Manuscript accepted on :05-09-2025
Published online on: 09-10-2025
Plagiarism Check: Yes
Reviewed by: Dr. Aybike Turkmen
Second Review by: Dr. M Mohan Varma
Final Approval by: Dr. Eman Refaat Youness
Reehana.Shaik1 , Ishwarya Obilineni3
, Ishwarya Obilineni3 , Naga Venkata Chenchu Lakshmi Kollipara2*
, Naga Venkata Chenchu Lakshmi Kollipara2* , K Ravishankar4
, K Ravishankar4 and Saipraneeth Tripuraneni4
and Saipraneeth Tripuraneni4
1Department of Pharmaceutical Analysis, KL College of Pharmacy, Koneru Lakshmaiah Education Foundation, Vaddeswaram, Andhra Pradesh, India
2Department of Pharmaceutical Chemistry, KL College of Pharmacy, Koneru Lakshmaiah Education Foundation, Vaddeswaram, Andhra Pradesh, India.
3Department of Pharmacology, KVSR Siddhartha College of Pharmaceutical Sciences, Vijayawada, Andhra Pradesh, India.
4Department of Pharmaceutics, KVSR Siddhartha College of Pharmaceutical Sciences, Vijayawada, Andhra Pradesh, India.
Corresponding Author E-mail:chenchu42@gmail.com
DOI : https://dx.doi.org/10.13005/bpj/3311
Abstract
The study's purpose is to conduct a thorough analysis of the Saussurea lappa root extract. The research focused on primary screening for phytochemicals followed by GC-MS analysis of both polar and non-polar extracts. A total of 34 components were found in the root extract; 1, 8-cyclopentadecadiyne was found to have the highest percentage in both extracts. The findings from SWISS target prediction, the PASS online database, and pdCSM-cancer are corroborated by the examination of structure-based biological activities, demonstrating that the anticancer capabilities of this plant root chemical are derived from structure-based drug advancements. The in-silico outcomes showed interactions involving alkyl bonds, binding affinity with EGFR kinase inhibitors, anti-proliferative proteins, and Steroid DELTA-isomerize target proteins were determined. A computational synergistic study was also performed to predict the possible drug combinations that can be administered with Saussurea lappa to treat cancer. This suggests an increased potential for developing anticancer drugs to treat breast cancer.
Keywords
Anticancer activity; BIOSIG online; GC-MS;In-silico; Molecular docking; Sassurea lappa; SWISS target prediction
Download this article as:| Copy the following to cite this article: Shaik R, Obilineni I, Kollipara N. V. C. L, Ravishankar K, Tripuraneni S. Comprehensive Analysis of Saussurea lappa: GC-MS Profiling, In Silico Evaluation, Toxicity Prediction, and Molecular Docking for Breast Cancer Therapy. Biomed Pharmacol J 2025;18(4). |
| Copy the following to cite this URL: Shaik R, Obilineni I, Kollipara N. V. C. L, Ravishankar K, Tripuraneni S. Comprehensive Analysis of Saussurea lappa: GC-MS Profiling, In Silico Evaluation, Toxicity Prediction, and Molecular Docking for Breast Cancer Therapy. Biomed Pharmacol J 2025;18(4). Available from: https://bit.ly/4nE8Ulj |
Introduction
Malignancy of breast is the most frequent disease detected in women globally; expected 2.3 million new cases were logged in 2020. Despite significant advancements in treatment modalities—including surgery, chemotherapy, hormonal therapy, and targeted treatments like PARP inhibitors and CDK4/6 blockers—the disease remains an significant cause of cancer-related mortality.1 The rise of resistance to traditional treatments is an important barrier in dealing breast cancer, particularly in aggressive subtypes like TNBC (triple-negative breast cancer) and HER2+ve breast cancer.2 Additionally, the severe side effects associated with chemotherapy, such as myeloid suppression, cardio toxicity, and neurotoxicity, underscore there is an urgent demand for more secure and efficient alternatives to therapy.
Historically, natural ingredients have proven essential to the research and development of anticancer regimens with approximately 60% of approved chemotherapeutic agents derived from plant bases. Phytochemicals for instance paclitaxel, vincristine and camptothecin exemplify a greater therapeutic potential in oncology.3 given the structural diversity and bioactivity of plant-derived compounds, medicinal plants continue to be an invaluable resource for identifying novel anticancer leads.
Saussurea lappa (Decne.) Sch. Bip., (commonly name costus or kuth), is a biennial plant native to the Himalayan geographical area and frequently employed in ISM (Indian system of medicine) and TCM(Traditional Chinese Medicine).4 The roots of S. lappa have been traditionally employed for treating inflammation, respiratory disorders, gastrointestinal ailments, and microbial infections.5 Recent pharmacological studies have also highlighted its anticancer potential, with crude extracts and isolated compounds demonstrating cytotoxic properties on varied cancer cell lines, including those from the breast, lung, and colon.6,7
Phytochemical investigations have identified several bioactive constituents in S. lappa, primarily Sesquiterpene lactones, including dehydrocostus lactone and costunolide, and saussureamines, which exhibit anti-inflammatory, antioxidant, and antitumor properties.8
Saussurea lappa roots are rich in various compounds of different chemical groups. They include monoterpenes,9 Sesquiterpenes,10 sesquiterpene lactones,11 triterpenes,12 sterols,13 flavonoids,14 coumarins,15 and phenylpropanoids.16 The oil extracted from S.lappa root was found useful in treating leprosy17 as well as an arthritic pain reliever.18 Several researchers have extensively studied the therapeutic activity of these roots against chronic gastritis, rheumatoid arthritis, asthma, stomach ulcers, etc.19
These chemicals modulate critical carcinogenic pathways such NF-κB, PI3K/AKT, and MAPK/ERK which are frequently dysregulated in breast cancer.20 However, despite these promising findings, a systematic analysis of S. lappa’s phytochemical composition and its molecular interactions with breast cancer targets remains limited.
GC-MS is a consistent analytical method frequently used to identify volatile and semi-volatile chemical substances in plant extracts.21 By combining GC-MS and metabolomics techniques, researchers can acquire a complete phytochemical profile, enabling the detection of potential bioactive molecules with therapeutic relevance.
Molecular docking, ADMET prediction, and other in-silico techniques and pharmacophore modeling, have become indispensable tools in modern drug discovery.22 these computational approaches allow for the rapid screening of phytochemicals against disease-specific molecular targets, predicting binding affinities, drug-likeness, and toxicity profiles before costly and time-consuming experimental justification.
Materials and Methods
Chemicals required
n-Hexane, Methanol, ethyl acetate, silica gel powder, Molisch’s reagent, concentrated sulphuric acid, Barfoed’s reagent, acetic anhydride, conc hydrochloric acid, chloroform, ninhydrin, acetone, copper sulphate solution, ethanol, KOH pellets, The reagents of Hager, Wagner, and Dragendorff, as well as, Mayer’s reagent, dilute ammonia, Baljet’s reagent, zinc dust, 1% gelatin solution, 10% NaCl, 5% FeCl3.
Collection and Preparation of plant extract
Saussurea lappa roots were purchased in a local market in Vijayawada, Andhra Pradesh. The roots were properly cleansed with water to eliminate debris before being shade-dried. The dried roots were pulverized and kept in a container.23
10g of Saussurea lappa (powder of root) weighed and mixed to 300 milliliter of distilled water. The resulting solution was reduced to its half by heating on a hot plate i.e., 150mL. The solution was filtered using clean muslin cloth to remove any impurities. The filtered extract was allowed to rest for 24 hours, so that the sediment settles in the bottom of the beaker. After 24 hours, carefully filter the solution and the filtrate was used for the study.
Preliminary phytochemical screening
Phytochemical profiling of the aqueous extract from Saussurea lappa roots was carried out using established methodologies.24, 25, 26 Molisch’s test involved adding a few drops of alcoholic α-naphthol to the test sample, followed by 2-3 drops of conc sulphuric acid through the test tube’s sides, if the sample contains carbohydrates purple to violet color ring appears at the junction solvents in test tube. The Libermann-Burchard test was performed by adding a small number of droplets of acetic anhydride to the sample with subsequent scalding and followed by chilling it. Then, add strong sulphuric acid to the test tube. The presence of steroids is indicated by the top layer turning green and a brown ring forming where the two levels meet, and a deep red colour appears, indicating the presence of triterpenoids. To conduct the hydrochloric acid test, a few droplets of hydrochloric acid had been added to the extract. The extract’s pink coloration suggests the presence of lignin molecules or resins.
Hesser’s test was carried out by adding concentrated sulfuric acid to the test tube’s side walls and treating the extract with chloroform. Progress the phytosterols presence by color blood red. To conduct the Biuret test, a drop of 2% copper sulphate solution, one millilitre of 95% ethanol and potassium hydroxide pellets were included to 2mL of the plant extract, appearance of blue to purple or violet indicates presence of amino acids. Hager’s test was carried out by adding Hager’s reagent (saturated picric acid solution) to a few milliliters of plant extract; the emergence of red color shows the presence of alkaloids. The zinc-hydrochloride reduction test was carried out by adding a little amount of zinc dust and concentrated HCl along the interior of the test tube to a few ml of plant extract. Together, these methodologies provided a detailed phytochemical profile of Saussurea lappa root extract, offering valuable insights into its medicinal properties and potential therapeutic applications.
GC/MS study
Sample readiness
Put 10 milliliters of the sample extract into each of two centrifuge tubes. Add 10 milliliter of a non-polar solvent, such as n-hexane, to one tube and 10 ml methanol, to the other. Thoroughly mix the sample extract and solvents in each tube by using a vortex mixer to ensure proper dissolution and distribution of the sample components. Followed by, centrifuge the mixtures using a REMI-8C Plus centrifuge machine at 5000 RPM for 10 minutes which separates the components based on their density. After centrifugation, the supernatant, which is the liquid layer on top of the sediment or pellet formed after centrifugation, is then transferred to separate tubes carefully for GC-MS analysis.27, 28
A NexiD GC-2030 gas chromatograph was used in conjunction with a GCMS-QP2020NX mass spectrometer to analyze polar and non-polar extracts of Saussurea lappa roots. The analysis utilized a capillary column containing 95% dimethyl-polysiloxane and 5% diphenyl (SH-Rxi-5Sil MS; 30 m x 0.250 mm x 0.25 µm). The injector temperature was set to 280°C, while the oven temperature began at 350°C and progressively increasing to 400°C at a rate of 3°C/min for 2 minutes, followed by a final increase to 300°C at 15°C/min for 2 minutes.
At a column flow rate of 0.5 mL/min, XL-grade helium was employed as the carrier gas for the 44-minute GC-MS study. The temperature of the GC-MS interface was kept constant at 310°C, and the MS was operated in scan mode using an electron ionization system. The MS quadrupole with source temperatures were both set at 280°C, with a mass range of 1050 m/z being investigated.
Identification of compounds
The chemicals were identified utilizing the “NIST-MS Library 2009”. 29 The relative percentage of each compound was measured based on the peak area from the Total Ion Chromatogram (TIC), with the calculations being done automatically.
In Silico Studies
The initial selection of targets was based on their physicochemical properties and drug-like characteristics, identified through tools like PASS online and Swiss Target Prediction.30, 31, 32, 33 this was followed by molecular docking using AutoDock Pyrx34 and BIOVIA Discovery Studio visualizer35 and ADME toxicity prediction. The study also explored potential drug combinations that could be used synergistically in the treatment of breast cancer.
ADMET and physicochemical property prediction
The virtual prediction technique was used to ascertain the lead molecule’s physicochemical characteristics. Molinspiration to assess its drug-likeness characteristics and Lipinski’s Rule of 5 and PkCSM to assess its toxicity. Toxicity was also assessed by toxtree to determine the toxicity class.
PASS analysis
The PASS web server36 was used to examine the compound’s possible biological characteristics. The PASS tool helps in the prediction of possible biological activities of compounds. The test was conducted by inputting the SMILES. The data were analyzed based on the Pa (Probable Activation) and Pi (Probable Inactivation) values. A higher Pa value suggests a greater likelihood that the compound exhibits the predicted biological activity.
SWISS target prediction
Swiss Target prediction, a web based application that has been accessible since 2014 and is aimed to predict the utmost probable protein targets for smaller chemical entities. The predictions are completed via reverse screening.37 Ligands are uploaded using the SMILES format. The 2019 version is a big update that includes enhancements to the fundamental facts, backend & online interface.
Docking
Molecular docking research was done out utilizing the commercial application Auto Dock Pyrex. This method determines the preferred orientation of one molecule, such as a drug, relative to another, such as a protein, in order to obtain insight into their molecular interactions.
Protein Preparation and Ligand Preparation
Targeted receptor crystal structures were obtained from the RCSB Protein Data Bank, 38 specifically the proliferative peroxisome protein [PDB: 6MS7] and steroid DELTA-isomerize [PDB: 1OH0]. The primary module was used to prepare the proteins by eliminating water molecules, balancing charges, and repairing any side chains and residues that were missing. The National Centre for Biotechnology Information’s PubChem database provided the GC-MS identification of the chemical 1, 8-Cyclopentadecadiyne for its known anti-inflammatory and anticancer properties. The results of the molecular docking analysis, along with graphical images, were processed using the visualization program/server BIOVIA, PDBsum.
Biosig Tools to predict cancer activity:
BioSig is an open-source platform (https://biosig.lab.uq.edu.au/tools) that offers various tools like pdCSM-cancer that provides information regarding the compound ability to act against cancer and PiscesCSM provides various synergistic and antagonistic molecules with respect to the ligand.
pdCSM-cancer
pdCSM- cancer is a new predictive technique that we used in this investigation. This technique accurately predicts which small composite molecules active against one or more malignant cell lines39 by using a plot-based signature depiction of the molecules’ chemical structure. Here, we run using cyclo-pentadiene smiles to identify the target molecules against cancer cell lines.
PiscesCSM
Graph-based signatures combined with physicochemical features allow us to predict synergistic medication combinations for classification tasks40 better than existing approaches. Furthermore, this online server solution offers an easy-to-use platform for searching for possible drug combinations that work well together to combat one or more malignant cell lines. https://biosig.lab.uq.edu.au/piscescsm/
Gusar Model of Acute Rat Toxicity
The compound’s acute toxicity in rats was studied using the GUSAR tool41.This investigation was based on quantitative atom descriptor neighborhoods and a substance activity spectrum prediction method. The predictions were compared to the SYMYX MDL toxicity database, and the compounds were then classified using the Organization for Economic Cooperation and Development (OECD) manual42 criteria. The median lethal dose (LD50), which represents the quantity of the medicine that kills 50% of the research animals exposed to it, is used to express hazardous dosages. The LD50 value is 43, 44 given as the weight of the drug entity per unit of body weight (mg/kg).
Results
Primary phytochemical analysis
Phytoconstituent screening of S.lappa roots of aqueous extract using various chemical reactions revealed that the extract confirms the phytoconstituent presence such as alkaloids, flavonoids, phytosterols, lignin, resins, carbohydrates, steroids and triterpenoids with observation as given in Table 1 and depicted in Figure 1.
Table 1: Preliminary phytochemical analysis of root extract of S.lappa
| S.No | Constituent | Result |
| i. | Alkaloids | + |
| ii. | Flavonoids | + |
| iii. | Phytosterols | + |
| iv. | Lignins | + |
| v. | Resins | + |
| vi. | Carbohydrates | + |
| vii. | Steroids | + |
| viii. | Protein | – |
| ix. | Triterpenoids | + |
GC-MS profiling of S.lappa
A total of 34 compounds 16 polar (Methanol) extract compounds and 15 non-polar (n-hexane) extract compounds were recognized from the GC-MS analysis of S.lappa roots. The tables 2 and 3 provide the chemical contents in S.lappa roots, including their Rt, MF, molecular weight, and concentration, as well as their online biological activities.
Table 2: GC-MS study listed phytochemical components in polar roots extract of Sassurea lappa and their biological activities
| S.No | Rt | Name | MF | Mol.wt (g/mol) | Peak Area% | Pass online bioactivity |
| 1 | 27.371 | 9-Octadecynoic acid | C18H32O2 | 280.4 | 0.15 | Inhibitors of Acrocylindropepsin, Saccharopepsin, Chymosin and Dextranase targets |
| 2 | 28.463 | 1,3-Bis-(2-cyclopropyl, 2-methylcyclopropyl)-t | C18H26O | 258.4 | 0.66 | Testosterone 17beta- dehydrogenase (NADP+) inhibitor, CYP2J substrate, Antieczematic, Acylcarnitine hydrolase inhibitor, Dermatologic, Beta glucuronidase inhibitor |
| 3 | 31.101 | Aromandendrene | C15H24 | 204.35 | 0.9 | Antieczematic, Dermatologic, Antineoplastic, Ovulation inhibitor, Testosterone 17beta-dehydrogenase (NADP+) inhibitor |
| 4 | 32.359 | Octasiloxane, 1,1,3,3,5,5,7,7,9 ,9,11,11,13,13,1 |
C16H50O7Si8 | 579.25 | 2.64 | Apoptosis agonist, Antineoplastic, Antineoplastic (colorectal cancer), Antineoplastic (colon cancer). |
| 5 | 33.16 | Cyclopentane-3′-spirotricyclo[3.1.0.0(2,4)]hex | C14H20 | 188.31 | 1.06 | Testosterone 17beta-dehydrogenase (NADP+) inhibitor, Nicotinic alpha6beta3beta4alpha5 receptor antagonist. |
| 6 | 33.28 | 1,3-Bis-(2-cyclopropyl,2-methylcyclopropyl)-t | C18H26O | 258.4 | 1.01 | Testosterone 17beta-dehydrogenase (NADP+) inhibitor, CYP2J substrate, Antieczematic, Acylcarnitine hydrolase inhibitor, Dermatologic, Beta glucuronidase inhibitor |
| 7 | 34.223 | n-Hexadecanoic acid | C16H32O2 | 256.424 | 0.68 | Alkenylglycerophosphocholine hydrolase inhibitor, Acylcarnitine hydrolaseinhibitor, Alkylacetyl glycerophosphatase inhibitor, CYP2J substrate, CYP2J2 substrate, Acrocylindropepsin inhibitor |
| 8 | 35.125 | Benzoic acid, 2,4-bis[(trimethylsilyl)oxy]-, trir | C16H30O4Si3 | 370.66 | 2.54 | Apoptosis agonist, Antineoplastic, Antineoplastic (colorectal cancer), Antineoplastic (colon cancer). |
| 9 | 35.213 | Cedranoxide, 8,14- | C15 H24 O | 220.35 | 0.44 | Steroid-like, Hepatic disorders treatment, Cachexia treatment, RNA-directed RNA polymerase stimulant, RNA agonist |
| 10 | 35.345 | 1,2-Benzenedicarboxylic acid, dipropyl ester | C14H18O4 | 250.290 | 0.67 | Sugar-phosphatase inhibitor, Antiseborrheic, Alkenylglycero phosphocholine hydrolase inhibitor, Aspulvinonedimethyl allyltransferase inhibitor. |
| 11 | 35.556 | Cyclopropane, 1-(2-methylene-3-butenyl)-1-(1 | C12H18 | 162.27 | 9.19 | CYP2E1 substrate, CYP2E substrate, Aspulvinonedimethylallyltransferase inhibitor, CYP2J substrate, G-protein-coupled receptor kinase inhibitor |
| 12 | 36.032 | cis-.alpha.-Bisabolene | C15H24 | 204.351 | 0.68 | Angiogenesis inhibitor, TP53 expression enhancer, Antineoplastic, Endothelial growth factor antagonist, CDP-glycerol glycerophosphotransferaseinhibitor, Antidiabetic symptomatic |
| 13 | 36.171 | 1.3-Benzodioxin-4-one, 4a, 5-dimethylperhydrc | C10H16O3 | 184.23 | 0.3 | Phosphatase inhibitor, Anti dyskinetic, Acylcarnitine hydrolase inhibitor, |
| 14 | 36.593 | 1.8-Cyclopentadecadiyne | C15H22 | 202.33 | 70.59 | Steroid DELTA-isomerase inhibitor, Aspulvinonedimethylallyltransferase inhibitor, Nicotinic alpha-6-beta-3-beta-4alpha5 receptor antagonist, Antineoplastic. |
| 15 | 43.869 | Benzoic acid, 2,4-bis[(trimethylsilyl)oxy]-, trir | C16H30O4Si3 | 370.66 | 4.1 | Apoptosis agonist, Antineoplastic, Antineoplastic (colorectal cancer), Antineoplastic (colon cancer). |
| 16 | 44.906 | Aminoguanidine | NH2NHC(=NH)NH2 · HCl | 110.55 | 2.61 | Apoptosis agonist, Antineoplastic,Antineoplastic (colorectal cancer),
Antineoplastic (colon cancer). |
Table 3: GC-MS study enlist phytochemical components in non-polar roots part of Sassurea lappa , as well as their biological activities (PASS online).
| S.No | Rt | Name | MF | Mol wt (g/mol ) | Peak Area% | Pass online bioactivity |
| 1 | 26.935 | Aromandendrene | C15H24 | 204.35 | 0.76 | Antieczematic, Dermatologic, Antineoplastic, Ovulation inhibitor, Testosterone 17beta-dehydrogenase (NADP+) inhibitor |
| 2 | 27.635 | 9,12,15-Octadecatrienoic acid, (Z,Z.Z)- | C18H30O2 | 278.4296 | 2.25 | Antieczematic,CYP2J substrate, CYP2J2 substrate, Linoleate diol synthase inhibitor, Phosphatidylglycerophosphatase inhibitor, |
| 3 | 28.469 | 1,11-Hexadecadiyne | C18H32O2 | 280.4 | 0.54 | Antieczematic, Dermatologic, Antineoplastic,
Ovulation inhibitor, Testosterone 17beta- dehydrogenase (NADP+) inhibitor |
| 4 | 29.547 | 3-Cyclohexene-1-carboxaldehyde, 1,3,4-trime | C10H16O | 152.2334 | 0.8 | Testosterone 17beta-dehydrogenase (NADP+)
inhibitor,Nicotinic alpha6beta3beta4alpha5 receptor antagonist |
| 5 | 30.826 | Cyclohexane, 1-ethenyl-1-methyl-2,4-bis(1-m | C15H24 | 204.35 | 2.07 | Antieczematic, Dermatologic, Antineoplastic,
Ovulation inhibitor, Testosterone 17beta- dehydrogenase (NADP+) inhibitor |
| 6 | 31.108 | Aromandendrene | C15H24 | 204.35 | 0.56 | Antieczematic, Dermatologic, Antineoplastic, Ovulation inhibitor, Testosterone 17beta-dehydrogenase (NADP+) inhibitor |
| 7 | 33.17 | Cyclopentane-3′-spirotricyclo[3.1.0.0(2,4)]he | C14H20 | 188.31 | 1.75 | Testosterone 17beta-dehydrogenase (NADP+) inhibitor,Nicotinic alpha6beta3beta4alpha5 receptor antagonist |
| 8 | 33.287 | 1,3-Bis-(2-cyclopropyl,2-methylcyclopropyl)- | C18H26O | 258.4 | 1.38 | Testosterone 17beta-dehydrogenase (NADP+) inhibitor, CYP2J substrate, Antieczematic, Acylcarnitine hydrolase inhibitor, Dermatologic, Beta glucuronidase inhibitor |
| 9 | 34.358-35.761 | 1,8-Cyclopentadecadiyne | C15H22 | 202.33 | 62.76 | Steroid DELTA-isomerase inhibitor, Aspulvinonedimethylallyltransferase inhibitor, Nicotinic alpha6beta3beta4alpha5 receptor antagonist, Testosterone 17beta-dehydrogenase (NADP+) inhibitor , Antineoplastic |
| 12 | 36.164 | Bicyclo[3.1.0]hexane-6-methanol, 2-hydroxy. | C10H18O2 | 170.25 | 0.88 | Alkenyl glycerophospho-choline hydrolase inhibitor, Alkylacetylglycerophosphatase inhibitor, Acylcarnitine hydrolase inhibitor, |
| 13 | 36.873 | .alpha.-Bisabolol | C15H26O | 222.37 | 2.42 | Apoptosis agonist, Antieczematic, CYP2J substrate, Retinol dehydrogenase inhibitor, Prenyl-diphosphatase inhibitor, |
| 14 | 37.487 | .alpha.-Bisabolol | C15H26O | 222.37 | 3.35 | Testosterone 17beta-dehydrogenase (NADP+)inhibitor,Nicotinic alpha6beta3beta4alpha5
receptor antagonist |
| 15 | 41.182 | N1-Benzyl-N2(bezylidenyl-benzylamino)-ben | C20H18N2O2 | 318.4 | 20.48 | CYP2C12 substrate, Calcium channel (voltage-sensitive) activator |
In silico studies
Physiochemical properties and drug likeness
The molecule satisfied Lipinski rule of five LogP (4.298), number of donors, acceptors, rotatable bonds are 0 with Molecule Weight of 202.341 and surface area of 94.841 such that the molecule is considered as a drug. The Lipinski rule of the molecule illustrated in Fig 1a & 1b.
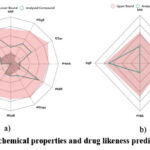 |
Figure 1: Physiochemical properties and drug likeness prediction by pKSCM |
Molecular docking
The compound 1,8-cyclopentadecadiyne was shown to be deeply lodged in the 6MS7 active site, as demonstrated by the molecular docking analysis. An alkyl interaction was also discovered, indicating the drug’s increased stability and propensity for binding through the receptor. The protein 6MS7’s amino acids LEU473 and LYS457 form an alkyl bond with the ligand at 3.69 and 4.29 Å, respectively, and the slip index is -5.79. Fig 2(a) & Fig 2(b) indicate 3D images of ligand (1,8-Cyclopentadecadiyne) and protein (PDB ID: 6MS7) interaction with distances (given in Table 4). Fig 3(b) represents the 2D interaction of ligand and neighboring amino acids.
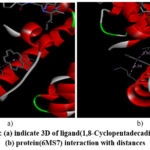 |
Figure 2: (a) indicate 3D of ligand(1,8-Cyclopentadecadiyne) and (b) protein(6MS7) interaction with distances |
The Ramachandran plot describing whether amino acid residues are in the “accepted region” or the “unaccepted” field was created using Drug Discovery Studio 4.1. Figure 5a depicts a Ramchandran plot, while Figure 5b depicts a two-dimensional interaction between ligand and adjacent amino acids (PDB ID:6MS7). A superior 3D protein modal should have more than 90% amino acid residues in the favourable quadrant of the Ramachandran plot.
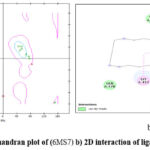 |
Figure 3: a) Ramchandran plot of (6MS7) b) 2D interaction of ligand and neighboring |
The molecular docking analysis revealed that the molecule 1,8-Cyclopentadecadiyne was profoundly concealed in the active locale of the 1OH0, alkyl interaction, indicating increased stability as well as the molecule’s affinity for binding to the receptor. TRP 92 of protein 1OH0 forms an alkyl bond with the ligand at 4.88 Å. Fig 4a indicate 3D of ligand and protein (PDB ID: 1OH0) interaction with distances and Fig 4b represents the 2D interaction of ligand and receptor amino acids.
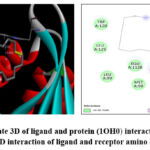 |
Figure 4: a) indicate 3D of ligand and protein (1OH0) interaction with distances b) 2D interaction of ligand and receptor amino acids |
The compound 1,8-Cyclopentadecadiyne was shown to be deeply buried in the 2J6M active site, as demonstrated by the molecular docking analysis. An alkyl interaction was identified, indicating the drug’s increased stability and affinity for binding through the receptor. The protein 2J6M’s amino acid PHE 910 and the ligand form a single alkyl bond that is 4.45 Å. Fig 5a indicate 3D of ligand and protein (PDB ID: 2J6M) complex with distances and Fig 5b represents the 2D interaction of the complex.
Table 4: Representing molecular docking results of 1,8 cyclopentadiene with various targets.
| Targets (PDB ID) | Distance (Å) | Pi-Alkyl interactions | Neighbouring aminoacids | Docking score |
| Antiproliteration PDB ID: 6MS7 | 3.69 & 4.29 | LEU465; LYS457 | TYR473, LEU453, MET463, GLN470 | -7.6 |
| Steroid DELTA-isomerase PDB ID: 1OH0 | 4.88 | TRP A:92 | TRP120, LEU125, LEU99, MET90, CYS97, VAL91 | -6.5 |
| EGFR Kinase PDB ID:2J6M | 4.45 | PHE A:910 | GLN982, ILE981, ASP807, LYS806, PRO937, | -6.5 |
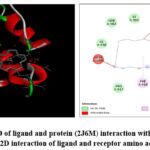 |
Figure 5: a) 3D of ligand and protein (2J6M) interaction with distances and b) 2D interaction of ligand and receptor amino acids |
Target identification results predicted that the molecule possess anti-proliferative that might be used to inhibit cell proliferation helpful to treat cancer. Steroid DELTA-isomerize inhibitor target was identified by PASS online target prediction The inhibition of the enzyme results prevents the oxidative conversion of ketosteroids that are responsible for synthesis of hormonal steroids. Based on the above two targets identified it was assumed that it might be effective against cancer. To identify its effectiveness against cancer pdCSM-cancer prediction tool was used that evaluates the structural features against multiple cell lines. Amongst them the molecule is active in contradiction of cell line MDA_MB_468 (breast) having a highest value of 5.98. Molecular docking studies were carried out against three protein targets 6MS7-antiproliferative, 1OH0- Steroid DELTA-isomerize, 2J6M- breast cancer cell line MDA_MB_468. 6MS7 target showed highest value among the three of about -7.6 depicting the molecule is effective against proliferation.
pdCSM-cancer
PdCSM-cancer, which successfully predicts compounds to be expected to be effective against multiple cancer cell lines by using a plot-based signature representation of a small molecule’s chemical structure. The results predicted that the target compound is active against cancer effective against various cancer cell lines and was found highest for breast cancer cell lines breast MDA_MB_468. Figure 6 represents the graph illustrating the activity of the molecule 1,8-cyclopentadiene on different cell lines.
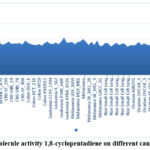 |
Figure 6: Molecule activity 1,8-cyclopentadiene on different cancer cell lines. |
PiscesCSM:
A total of 7932 molecules were found to possess both antagonist and synergistic properties in the treatment of breast cancer. 2524 molecules were found to be synergistic and remaining 5410 molecules were antagonistic. Based on mol log p values the molecules with synergistic effect values ranged from 12.6058 to -9.9602.the highest value is with beta-carotene and second was squalene. Antagonistic value ranged from 10.574 to -8.72. Anchovixanthin, cis-lutein. Fig. 7 represents the pic chart with molecules having synergistic and antagonist molecules.
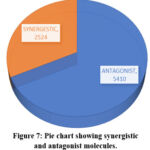 |
Figure 7: Pie chart showing synergistic and antagonist molecules. |
Toxicity prediction using GUSAR model
The Generally Unrestricted Structure Activity Relationship (GUSAR) model for acute rat toxicity was employed to forecast the LD50 values for each lead molecule delivered by subcutaneous (SC), intraperitoneal (IP), oral, and intramuscular (IV) routes.Fig 8 graph represents acute toxicity prediction in rats using GUSAR model.
The molecule toxicity based on route of administration was evaluated using GUSAR model that indicated that the molecule is less toxic and belongs to IP Class 5, IV Class 3, Oral Class 4, and SC Class 3. Table 5 represents GUSAR model dosage form with LD50, classification and class of toxicity prediction in rats.
Table 5: GUSAR model with dosage form, LD50, toxicity classification prediction in rats
| Route of administration | IP | IV | Oral | SC |
| Class | 5 | 3 | 4 | 3 |
| LD50 (mg/kg) | 565,100 | 37,810 | 1708,000 | 82,350 |
| LD50 Log10(mmol/kg) | 0,446 | -0,728 | 0,926 | -0,390 |
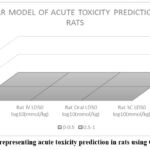 |
Figure 8: Graph representing acute toxicity prediction in rats using GUSAR model. |
Discussion
The investigations on the roots of Saussurea lappa indicated the presence of various phytochemicals such as alkaloids, flavonoids, sterols, triterpenoids, and resins, all of which are associated with various biological activities and medicinal properties. Some of the documented biological activities of the constituents obtained from the GC-MS analysis such as 1,8-Cyclopentadecadiyne, Aromandendrene, and Bisabolol, include anticancer, anti-inflammatory and other dermatological activities. 1,8-Cyclopentadecadiyne was particularly prominent in both the polar and non-polar extracts, reinforcing its position as the major active compound of the extracts.
Comparative in silico studies suggest 1,8-Cyclopentadecadiyne also has a potential inhibitory effect on antiproliferative proteins (6MS7), Steroid Delta-isomerase (1OH0), and EGFR kinase (2J6M) which based on expression analysis in documented databases are associated with the breast cancer cell line MDA-MB-468. The strong docking score (-7.6) against 6MS7, correlating with one of its predicted activities against breast cancer cell line MDA-MB-468, further supports these claims and highlights the potential role of this compound in inhibiting cancer cell proliferation.
Toxicity prediction using the GUSAR tool also demonstrated acceptable safety margins with classification from Class 3-5 for different modes of administration indicating moderate to low acute toxicity
This is still an excerpt, and substantiating, particularly 1, 8-Cyclopentadecadiyne, demonstrates potential for anticancer agent formulation, albeit needing in vitro and in vivo studies to authenticate such claims.
Conclusion
This study stresses the possibilities of Saussurea lappa root extracts as a viable source of bioactive chemicals with anticancer activity in breast cancer. Phytochemical analysis verified the existence of important secondary metabolites with established medicinal uses, including terpenoids, alkaloids, flavonoids, and phenolic compounds. GC-MS profiling revealed a diverse range of phytoconstituents, several of which have been previously associated with cytotoxic and anti-proliferative properties.
Overall, the GC-MS study of the S. lappa aqueous educe indicates that it contains a variety of phytoconstituents, with 1,8-cyclopentadecadiene showing particular interest due to its exceptional affinity for binding to enzymes linked to cancer (e.g., proliferation, steroid DELTA-isomerize, and EGFR) as determined by docking studies and the PASS evaluation approach. These results demonstrate the phytochemicals under study’s considerable promise. 1,8-cyclopentadecadiene exhibits a significant binding affinity to tumour proteins through alkyl interactions, according to molecular docking findings. In light of this, 1,8-cyclopentadecadiene appears to be a viable option for future investigation as an anticancer medication.
Acknowledgement
The author would like to thank KL University for providing research facilities. We also thanks to the accurate labs, Vijayawada for providing GC-MS facilities.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest: The author(s) do not have any conflict of interest.
Data Availability Statement This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required
Clinical Trial Registration
This research does not involve any clinical trials
Permission to Reproduce Sources
Not applicable
Authors’ Contribution
- Reehana Shaik’s contributions include conceptualization, literature review, experimental design, data analysis, and article drafting.
- Ishwarya Obilineni performs GC-MS profiling, data interpretation, software analysis, and figure production.
- Naga Venkata Chenchu Lakshmi Kollipara is responsible for supervision, project administration, critical manuscript revision, and correspondence.
- K Ravishankar: In-silico evaluation, molecular docking research, and data curation.
- Saipraneeth Tripuraneni: Toxicity prediction analysis, result validation, proofreading, and formatting.
References
- Waks AG, Winer EP. Breast cancer treatment: a review. JAMA. 2019;321(3):288-300.
CrossRef - Lehmann BD, Bauer JA, Chen X, et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest. 2011;121(7):2750-2767.
CrossRef - Cragg GM, Pezzuto JM. Natural products as a vital source for the discovery of cancer chemotherapeutic and chemopreventive agents. Med Princ Pract. 2016;25(Suppl. 2):41-59.
CrossRef - Pandey A, Chowdary VM, Mal BC. Identification of critical erosion prone areas in the small agricultural watershed using USLE, GIS and remote sensing. Water Resour Manage. 2007;21:729-746.
CrossRef - Shah SC, Colombel JF, Sands BE, Narula N. Mucosal healing is associated with improved long-term outcomes of patients with ulcerative colitis: a systematic review and meta-analysis. Clin Gastroenterol Hepatol. 2016;14(9):1245-1255.
CrossRef - Cho JY, Kim AR, Jung JH, et al. Cytotoxic and pro-apoptotic activities of cynaropicrin, a sesquiterpene lactone, on the viability of leukocyte cancer cell lines. Eur J Pharmacol. 2004;492(2-3):85-94.
CrossRef - Yang DS, Peng WB, Yang YP, Liu KC, Li XL, Xiao WL. Cytotoxic prenylated flavonoids from Macaranga indica. Fitoterapia. 2015;103:187-191.
CrossRef - Liu XN, Li HM, Wang SP, Zhang JZ, Liu DL. Sesquiterpene lactones of Aucklandia lappa: Pharmacology, pharmacokinetics, toxicity, and structure–activity relationship. Chin Herb Med. 2021;13(2):167-176.
CrossRef - Ali SI, Venkatesalu V. Botany, traditional uses, phytochemistry and pharmacological properties of Saussurea costus–An endangered plant from Himalaya–A review. Phytochem Lett. 2022;47:140-155.
CrossRef - Ali S, Ahmed D. Comparing DES-mediated ultrasound- and heat-assisted extraction of bioactive metabolites from Saussurea lappa and optimization by RSM and validation studies. Green Anal Chem. 2023;7:100080.
CrossRef - Chang KM, Kim GH. Comparison of volatile aroma components from Saussurea lappa CB Clarke root oils. Prev Nutr Food Sci. 2008;13(2):128-133.
CrossRef - Kumar A, Agnihotri VK. NMR based profiling of sesquiterpene lactones in Saussurea lappa roots collected from different location of Western Himalaya. Nat Prod Res. 2022;36(2):621-624.
CrossRef - Nadda RK, Ali A, Goyal RC, Khosla PK, Goyal R. Aucklandia costus (syn. Saussurea costus): Ethnopharmacology of an endangered medicinal plant of the Himalayan region. J Ethnopharmacol. 2020;263:113199.
CrossRef - Singh R, Chahal KK, Singla N. Chemical composition and pharmacological activities of Saussurea lappa: A review. J Pharmacogn Phytochem. 2017;6(4):1298-1308.
- Hassan R, Masoodi MH. Saussurea lappa: A comprehensive review on its pharmacological activity and phytochemistry. Curr Tradit Med. 2020;6(1):13-23.
CrossRef - Alaagib RM, Ayoub SM. On the chemical composition and antibacterial activity of Saussurea lappa (Asteraceae). Pharma Innov. 2015;4(2, Part C):73.
- Naseer S, Iqbal J, Naseer A, et al. Deciphering chemical profiling, pharmacological responses and potential bioactive constituents of Saussurea lappa Extracts through in vitro approaches. Saudi J Biol Sci. 2022;29(3):1355-1366.
CrossRef - Jeong SJ, Itokawa T, Shibuya M, et al. Costunolide, a sesquiterpene lactone from Saussurea lappa, inhibits the VEGFR KDR/Flk-1 signaling pathway. Cancer Lett. 2002;187(1-2):129-133.
CrossRef - Mishra AP, Nigam M, Devkota HP, Matsabisa MG. Dolomiaea costus (Falc.) Kasana and AK Pandey: Ethnomedicinal uses, bioactive chemical constituents, and pharmacological activities. In: Medicinal Plants of the Asteraceae Family: Traditional Uses, Phytochemistry and Pharmacological Activities. Singapore: Springer Nature Singapore; 2022:153-172.
CrossRef - Butturini E, Butera G, Pacchiana R, et al. Redox sensitive cysteine residues as crucial regulators of wild-type and mutant p53 isoforms. Cells. 2021;10(11):3149.
CrossRef - Fiehn O. Metabolomics by gas chromatography–mass spectrometry: Combined targeted and untargeted profiling. Curr Protoc Mol Biol. 2016;114(1):30-4.
CrossRef - Adelusi TI, Oyedele AQ, Boyenle ID, et al. Molecular modeling in drug discovery. Inform Med Unlocked. 2022;29:100880.
CrossRef - Chen SL, Yu H, Luo HM, et al. Conservation and sustainable use of medicinal plants: problems, progress, and prospects. Chin Med. 2016;11:1-10.
CrossRef - Al-Adhroey AH, Al-Abhar YM, Noman NM, Al-Mekhlafi HM. Ethnopharmacological survey of herbal remedies used in Yemen for treating malaria. J Herb Med. 2020;21:100344.
CrossRef - Saleem TM, Lokanath N, Prasanthi A, et al. Aqueous extract of Saussurea lappa root ameliorate oxidative myocardial injury induced by isoproterenol in rats. J Adv Pharm Technol Res. 2013;4(2):94-100.
CrossRef - Mujeeb F, Bajpai P, Pathak N. Phytochemical evaluation, antimicrobial activity, and determination of bioactive components from leaves of Aegle marmelos. Biomed Res Int. 2014;2014:497606.
CrossRef - Farag RS, Abdel-Latif MS, Abd El Baky HH, Tawfeek LS. Phytochemical screening and antioxidant activity of some medicinal plants’ crude juices. Biotechnol Rep. 2020;28:e00536.
CrossRef - Soni A, Sosa S. Phytochemical analysis and free radical scavenging potential of herbal and medicinal plant extracts. J Pharmacogn Phytochem. 2013;2(4):22-29.
- Mastovska K, Lehotay SJ. Rapid sample preparation method for LC−MS/MS or GC−MS analysis of acrylamide in various food matrices. J Agric Food Chem. 2006;54(19):7001-7008.
CrossRef - Jung MY, Kim GB, Jang ES, et al. Improved extraction method with hexane for gas chromatographic analysis of conjugated linoleic acids. J Dairy Sci. 2006;89(1):90-94.
CrossRef - Wallace WE, Moorthy AS. NIST Mass Spectrometry Data Center standard reference libraries and software tools: Application to seized drug analysis. J Forensic Sci. 2023;68(5):1484-1493.
CrossRef - Lohohola PO, Mbala BM, Bambi SM, et al. In silico ADME/T properties of quinine derivatives using SwissADME and pkCSM webservers. Int J Trop Dis Health. 2021;42(11):1-2.
CrossRef - Lipinski CA. Lead-and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol. 2004;1(4):337-341.
CrossRef - Gfeller D, Grosdidier A, Wirth M, et al. SwissTargetPrediction: a web server for target prediction of bioactive small molecules. Nucleic Acids Res. 2014;42(W1):W32-W38.
CrossRef - Tousif MI, Saleem M, Tauseef S, et al. Unrivalled insight into potential biopharmaceutical application of Saussurea albescens (DC.) Sch. Bip.: Chemical profiling, antioxidant and enzyme inhibition activities and in silico studies. Biocatal Agric Biotechnol. 2023;54:102964.
CrossRef - Baroroh U, Biotek M, Muscifa ZS, et al. Molecular interaction analysis and visualization of protein-ligand docking using Biovia Discovery Studio Visualizer. Indones J Comput Biol. 2023;2(1):22-30.
CrossRef - Lagunin A, Stepanchikova A, Filimonov D, Poroikov V. PASS: prediction of activity spectra for biologically active substances. Bioinformatics. 2000;16(8):747-748.
CrossRef - Daina A, Michielin O, Zoete V. SwissTargetPrediction: updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res. 2019;47(W1):W357-W364.
CrossRef - Jamil F, Raheel S, Rasheed H. Docking of smaller ligands in the heme pocket of Hell’s Gate Globin IV. Pak J Zool. 2018;50(4).
CrossRef - Al-Jarf R, de Sá AG, Pires DE, Ascher DB. pdCSM-cancer: using graph-based signatures to identify small molecules with anticancer properties. J Chem Inf Model. 2021;61(7):3314-3322.
CrossRef - AlJarf R, Rodrigues CH, Myung Y, et al. piscesCSM: prediction of anticancer synergistic drug combinations. J Cheminform. 2024;16(1):81.
CrossRef - Lagunin A, Zakharov A, Filimonov D, Poroikov V. QSAR modelling of rat acute toxicity on the basis of PASS prediction. Mol Inform. 2011;30(2‐3):241-250.
CrossRef - Erhirhie EO, Ihekwereme CP, Ilodigwe EE. Advances in acute toxicity testing: strengths, weaknesses and regulatory acceptance. Interdiscip Toxicol. 2018;11(1):5-12.
CrossRef - Hamidi MR, Jovanova B, Panovska TK. Toxicological evaluation of the plant products using Brine Shrimp (Artemia salina) model. Maced Pharm Bull. 2014;60(1).
CrossRef







