to view Figure |
New formations of “smart” delivery mechanism are appearing, which can react to neural microenvironments (change in pH, enzyme levels, or redox value) in order to have spatiotemporal control over the editing of genes. Specificity and reduction of off-target effect may be achieved using these systems since the system only releases the payload in diseased or inflamed regions within the brain. For example, enzyme-responsive nanoparticles can release CRISPR components in response to matrix metalloproteinases upregulated in neurodegenerative lesions.34
A challenge to overcome the BBB is central. Tactics to get around or temporarily halt the barrier are focused ultrasound with microbubbles, receptor-mediated transcytosis with transferrin or insulin-like growth factor ligands and cell-penetrating peptides. Engineering vectors or carriers that exploit endogenous BBB transport mechanisms is a promising avenue for enhancing brain specificity without compromising barrier integrity.35
Therapeutic Potential Vs. Off-Target Risks
Although CRISPR- Cas 9 has outstanding potential in treating neurological disorders, safety issues (particularly, the ones related to off-target effects) are one of the limiting factors in clinical translation. When Cas9 catalyzes unintended mutations in the genome, known as off-target mutagenesis, whereby Cas9 generates unpredictable mutations at loci that partially match the target locus, but where double-stranded breaks are generated.36 In neurons and other post-mitotic cells, such off-target effects are particularly concerning due to limited DNA repair capacity and the inability to replace damaged cells.
Many strategies have been generated to overcome these difficulties to predict, detect and reduce off-target activity. The analysis of homology and chromatin accessibility allows predicting the potential off-targets using in silico tools: CRISPRoff, Cas-OFFinder, and CCTop. Experimental techniques like GUIDE-seq (genome-wide, unbiased identification of DSBs enabled by sequencing) and Digenome-seq allow for empirical mapping of off-target cleavage events across the genome, providing a more comprehensive assessment of CRISPR safety profiles.37
Another important area of development is an increase in fidelity of CRISPR systems. In order to limit off-target effects without affecting on-target activity, engineered derivatives of SpCas9 have been created which include: SpCas9-HF1 (high-fidelity), eSpCas9 (enhanced specificity), and xCas9.38 These variants feature amino acid substitutions that weaken non-specific DNA interactions, thereby improving target discrimination and reducing off-target cleavage.
On-target risks, like mosaicism and unwanted rearrangements of the genome, are also present despite any increment in targeting specificity. Mosaicism in which some of the cells of a given population treated do not have the same genetic alteration is particularly troublesome in in-vivo studies and potentially can yield nonconsistent results in therapies. Additionally, large deletions, insertions, or chromosomal translocations at the target site may occur due to aberrant repair of double-strand breaks, complicating interpretation of phenotypic outcomes.39
In neurological applications, long-term operation of gene control is especially important because long-term Cas9 activity can increase risks. To that end temporal control of gene editing has inducible CRISPR (such as doxycycline-controlled, light-based (optogenetic) or small molecule) systems provide the ability to activate or deactivate Cas9 expression when required.40 Self-limiting systems that degrade Cas9 after target editing or restrict expression to transient windows provide an additional safety layer, minimizing prolonged exposure to the editing machinery.
Ethical, Legal, and Regulatory Considerations
The use of CRISPR-Cas9 in the treatment of neurological disorders is associated with complicated ethical, legislative, and regulatory issues, which have to be thought through with special attention. One of the significant ethical lines of confrontation is related to the difference between somatic and germline gene editing. Whereas somatic editing of non-reproductive cells is commonly regarded as acceptable when used to treat a disease, germline editing poses serious ethical questions owing to its heritable status.41 In the context of the brain, this debate is amplified by the potential for unintended changes in cognition, personality, or behavior, especially when editing is performed during early neurodevelopmental stages.
The topic of cognitive enhancement and neuroethics is becoming a hot issue, as the CRISPR technology is more accurate and accessible. When the potential of gene editing is expanded to a capability of improving cognitive abilities to such levels as memory, attention, or intelligence; there are concerns of a future of genetic inequality, coercion, and eugenics. These concerns are especially relevant in neurological applications, where the line between therapy and enhancement can become blurred.42
There is also the problem of informed consent concerning clinical trials of neurodegenerative diseases. These conditions tend to damage the cognitive process, memory, or judgment of the patients, thus making it complex to comprehend well and consent to experimental treatments with freedom of their own will. Ethical trial design must therefore include surrogate consent provisions, patient advocacy involvement, and continuous monitoring of participants’ capacity to consent, especially in progressive conditions like Alzheimer’s or ALS.43
Internationally, the regulatory systems wielding CRISPR in the CNS are not fully developed. The NIH in the United States encourages somatic gene therapy and has a well regulated approach to it, whereas FDA insists on high leveled preclinical evidence and safety testing over several clinical years of assessment. The European Medicines Agency (EMA) emphasizes pharmacovigilance, quality control, and ethical trial protocols. India’s ICMR guidelines permit somatic cell gene editing under ethical review board approval but ban germline editing.44 World Health Organization (WHO) has proposed a registry of human genome editing trials worldwide and insisted on transparency, equality, and involvement of people in governance of gene editing.
Clinical Trials and Translational Pipeline
The former is a switch from the CRISPR-Cas9 being a potent experimental tool towards a practical treatment in the neurological field. The translational pipeline is developing soon, even though its clinical application in neurological disorders is still in its early stages. According to data from ClinicalTrials.gov, several early-phase trials are now exploring the feasibility, safety, and efficacy of CRISPR-based therapies for neurogenetic conditions, including amyotrophic lateral sclerosis (ALS), Huntington’s disease, and Leber congenital amaurosis.45 Although the majority of trials now remain in ex vivo editing or non-neuronal medicine, in vivo focus on the CNS loom ahead, with recent advances in delivery methods and preclinical verification.
Progression from bench to bedside requires a robust demonstration of therapeutic efficacy, long-term safety, and the scalability of gene-editing platforms. Not only the precise and long-term correction of pathogenic mutations but also the functional improvement of the neurological situation without side effects should be observed in preclinical models.46 Also, the manufacturing consistency, quality of vectors and dose standardization also have to be solved to receive a regulatory approval.
The clinical trajectory is also affected by regulatory and intellectual property (IP) landscape. The continuous patent conflicts especially between the Broad Institute and the University of California Berkeley have brought ambiguity to the laws that influence licensing and commercialization among several others. These disputes over foundational CRISPR patents underscore the importance of IP clarity for companies and academic institutions engaged in translational research.47-48
Another major role of collaborative consortia and public-private partnerships is in linking research and clinical application. The National Institute of Neurological Disorders and Stroke (NINDS) supports a variety of translational research initiatives to speed up the process of developing gene therapy in rare neurological disorders.49 Similarly, organizations like CureRare and EdiGENE are working to advance gene editing platforms into human trials through interdisciplinary collaboration, biomarker discovery, and patient registry development (Figure 2).
 |
Figure 2: CRISPR-Cas9 in neurological disordersClick here to view Figure |
Future Directions
With the evolving CRISPR-Cas9 technology combined with other emerging advanced fields, it seems that it can be used to facilitate how neurological diseases are treated. Among the options related to future directions that are the most disruptive, a combination of CRISPR with artificial intelligence (AI) and machine learning (ML) algorithms should be noted. These tools enable highly accurate prediction of guide RNA efficiency, off-target sites, and optimal editing windows, thus enhancing precision and reducing trial-and-error in therapeutic design.50-51 AI-based systems are as well being used to discover new therapeutic aims, and forecast gene-condition links in polygenic neurological disorders.
Multi-gene editing introduces a potent approach to address polygenic and complex disorders that have recently gained relevance to treat such diseases, as schizophrenia, epilepsy, and ASDs (autism spectrum disorders), which are commonly associated with the deregulation of many interconnected genes. Advances in multiplexed CRISPR systems, including the use of multiple guide RNAs or engineered Cas variants, allow simultaneous modulation of several targets, offering a new level of therapeutic customization.52-54
Beyond DNA editing, CRISPR-Cas13 systems offer RNA-targeting capabilities, opening avenues for reversible and transient interventions. The Cas13-based editing can be used to regulate aberrant transcriptome without editing the underlying genome, something is especially of interest when treating diseases related to toxic RNA species, like in certain types of ALS and frontotemporal dementia.
Combining CRISPR with neurotechnologies such as neuroprosthetics, optogenetics, and nanotechnology is another frontier.55-56 Such bi-systems have the capability of displaying real time feedback, spatial accuracy, and activity dependent gene regulation. As an example, it is possible to use optogenetically regulated CRISPR system which can provide temporal regulation of gene expression under light and nanomaterial-based carriers that ensure that they could be delivered with improved specificity and less immunogenicity.
Finally, CRISPR is anticipated to become a cornerstone of precision medicine in neurology. As patient specific genomic data and induced pluripotent stem cell (iPSC) models become more widely obtained, patient specific gene therapy genetically and epigenetically-specific diseases can be achievable.57-58 This method has close potential in the case of ultra-rare neurological disorders, where conventional medication is unusable.59-60
Conclusion
The innovation of CRISPR-Cas9 has brought the revolutionary kind of solution in the neurological research and therapy landscape by providing precision and versatility to gene editing like never before. Ranging between explaining the molecular mechanisms of neurological disorders that are very complicated to the fixation of pathologic mutations in preclinical designs, its uses have flourished at an extremely quick pace, and holds the prospect of offering new biomedical interventions to illnesses that were hitherto untreatable. The ability to manipulate the genome and, increasingly, the transcriptome, places CRISPR at the forefront of next-generation neurotherapeutics.61-62
Nevertheless, such promise should be sweetened with the consideration of the ethical, legal and safety challenges. The complexity of off-target effects, the impediments of delivery beyond the blood-brain barrier, and the neuroethical considerations of the neuroscience of cognitive modification or enhancement require dependable regulatory responses and tightly scientifically scrutinized. Ensuring informed consent, particularly in vulnerable populations affected by neurodegenerative disorders, is a crucial component of responsible clinical deployment.63-64
To conclude and project into the future, CRISPR can only be used to its full potential in neurology when collaboration between disciplines occurs. The translation of the bench side innovations into safe effective and secure bedside interventions will necessitate an inclusion of insights in molecular neuroscience, bioengineering, artificial intelligence, regulatory science and ethics.65 CRISPR-mediated therapies are on the horizon with careful stewardship and continued advances looking forward to revolutionizing the clinical implications of managing neurological disorders within ten years.
Acknowledgement
Authors are thankful to V. V. Institute of Pharmaceutical Sciences, Gudlavalleru, India.
Funding Source
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable
Author Contributions:
- Shaik Aminabee: Conceptualized the study, supervised the manuscript development, and critically reviewed the final draft.
- Mittapalli Meghana Sri Naga Chaitanya and Mohammad Ayesha Begum: Contributed to the literature search, data collection, and drafting of the initial manuscript.
- Sree Kavya Jonnalagadda and Mudunuri Syamala Devi: Assisted in organizing content and formatting the article as per journal guidelines.
- Polukonda Thanmai and Moka Bhagya Naga Deepika: Participated in compiling references and preparing tables and figures. \
- Saladhi Veera Mounika Lakshmi: Contributed to the final proofreading and revision.
References
- Misra MK, Damotte V, Hollenbach JA. The immunogenetics of neurological disease. Immunology. 2018;153(4):399-414.
CrossRef - Feigin VL, Vos T, Nichols E, et al. The global burden of neurological disorders: translating evidence into policy. Lancet Neurol. 2020;19(3):255-65.
CrossRef - Aminabee S, Prabhakar MC, Prasad RGSV, Lakshmana Rao A. Screening the pharmacological activity of cerium oxide nanoparticles in-vitro. Biomed Pharmacol J. 2011;4(2):287-9.
CrossRef - Deng H-X, Zhai H, Shi Y, Liu G, Lowry J, Liu B, et al. Efficacy and long-term safety of CRISPR/Cas9 genome editing in the SOD1-linked mouse models of ALS. Commun Biol. 2021;4(1):396.
CrossRef - Huang H, Sharma HS, Chen L, Saberi H, Mao G. 2018 Yearbook of Neurorestoratology. J Neurorestoratol. 2019;7(1):8-17.
CrossRef - Aminabee S, Lakshmana Rao A, Sowmya K, Nymisha D, Kusuma Naga Lakshmi K, Manikanta KVNS, Praveen Kumar P. Evaluation of analgesic activity of Ficus palmate. Iran J Pharm Sci. 2019;15(3):47-60.
- Guo N, Liu J-B, Li W, Ma Y-S, Fu D. The power and the promise of CRISPR/Cas9 genome editing for clinical application with gene therapy. J Adv Res. 2022;40:135-52.
CrossRef - Huang H, Chen L, Mao G, Sharma H. Clinical neurorestorative cell therapies: developmental process, current state, and future prospective. J Neurorestoratol. 2020;8(2):61-82.
CrossRef - Aminabee S, Lakshmana Rao A, Chinna Eswaraiah M. In-vivo antioxidant activity of different fractions of Indigofera barberi against paracetamol induced toxicity in rats. Turk J Pharm Sci. 2020;17(2):136-40.
CrossRef - Huang H, Chen L, Chopp M, et al. The 2020 Yearbook of Neurorestoratology. J Neurorestoratol. 2021;9(1):1-12.
CrossRef - Liebner S, Dijkhuizen RM, Reiss Y, Plate KH, Agalliu D, Constantin G. Functional morphology of the blood-brain barrier in health and disease. Acta Neuropathol. 2018;135(3):311-36.
CrossRef - Prasanth DSNBK, Aminabee S, Lakshmana Rao A, Teja N, Bhargavi K, Monika C, Pujitha B, Sandhya T, Lalitha A, Siva Prasad P. Anthelmintic activity of Mansoa alliacea against Pheretima posthuma: In-vitro and in-silico approach. Thai J Pharm Sci. 2020;4(3):186-96.
CrossRef - He L, Wang S, Peng L, Zhao H, Li S, Han X, et al. CRISPR/Cas9 mediated gene correction ameliorates abnormal phenotypes in spinocerebellar ataxia type 3 patient-derived induced pluripotent stem cells. Transl Psychiatry. 2021;11(1):479.
CrossRef - Yu S, Price MA, Wang Y, et al. CRISPR-dCas9 mediated cytosine deaminase base editing in Bacillus subtilis. ACS Synth Biol. 2020;9(7):1781-9.
CrossRef - Prasanth DSNBK, Aminabee S, Lakshmana Rao A, Guntupalli C, Reddy AR, Kulandaivelu U, Rao SNK, Rajeshwari P. Inhibitory effects of Mansoa alliacea in Freund’s adjuvant arthritis on inflammatory markers and its confirmation by in-silico strategy. Thai J Pharm Sci. 2021;45(6):532-44.
CrossRef - Giau VV, Lee H, Shim KH, Bagyinszky E, An SSA. Genome-editing applications of CRISPR-Cas9 to promote in vitro studies of Alzheimer’s disease. Clin Interv Aging. 2018;13:221-33.
CrossRef - Gyorgy B, Loov C, Zaborowski MP, et al. CRISPR/Cas9 mediated disruption of the Swedish APP allele as a therapeutic approach for early-onset Alzheimer’s disease. Mol Ther Nucleic Acids. 2018;11:429-40.
CrossRef - Aminabee S, Santhi Krupa D, Hanumanth Rao K, Sirisha V, Anitha Kumari V, Nagasen D, Sujiya B, Leelavati TS. Pharmacoeconomic analysis of biologic vs. biosimilar therapies in rheumatoid arthritis. Int J Chem Biochem Sci. 2023;24(4):395-400.
- Newcombe EA, Camats-Perna J, Silva ML, Valmas N, Huat TJ, Medeiros R. Inflammation: the link between comorbidities, genetics, and Alzheimer’s disease. J Neuroinflammation. 2018;15(1):276.
CrossRef - Raikwar SP, Thangavel R, Dubova I, et al. Targeted gene editing of glia maturation factor in microglia: a novel Alzheimer’s disease therapeutic target. Mol Neurobiol. 2019;56(1):378-93.
CrossRef - Aminabee S, Sirisha K, Ravi Shankar K, Revanth G, Yaswanth K, Jaswanth Kumar K, Rishitha K. Pharmacological interventions for relieving neuropathic pain in diabetic patients. Int J Chem Biochem Sci. 2023;24(4):414-9.
- Konstantinidis E, Molisak A, Perrin F, Streubel-Gallasch L, Fayad S, Kim DY, et al. CRISPR-Cas9 treatment partially restores amyloid-β 42/40 in human fibroblasts with the Alzheimer’s disease PSEN1 M146L mutation. Mol Ther Nucleic Acids. 2022;28:450-61.
CrossRef - Park H, Oh J, Shim G, et al. In vivo neuronal gene editing via CRISPR-Cas9 amphiphilic nanocomplexes alleviates deficits in mouse models of Alzheimer’s disease. Nat Neurosci. 2019;22(4):524-8.
CrossRef - Leelavati TS, Madhavi S, Kamal G, Raju PVM, Susmitha K, Vinod M, Aminabee S. Revolutionizing healthcare delivery: Telemedicine’s influence on access and patient satisfaction. Int J Chem Biochem Sci. 2023;24(5):106-15.
- Balendra R, Isaacs AM. C9orf72-mediated ALS and FTD: multiple pathways to disease. Nat Rev Neurol. 2018;14(9):544-58.
CrossRef - Lopez-Gonzalez R, Yang D, Pribadi M, et al. Partial inhibition of the overactivated Ku80-dependent DNA repair pathway rescues neurodegeneration in C9ORF72-ALS/FTD. Proc Natl Acad Sci U S A. 2019;116(19):9628-33.
CrossRef - Leelavati TS, Madhavi S, Susmitha K, Venkateswara Kumar KS, Vara Prasad Goud P, Ganga Raju K, Aminabee S. Exploring university student attitudes, beliefs, and alcohol usage patterns: An investigation into alcohol and drug use within the student lifestyle. J Drug Alcohol Res. 2023;12(8):1-6.
- Krishnan G, Zhang Y, Gu Y, Kankel MW, Gao FB, Almeida S. CRISPR deletion of the C9ORF72 promoter in ALS/FTD patient motor neurons abolishes production of dipeptide repeat proteins and rescues neurodegeneration. Acta Neuropathol. 2020;140(1):81-4.
CrossRef - Ababneh NA, Scaber J, Flynn R, et al. Correction of amyotrophic lateral sclerosis related phenotypes in induced pluripotent stem cell-derived motor neurons carrying a hexanucleotide expansion mutation in C9orf72 by CRISPR/Cas9 genome editing using homology-directed repair. Hum Mol Genet. 2020;29(13):2200-17.
CrossRef - Aminabee S, Devi Mallika K, Lakshmana Rao A. Natural resources in management and treatment of diseases: A review. Pharma Times. 2023;55(11):7-10.
- Nance ME, Shi R, Hakim CH, et al. AAV9 edits muscle stem cells in normal and dystrophic adult mice. Mol Ther. 2019;27(9):1568-85.
CrossRef - Matre PR, Mu X, Wu J, et al. CRISPR/Cas9-based dystrophin restoration reveals a novel role for dystrophin in bioenergetics and stress resistance of muscle progenitors. Stem Cells. 2019;37(12):1615-28.
CrossRef - Aminabee S, Lakshmana Rao A. Computational identification of selected bioactive compounds from Cedrus deodara as inhibitors against SARS-CoV-2 main protease: A pharmacoinformatics study. Indian Drugs. 2024;61(2):78-91.
CrossRef - Xu L, Lau YS, Gao Y, Li H, Han R. Life-long AAV-mediated CRISPR genome editing in dystrophic heart improves cardiomyopathy without causing serious lesions in mdx mice. Mol Ther. 2019;27(8):1407-14.
CrossRef - Nelson CE, Wu Y, Gemberling MP, et al. Long-term evaluation of AAV-CRISPR genome editing for Duchenne muscular dystrophy. Nat Med. 2019;25(3):427-32.
CrossRef - Madhavi S, Leelavati TS, Varaprasad Goud P, Kavitha C, Vijay Kumar M, Lakshmi B, Aminabee S. Unravelling the impact of alcohol use on college students’ educational success: A systematic analysis. J Drug Alcohol Res. 2024;13(2):1-5.
- Lu L, Yu X, Cai Y, Sun M, Yang H. Application of CRISPR/Cas9 in Alzheimer’s Disease. Front Neurosci. 2021;15:803894.
CrossRef - Petkova MV, Stantzou A, Morin A, et al. Live-imaging of revertant and therapeutically restored dystrophin in the Dmd(EGFP-mdx) mouse model for Duchenne muscular dystrophy. Neuropathol Appl Neurobiol. 2020;46(6):602-14.
CrossRef - Ganga Raju K, Varaprasad Goud P, Parween S, Ramarao M, Leelavati TS, Madhavi S, Aminabee S. Economic analysis of pharmacological interventions for substance use disorders: Cost-effectiveness and market dynamics. J Drug Alcohol Res. 2024;13(7):1-4.
- Zhu P, Wu F, Mosenson J, et al. CRISPR/Cas9-mediated genome editing corrects dystrophin mutation in skeletal muscle stem cells in a mouse model of muscle dystrophy. Mol Ther Nucleic Acids. 2017;7:31-41.
CrossRef - Lee K, Conboy M, Park HM, et al. Nanoparticle delivery of Cas9 ribonucleoprotein and donor DNA in vivo induces homology-directed DNA repair. Nat Biomed Eng. 2017;1:889-901.
CrossRef - Min YL, Chemello F, Li H, et al. Correction of three prominent mutations in mouse and human models of Duchenne muscular dystrophy by single-cut genome editing. Mol Ther. 2020;28(9):2044-55.
CrossRef - Ravi Shankar K, Aminabee S, Ramana G, Chenchu Lakshmi KNV, Indusree G. Formulation and evaluation of propranolol hydrochloride floating tablets by 32 factorial design. Int J Drug Deliv Technol. 2024;14(4):2060-7.
CrossRef - Lattanzi A, Duguez S, Moiani A, et al. Correction of the exon 2 duplication in DMD myoblasts by a single CRISPR/Cas9 system. Mol Ther Nucleic Acids. 2017;7:11-19.
CrossRef - Duchene BL, Cherif K, Iyombe-Engembe JP, et al. CRISPR-induced deletion with SaCas9 restores dystrophin expression in dystrophic models in vitro and in vivo. Mol Ther. 2018;26(11):2604-16.
CrossRef - Leelavati TS, Madhavi S, Aminabee S. Fostering ethical decision-making skills in pharmaceutical education: Vital for future pharmacists. Indian J Pharm Educ Res. 2025;59(2):433-40.
CrossRef - Brescia M, Janssen JM, Liu J, Goncalves M. High-capacity adenoviral vectors permit robust and versatile testing of DMD gene repair tools and strategies in human cells. Cells. 2020;9(4):869.
CrossRef - Xiang X, Zhao X, Pan X, et al. Efficient correction of Duchenne muscular dystrophy mutations by SpCas9 and dual gRNAs. Mol Ther Nucleic Acids. 2021;24:403-15.
CrossRef - Aminabee S, Lakshmana Rao A, Chinna Eswaraiah M. Hepatoprotective activity of Michelia nilagirica against paracetamol induced hepatic injury in rats. Pharmacogn J. 2015;7(4):1-7.
CrossRef - Scheffer IE, Nabbout R. SCN1A-related phenotypes: epilepsy and beyond. Epilepsia. 2019;60(Suppl 3):S17-S24.
CrossRef - Mei D, Cetica V, Marini C, Guerrini R. Dravet syndrome as part of the clinical and genetic spectrum of sodium channel epilepsies and encephalopathies. Epilepsia. 2019;60(Suppl 3):S2-S7.
CrossRef - Aminabee S, Lakshmana Rao A, Chinna Eswaraiah M. Antidepressant activity of chloroform extract of Indigofera barberi in experimental animal models. Int J Chem Sci. 2016;14(2):739-50.
- Colasante G, Lignani G, Brusco S, et al. dCas9-based scn1a gene activation restores inhibitory interneuron excitability and attenuates seizures in Dravet syndrome mice. Mol Ther. 2020;28(1):235-53.
CrossRef - Colasante G, Qiu Y, Massimino L, et al. In vivo CRISPRa decreases seizures and rescues cognitive deficits in a rodent model of epilepsy. Brain. 2020;143(3):891-905.
CrossRef - Tabrizi SJ, Ghosh R, Leavitt BR. Huntingtin lowering strategies for disease modification in Huntington’s disease. Neuron. 2019;102(4):899.
CrossRef - Devi NKD, Aminabee S, Radha Madhavi B, Lakshmi Pavani A, Anasurya T, Kamepalli S, Swathi Lakshmi V, Sravya V, Sai Krishna S, Manasa GML, Nagalakshmi A, Surendra K. Comparative in-vitro antihelmintic activity of prepared, branded and generic albendazole samples in Pheretima posthuma. J Drug Alcohol Res. 2023;12(6):1-5.
- Ekman FK, Ojala DS, Adil MM, et al. CRISPR-Cas9-mediated genome editing increases lifespan and improves motor deficits in a Huntington’s disease mouse model. Mol Ther Nucleic Acids. 2019;17:829-39.
CrossRef - Kolli N, Lu M, Maiti P, Rossignol J, Dunbar GL. Application of the gene editing tool, CRISPR-Cas9, for treating neurodegenerative diseases. Neurochem Int. 2018;112:187-96.
CrossRef - Aminabee S, Rammohan Rao C, Lakshmana Rao A. Influence of Allium sativum on the antihyperglycaemic activity of gliclazide in diabetic rats: An herb-drug interaction study. Indian Drugs. 2023;60(12):82-4.
CrossRef - Safari F, Hatam G, Behbahani AB, et al. CRISPR system: a high-throughput toolbox for research and treatment of Parkinson’s disease. Cell Mol Neurobiol. 2020;40(4):477-93.
CrossRef - Kantor B, Tagliafierro L, Gu J, et al. Downregulation of SNCA expression by targeted editing of DNA methylation: a potential strategy for precision therapy in PD. Mol Ther. 2018;26(11):2638-49.
CrossRef - Li H, Yahaya BH, Ng WH, Yusoff NM, Lin J. Conditioned medium of human menstrual blood-derived endometrial stem cells protects against MPP(+)-induced cytotoxicity in vitro. Front Mol Neurosci. 2019;12:80.
CrossRef - Aminabee S, Ravi Shankar K, Chenchu Lakshmi KNV, Saritha K, Kavya R, Chaitanya Babu K, Santhi Krupa D. Influence of Allium sativum on the hypoglycaemic activity of gliclazide in normal rats: A possible approach to herb-drug interaction. Biomed Pharmacol J. 2024;17(1):377-82.
CrossRef - Lee J, Bayarsaikhan D, Arivazhagan R, et al. CRISPR/Cas9 edited sRAGE-MSCs protect neuronal death in Parkinson’s disease model. Int J Stem Cells. 2019;12(1):114-24.
CrossRef - Hajiahmadi Z, Movahedi A, Wei H, et al. Strategies to increase on-target and reduce off-target effects of the CRISPR/Cas9 system in plants. Int J Mol Sci. 2019;20(15):3719.
CrossRef
Abbreviations List:
CRISPR-Cas9: Clustered Regularly Interspaced Short Palindromic Repeats-CRISPR-associated protein 9, ALS: Amyotrophic Lateral Sclerosis, ZFNs: Zinc Finger Nucleases, TALENs: Transcription Activator-Like Effector Nucleases, APP: Amyloid Precursor Protein, sgRNA: Single-Guide RNA, PAM: Protospacer Adjacent Motif, NHEJ: Non-Homologous End Joining, HDR: Homology-Directed Repair, pegRNA: Prime Editing Guide RNA, IPSC: Induced Pluripotent Stem Cell, IC: Intracerebroventricular, AAV: Adeno-Associated Virus, LNPs: Lipid Nanoparticles, BBB: Blood-Brain-Barrier, GUIDE-seq: Genome-Wide, Unbiased Identification of DSBs Enabled by Sequencing, EMA: European Medicines Agency, WHO: World Health Organization, IP: Intellectual Property, NINDS: Neurological Disorders and Stroke, AI: Artificial Intelligence, ML: Machine Learning, ASDs: Autism Spectrum Disorders







