Manuscript accepted on :26-08-2025
Published online on: 08-09-2025
Plagiarism Check: Yes
Reviewed by: Dr. Ameer Ali Shakr Hadi and Dr. Shail Modi
Second Review by: Dr. Hind Shakir
Final Approval by: Dr. Mariia Shanaida
Preeti Chaudhary* , Akash Kumavat
, Akash Kumavat , Pritam Khandave
, Pritam Khandave , Rupesh Pingale
, Rupesh Pingale , Sakshi Singh
, Sakshi Singh , Omkar Dudhal
, Omkar Dudhal , Sonali Korlekar
, Sonali Korlekar and Shubham Tivlekar
and Shubham Tivlekar
Department of Pharmacy, NCRD's Sterling Institute of Pharmacy, Nerul, Navi Mumbai, Maharashtra, India
Corresponding Author E-mail: preeti.chaudhary@ncrdsip.com
DOI : https://dx.doi.org/10.13005/bpj/3248
Abstract
Anti-inflammatory agents are essential for managing inflammation-driven diseases, with COX-2 (cyclooxygenase-2) and NF-κB (nuclear factor kappa B) serving as key molecular targets. Vindoline, a phytochemical derived from Catharanthus roseus, has demonstrated potential pharmacological properties, including anti-inflammatory activity. This study aims to evaluate the binding affinity and interactions of vindoline with COX-2 and NF-κB through molecular docking studies using AutoDock Vina. Diclofenac sodium was used as the reference standard. The 3D structure of vindoline was obtained from PubChem, and protein structures of COX-2 (PDB ID: 6COX) and NF-κB (PDB ID: 1NFK) were retrieved from the Protein Data Bank. Molecular docking was performed using AutoDock Vina to analyze binding affinities, and protein-ligand interactions were visualized using BIOVIA Discovery Studio 2025. ADME/T analysis was performed using SwissDock and Protox 3.0. The results demonstrated strong binding interactions of vindoline with COX-2 (-9.4 kcal/mol) and NF-κB (-7.0 kcal/mol) compared to Diclofenac Sodium (-8.0 kcal/mol and -6.4 kcal/mol, respectively). Key interactions, including hydrogen bonding and hydrophobic contacts at the active sites, supported its potential inhibitory activity. The results of egg albumin denaturation assay revealed that Vindoline (50–250 µg/mL) significantly inhibits protein denaturation and exhibits superior inhibition compared to Diclofenac Sodium. Vindoline also demonstrated high solubility and a potentially safer profile than Diclofenac Sodium. It suggests its potential as a lead compound for the development of anti-inflammatory drugs.
Keywords
ADME/T Analysis; AutoDock Vina; Cyclooxygenase-2; Egg Albumin Denaturation Assay; Molecular Docking; Nuclear Factor Kappa B; Vindoline
Download this article as:| Copy the following to cite this article: Chaudhary P, Kumavat A, Khandave P, Pingale R, Singh S, Dudhal O, Korlekar S, Tivlekar S. In-Silico and In-Vitro Evaluation of Vindoline as a Potential Anti-Inflammatory Agent Targeting COX-2 and NF-κB. Biomed Pharmacol J 2025;18(3). |
| Copy the following to cite this URL: Chaudhary P, Kumavat A, Khandave P, Pingale R, Singh S, Dudhal O, Korlekar S, Tivlekar S. In-Silico and In-Vitro Evaluation of Vindoline as a Potential Anti-Inflammatory Agent Targeting COX-2 and NF-κB. Biomed Pharmacol J 2025;18(3). Available from: https://bit.ly/41GjVtF |
Introduction
Inflammation is a multifaceted biological response triggered by harmful agents such as pathogens, injured cells, or irritants.1,2 While short-term (acute) inflammation plays a crucial role in defending the body and promoting tissue healing, 3,4 prolonged (chronic) inflammation is associated with the development of numerous health conditions, including rheumatoid arthritis, heart disease, neurodegenerative disorders, and cancer.5,6,7 Key molecular mediators in the inflammatory cascade include cyclooxygenase-2 (COX-2) and nuclear factor kappa B (NF-κB).8,9 COX-2, an inducible enzyme, catalyzes the conversion of arachidonic acid to pro-inflammatory prostaglandins,10,11 while the transcription factor NF-κB plays a key role in controlling the activity of genes that govern inflammatory responses, immune function, and cell survival.12,13 These molecules have emerged as critical therapeutic targets for the development of anti-inflammatory drugs.
Vindoline, an alkaloid derived from Catharanthus roseus (commonly known as the Madagascar periwinkle), is one of the key phytoconstituents of this medicinal plant, renowned for its diverse therapeutic properties.14,15 Catharanthus roseus has long been utilized in traditional medicine, and its alkaloids, such as vincristine and vinblastine, are well-established chemotherapeutic agents.16,17 Vindoline, though less explored, has shown potential pharmacological activities, including anti-inflammatory, antioxidant, and anticancer effects.18,19 Given its promising biological properties, investigating vindoline as a potential inhibitor of COX-2 and NF-κB offers a compelling avenue for drug discovery. Molecular docking has revolutionized drug discovery by facilitating the virtual screening of compounds and identifying key protein-ligand interactions.20
This study aims to evaluate the anti-inflammatory potential of vindoline through in-silico molecular docking studies and in-vitro study. By evaluating the binding affinities and interactions of vindoline with COX-2 and NF-κB, this research aims to provide insights into the mechanism of action of vindoline and establish its potential as a lead compound for anti-inflammatory drug development.
Materials and Methods
Software and Tools
For in-silico evaluation of Vindoline as a potential anti-inflammatory agent targeting COX-2 and NF-κB the following software, tools and applications were used: PubChem, RCSB Protein Data Bank (PDB), AutoDock MGL Tools, AutoDock Vina, BIOVIA Discovery Studio 2025, SwissADME Online Tool, and Protox 3.0 Online Tool.
In-Silico Anti-Inflammatory Activity
Stepwise Procedure for Molecular Docking
Molecular docking of vindoline with 6COX (Cyclooxygenase-2, COX-2) and 1NFK (Nuclear factor-kappaB, NF-κB) involved several key steps, including protein preparation, ligand preparation, docking, and visualization.
Step 1: Preparation of Protein (Target Molecules)
The protein receptor and ligand molecules were selected based on their relevance to the study. Two proteins Cyclooxygenase-2 (COX-2) and Nuclear Factor-kappaB (NF-κB) were chosen for this study.
Retrieval of the Protein Structure
The three-dimensional structures of COX-2 (PDB ID: 6COX) and NF-κB (PDB ID: 1NFK) were retrieved from the RCSB Protein Data Bank, accessible at https://www.rcsb.org/.21 The corresponding PDB files were downloaded and used for docking analysis (Table 1).
Table 1: Target Proteins and their Respective Functions
| Proteins | Protein Description | Protein Function | PDB ID |
| COX-2 | Cyclooxygenase-2 | Inducible enzyme that converts arachidonic acid into pro-inflammatory prostaglandins (e.g., PGE₂), leading to pain, fever, and swelling. Upregulated in response to cytokines and stress.22,23 | 6COX |
| NF-κB | Nuclear factor-kappaB | A transcription factor responsible for controlling the expression of pro-inflammatory cytokines such as TNF-α, IL-1β, and IL-6, chemokines, and adhesion molecules. Activates immune responses and sustains chronic inflammation. Also induces COX-2 expression, amplifying inflammation.24 | 1NFK |
Remove Unnecessary Molecules and Prepare the Protein for Docking:
The protein structures (6COX.pdb and 1NFK.pdb) were loaded in AutoDock Tools. Water molecules were removed (Edit → Delete Water). Polar hydrogens were added (Edit → Hydrogens → Add → Polar only). Kollman charges were assigned (Edit → Charges → Add Kollman Charges). The modified protein structures were then saved in pdbqt format as 6COX_clean.pdbqt and 1NFK_clean.pdbqt.25,26
Step 2: Preparation of Ligand (Vindoline)
Obtain the Ligand Structure:
The 3D structures of Vindoline (PubChem ID: 260535) and Diclofenac Sodium (PubChem ID: 5018304) were sourced from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/).27 The structure was initially downloaded in SDF format and later converted to PDB format using Discovery Studio 2025 (Figure 1).
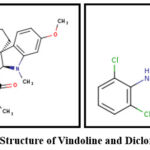 |
Figure 1: 2D Structure of Vindoline and Diclofenac Sodium |
Prepare Ligand for Docking (Using AutoDock Tools)
AutoDock Tools (ADT) was used for ligand preparation. The vindoline.pdb file was loaded into the software, hydrogens were added, and Gasteiger charges were assigned. The processed structure was then converted and saved in PDBQT format as vindoline.pdbqt.
Step 3: Docking Procedure
AutoDock Vina was used for molecular docking simulations to predict the binding affinity and orientation of Vindoline and Diclofenac sodium within the active site of COX-2 and NF-κB. This software facilitates the identification of potential lead compounds, enabling the rational design of novel inhibitors with improved efficacy.28,29,30
Grid Box Preparation (Defining Binding Site)
The prepared protein structures (6COX_clean.pdbqt or 1NFK_clean.pdbqt) and the ligand (vindoline.pdbqt) were loaded into the software (Figure 2). The grid box was set (Grid → Set Grid Box) and centered around the active site of COX-2 or NF-κB. The x, y, and z dimensions (x=47.061, y = 25.506, z = 36.889) were adjusted to encompass the binding site, and default parameters in AutoDock Vina were used for the simulations. The configuration file was then saved as config.txt.
 |
Figure 2: Import of Protein and Ligand in AutoDock Vina |
Docking Execution
The docking simulation was carried out in the Command Prompt. The working directory containing vina.exe was accessed using the command cd C:\path\to\AutoDockVina. Docking was executed with the following command: vina.exe –receptor 6COX.pdbqt –ligand vindoline.pdbqt –config config.txt –log vina_log.txt –out docked_output.pdbqt. The configuration file (config.txt) defined docking parameters, including grid box coordinates, exhaustiveness, and center position. After completion, the vina_log.txt file was analyzed to obtain binding affinity scores.31
Post-Docking Analysis
The binding affinity scores (in kcal/mol) were extracted from the vina_log.txt file and protein-ligand interactions were visualized to identify hydrogen bonds and hydrophobic interactions. The binding affinity scores of Vindoline were compared with those of the Standard Ligand (Diclofenac Sodium).
Step 4: Visualization in Discovery Studio 2025:
Molecular interactions were analyzed using BIOVIA Discovery Studio 2025. The prepared protein structure (6COX_clean.pdbqt or 1NFK_clean.pdbqt) and the docked ligand (docked_output.pdbqt) were loaded into the software. Ligand binding was examined through the Ligand-protein Interactions tool. Hydrogen bonding sites were identified via Receptor-Ligand Interactions. Additionally, 2D and 3D interaction maps were generated. High-quality images for publication were saved.32
ADME/T Analysis
The ADME analysis was performed using the SwissADME online server. Lipinski’s rule of five, ADME/T (Absorption, Distribution, Metabolism, Excretion and Toxicity), Drug-likeness properties, pharmacokinetics, and physicochemical properties of Vindoline and Standard Diclofenac Sodium were evaluated. Toxicity data for both drugs were obtained from the Protox 3.0 online server.33,34
In-Vitro Anti-Inflammatory Activity
Egg Albumin Denaturation Assay
This assay evaluates the anti-inflammatory potential of test compounds by determining their ability to prevent or inhibit the denaturation of egg albumin under specific conditions. Inflammation can cause denaturation of proteins. Denaturation is the process in which a protein loses its biological activity and undergoes structural changes.35 In this assay, egg albumin was used as a model protein, and denaturation was induced by exposing it to extreme conditions such as heat, pH variations, or other denaturing agents. This structural alteration resulted in changes to its physical properties and a loss of functional activity.36 Inflammation is often associated with protein denaturation, substances with anti-inflammatory properties are believed to stabilize protein structures and inhibit denaturation.37
Collection and Preparation of 1% Egg Albumin Solution
A fresh hen’s egg was purchased from the local market, and the egg white (albumin) was carefully separated into a beaker without disturbing the yolk. To prepare a 1% egg albumin solution, 1 ml of the translucent egg white was transferred into a 100 ml volumetric flask. Cold-distilled water was then added to bring the volume to 100 ml. The solution was thoroughly stirred with a magnetic stirrer to ensure uniform mixing.
Procedure of Egg Albumin Assay:
The anti-inflammatory activity of isolated Vindoline was evaluated in vitro by assessing its ability to inhibit egg albumin denaturation.
A reaction mixture was prepared by combining 0.2 mL of a 1% egg albumin solution with 2 ml of either Vindoline or the reference drug, diclofenac sodium, at varying concentrations ranging from 50 to 250 μg/ml. This was followed by the addition of 2.8 ml of phosphate-buffered saline (PBS) at pH 7.4, resulting in a final volume of 5 ml. The pH was carefully adjusted to 7.4 using a minimal volume of 1N HCl. A control solution was also prepared by mixing 2 ml of triple-distilled water with 0.2 ml of the 1% egg albumin and 2.8 ml of PBS. All mixtures were incubated at 37 ± 2°C for 30 minutes and then subjected to heating in a water bath at 70 ± 2°C for 15 minutes. Once cooled, the absorbance was recorded at 660 nm using a UV/Vis spectrophotometer, with triple-distilled water serving as the blank. The percentage inhibition of protein denaturation was calculated using the following formula.

Statistical Analysis
All statistical analyses were conducted using Microsoft Excel. The results are presented as the mean ± standard error of the mean (SEM). Differences between groups were assessed using one-way ANOVA (Single Factor). P-values were calculated for each comparison, and a p-value less than 0.05 was considered statistically significant.
Results
The molecular docking analysis was performed to evaluate the binding affinity and interaction of Vindoline with Cyclooxygenase-2 (6COX) and Nuclear Factor-kappa B (1NFK) in comparison with Diclofenac Sodium as a standard. Against 6COX, Vindoline exhibited a higher binding affinity (-9.4 kcal/mol) than Diclofenac Sodium (-8.0 kcal/mol), indicating a stronger interaction with the target (Table 2).
Table 2: Binding Affinity of Vindoline and Diclofenac Sodium with 6COX (Cyclooxygenase-2)
| Molecule Name | Binding Affinity (kcal/mol) | Hydrogen Bond Interactions (Protein Residues) | Bond Length (Aº) | Amino Acid Residue Interactions (Hydrophobic / Pi-Cation / Pi-Anion / Pi-Alkyl) |
| Vindoline
(Test Compound) |
-9.4 | B: LEU224 | 3.57 | A: PHE142, A: LEU145, B: LEU145, B: PHE142 |
| B: GLN374 | 3.78 | |||
| A: ASN375 | 3.49 | |||
| Diclofenac Sodium
(Standard) |
-8.0 | B: SER143 | 3.03 | A: LEU238 |
| B: ASN144 | 3.15 | |||
| B: GLU140 | 2.01 | |||
| A: GLU236 | 1.76 |
Vindoline formed hydrogen bonds with residues B: LEU224 (3.57 Å), B: GLN374 (3.78 Å), and A: ASN375 (3.49 Å), along with hydrophobic interactions involving A: PHE142, A: LEU145, B: LEU145, and B: PHE142 (Figure 3).
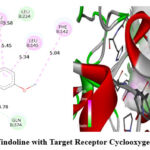 |
Figure 3: Interaction of Vindoline with Target Receptor Cyclooxygenase-2 (PDB ID- 6COX). |
In contrast, Diclofenac Sodium exhibited hydrogen bonding with B: SER143 (3.03 Å), B: ASN144 (3.15 Å), B: GLU140 (2.01 Å), and A: GLU236 (1.76 Å), with hydrophobic interaction involving A: LEU238 (Figure 4).
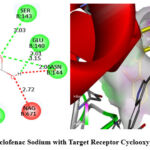 |
Figure 4: Interaction of Diclofenac Sodium with Target Receptor Cyclooxygenase-2 (PDB ID- 6COX) |
Similarly, docking results with 1NFK revealed that Vindoline had a binding affinity of -7.0 kcal/mol, which was higher than Diclofenac Sodium (-6.4 kcal/mol) (Table 3). Hydrogen bonding interactions for Vindoline were observed with A: ARG54 (3.60 Å), B: SER72 (2.30 Å), and B: GLY66 (3.57 Å), alongside hydrophobic interactions with B: ARG56, B: PHE53, and C: DG2 (Figure 5). Diclofenac Sodium, on the other hand, formed hydrogen bonds with A: ASN136 (2.16 Å, 2.72 Å), A: GLY66 (2.11 Å), and A: LEU67 (2.06 Å), with additional hydrophobic interactions with A: LYS77 and A: PRO68 (Figure 6).
Table 3: Binding Affinity of Vindoline and Diclofenac Sodium with 1NFK (Nuclear factor-kappaB)
| Molecule Name | Binding Affinity (kcal/mol) | Hydrogen Bond Interactions (Protein Residues) | Bond Length (Aº) | Amino Acid Residue Interactions (Hydrophobic / Pi-Cation / Pi-Anion / Pi-Alkyl) |
| Vindoline
(Test Compound) |
-7.0 | A: ARG54 | 3.60 | B: ARG56, B: PHE53, C: DG2 |
| B: SER72 | 2.30 | |||
| B: GLY66 | 3.57 | |||
| Diclofenac Sodium
(Standard) |
-6.4 | A: ASN136 | 2.16 | A: LYS77, A: PRO68 |
| A: ASN136 | 2.72 | |||
| A: GLY66 | 2.11 | |||
| A: LEU67 | 2.06 |
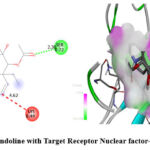 |
Figure 5: Interaction of Vindoline with Target Receptor Nuclear factor-kappaB (PDB ID- 1NFK) |
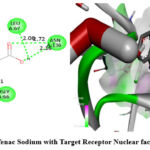 |
Figure 6: Interaction of Diclofenac Sodium with Target Receptor Nuclear factor-kappaB (PDB ID- 1NFK) |
The overall results suggest that Vindoline exhibits a stronger binding affinity and favorable interactions with both targets (COX-2 and NF-κB), potentially indicating its role as a promising anti-inflammatory agent. Figures 3-6 illustrate the binding interactions of Vindoline and Diclofenac Sodium with their respective target receptors.
Results of ADME/T Analysis
Vindoline and Diclofenac Sodium exhibit distinct pharmacokinetic and physicochemical properties (Table 4). Vindoline (Plant-derived alkaloid) has a higher molecular weight (456.53 g/mol) and greater lipophilicity than Diclofenac Sodium (318.13 g/mol). Both drugs show high gastrointestinal absorption. Their metabolic profiles also vary; Vindoline primarily inhibits the CYP3A4 and CYP2D6 enzymes, whereas Diclofenac Sodium predominantly inhibits the CYP1A2 enzyme. Both compounds adhere to Lipinski’s Rule of Five, indicating favorable drug-likeness, and have a similar bioavailability score of 0.55 (Figure 7 & 8). Diclofenac Sodium can cross the blood-brain barrier. Vindoline is more suitable for peripheral use as it does not cross the BBB.
Table 4: ADME Analysis of Vindoline and Diclofenac Sodium
| Drug | Lipinski’s Rules | Lipinski’s Rule Violations | GI Absorption | BBB Permeability | P-gp Substrate | CYP3A4 Inhibitor | Bioavailability Score | |||
| MW
(<500) |
Log P
(≤ 5) |
HBA
(≤ 10) |
HBD
(≤ 5) |
|||||||
| Vindoline | 456.53 g/mol | 1.78 | 7 | 1 | 0 | High | No | No | Yes | 0.55 |
| Diclofenac Sodium | 318.13 g/mol | 0.65 | 2 | 1 | 0 | High | Yes | Yes | No | 0.55 |
 |
Figure 7: ADME Analysis of Vindoline |
 |
Figure 8: ADME Analysis of Diclofenac Sodium |
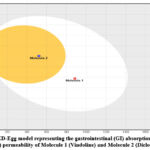 |
Figure 9: BOILED-Egg model representing the gastrointestinal (GI) absorption and blood-brain barrier (BBB) permeability of Molecule 1 (Vindoline) and Molecule 2 (Diclofenac Sodium) |
The Boiled-Egg model is plotted based on WLOGP (lipophilicity) vs. TPSA (topological polar surface area). The Boiled-Egg model (Figure 9) suggests that Vindoline (Molecule 1) falls within the white region, indicating good gastrointestinal absorption but poor blood-brain barrier (BBB) permeability. While, Diclofenac Sodium (Molecule 2) is positioned in the yellow area, indicating potential BBB penetration, it is suitable for CNS-related conditions (Table 5).
Table 5: Toxicity Prediction Data of the Vindoline and Diclofenac Sodium as per Protox server Results
| Drug | Vindoline | Diclofenac Sodium |
| Predicted LD50 (mg/kg) | 150mg/kg | 53mg/kg |
| Predicted Toxicity Class | 3 | 3 |
| Average Similarity % | 43.13% | 98.59% |
| Prediction Accuracy | 54.26% | 72.9% |
| Hepatotoxicity | Inactive | Active |
| Carcinogenicity | Active | Inactive |
| Immunotoxicity | Active | Inactive |
| Neurotoxicity | Active | Active |
| Nephrotoxicity | Active | Active |
| Respiratory Toxicity | Active | Active |
| Cardiotoxicity | Inactive | Inactive |
| Mutagenicity | Inactive | Inactive |
| Cytotoxicity | Inactive | Inactive |
Results of Egg Albumin Denaturation Assay
The ability of Vindoline to inhibit egg albumin denaturation was evaluated and compared with that of the standard drug, Diclofenac Sodium. The experiment was carried out in triplicate to ensure reliability. The outcomes, indicating the percentage inhibition of protein denaturation, are presented in Table 6.
Table 6: Effect of Vindoline & Diclofenac Sodium on Egg Albumin Denaturation
| S. No. | Treatment Group | Concentration (μg/ml) | Absorbance recorded at 660nm | % Inhibition on Egg Albumin Denaturation ± SEM |
| 1 | Control | 0.023 | – | |
| 2 | Standard (Diclofenac Sodium) | 50 | 0.015 | 34.78±0.141 |
| 100 | 0.012 | 47.82±0.292 | ||
| 150 | 0.010 | 56.52±0.413 | ||
| 200 | 0.009 | 60.86±0.219 | ||
| 250 | 0.007 | 69.56±0.269 | ||
| 3 | Test (Vindoline) | 50 | 0.013 | 43.47±0.265 |
| 100 | 0.011 | 52.17±0.467 | ||
| 150 | 0.008 | 65.21±0.561 | ||
| 200 | 0.007 | 69.56±0.338 | ||
| 250 | 0.005 | 78.26±0.057 | ||
All values are expressed in terms of Mean of % Inhibition ± SEM and are found to be significant when compared to standard p<0.05
The findings revealed that Diclofenac Sodium, used as the standard, exhibited protein denaturation inhibition of 34.78%, 47.82%, 56.52%, 60.86%, and 69.56% at concentrations of 50, 100, 150, 200, and 250 μg/ml, respectively. In comparison, Vindoline demonstrated inhibition values of 43.47%, 52.17%, 65.21%, 69.56%, and 78.26% at the same concentration levels. The highest inhibitory effects were recorded at 250 μg/ml, where Vindoline and Diclofenac Sodium showed inhibition percentages of 78.26% and 69.56%, respectively. These results suggest that Vindoline exhibits greater anti-inflammatory activity than the standard drug at the highest tested concentration. A graphical representation comparing the percentage inhibition of protein denaturation by Vindoline and Diclofenac Sodium is provided in Figure 10.
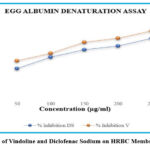 |
Figure 10: Effect of Vindoline and Diclofenac Sodium on HRBC Membrane Stabilization |
Discussion
Inflammation is a natural defense mechanism, but when it becomes uncontrolled, it can contribute to the development of chronic diseases. Non-steroidal anti-inflammatory drugs (NSAIDs) like Diclofenac Sodium are commonly used to manage inflammation, but their potential side effects and toxicity raise concerns, prompting the need for safer alternatives. This study investigated the anti-inflammatory potential of Vindoline, a naturally occurring alkaloid. We compared its effectiveness with that of Diclofenac Sodium by analyzing their molecular interactions with key inflammatory targets, COX-2 and NF-κB and its effect on HRBC membrane lysis.
Our findings revealed that Vindoline binds more strongly to both COX-2 (-9.4) and NF-κB (-7.0) than Diclofenac Sodium (-8.0 and -6.4, respectively), suggesting it may be more effective in inhibiting these inflammation-related proteins.
Protein denaturation refers to the loss of a protein’s secondary and tertiary structure, often resulting in the loss of its biological activity. This process is widely recognized as an indicator of inflammation. When proteins become denatured, they can give rise to autoantigens, which may trigger inflammatory responses and contribute to conditions such as rheumatoid arthritis, cancer, and diabetes. Therefore, inhibiting protein denaturation is considered a potential strategy for controlling inflammation.39 The egg albumin denaturation method offers a cost-effective approach to evaluating the anti-inflammatory potential of isolated compounds. In this assay, proteins are denatured by exposing them to high temperatures.
The results obtained demonstrated that Vindoline can statistically significantly (p<0.05) inhibit protein denaturation. Standard drug Diclofenac sodium showed 34.78, 47.82, 56.52, 60.86 and 69.56% inhibition of protein denaturation at 50, 100, 150, 200 and 250 µg/ml of sample concentration whereas Vindoline showed 43.47, 52.17, 65.21, 69.56 and 78.26 % at 50, 100, 150, 200 and 250 µg/ml of sample concentration in the concentration dependent manner. This study demonstrates that the percentage inhibition of protein denaturation was significantly increased with Vindoline compared to the standard drug diclofenac sodium. The highest level of inhibition was recorded at a sample concentration of 250 μg/ml. According to the results of the egg albumin denaturation assay, Vindoline was found to be more effective in curing inflammation, exhibiting the highest % inhibition of protein denaturation compared to the conventional standard drug Diclofenac Sodium.
Additionally, Vindoline demonstrated better water solubility (ESOL: -3.35) compared to Diclofenac Sodium (ESOL: -4.76), which could lead to better absorption and bioavailability—an important factor in drug effectiveness.
One significant advantage of Vindoline is that it does not cross the blood-brain barrier (BBB), making it a safer choice for treating peripheral inflammation without the risk of central nervous system (CNS) side effects. In contrast, Diclofenac Sodium can penetrate the BBB, which may contribute to neurological side effects over long-term use. In terms of toxicity, while both drugs exhibit certain risks, Vindoline has a higher LD50 (150 mg/kg) than Diclofenac Sodium (53 mg/kg), indicating a lower acute toxicity. However, while Diclofenac Sodium is primarily associated with liver toxicity, Vindoline carries potential neurotoxic, nephrotoxic, and immunotoxic risks, highlighting the need for further safety studies.
Overall, Vindoline shows promise as a potential anti-inflammatory agent with strong molecular interactions, improved solubility, and a potentially better safety profile than Diclofenac Sodium. Its lack of BBB permeability makes it a safer option for managing inflammation outside the CNS. However, further in-depth studies, including in vitro and in vivo research, are needed to fully evaluate its efficacy, pharmacokinetics, and long-term safety before it can be considered for clinical use.
Conclusion
Vindoline exhibits stronger binding affinity to COX-2 and NF-κB, and a better percentage of inhibition of protein denaturation indicating superior anti-inflammatory potential compared to diclofenac sodium. Its higher solubility enhances bioavailability, and its inability to cross the BBB minimizes CNS-related side effects, making it more suitable for non-CNS inflammation. With an optimal LogP (~1.8) and a higher LD50 (150 mg/kg), Vindoline demonstrates better permeability and lower toxicity, lacking hepatotoxicity, a significant concern with diclofenac sodium. Given its improved efficacy, solubility, safety profile, and reduced systemic risks, Vindoline emerges as a promising candidate for anti-inflammatory therapy.
Acknowledgement
The authors sincerely thank Dr. Rakhi Mishra, Professor, Noida Institute of Engineering and Technology Pharmacy Institute, Greater Noida for her valuable assistance in the visualization of molecular docking results. Her expertise and support greatly contributed to the analysis and interpretation of the findings in this study.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to Reproduce Material from Other Sources
Not Applicable
Author’s Contribution
- Preeti Chaudhary and Akash Kumavat performed molecular docking and preparation of the manuscript.
- Pritam Khandave visualized the docking results.
- Rupesh Pingale provided the facilities, expertise, and software to perform all computational studies.
- Sakshi Singh and Omkar Dudhal performed ADME analysis.
- Sonali Korlekar and Shubham Tivlekar performed toxicity studies.
References
- Medzhitov R. Origin and physiological roles of inflammation. Nature. 2008;454(7203):428-435.
CrossRef - Chen L, Deng H, Cui H, Fang J, Zuo Z, Deng J, et al. Inflammatory responses and inflammation-associated diseases in organs. Oncotarget. 2017;9(6):7204-7218.
CrossRef - Sansbury BE, Spite M. Resolution of acute inflammation and the role of resolvins in immunity, thrombosis, and vascular biology. Circ Res. 2016;119(1):113-130.
CrossRef - Soliman AM, Barreda DR. Acute inflammation in tissue healing. Int J Mol Sci. 2022;24(1):641.
CrossRef - Carol AF, Timothy MW. Cytokines in acute and chronic inflammation. Front Biosci (Landmark Ed). 1997;2(4):12-26.
CrossRef - Chavda VP, Feehan J, Apostolopoulos V. Inflammation: the cause of all diseases. Cells. 2024;13(22):1906.
CrossRef - Furman D, Campisi J, Verdin E, et al. Chronic inflammation in the etiology of disease across the life span. Nat Med. 2019;25(12):1822-1832.
CrossRef - Desai SJ, Prickril B, Rasooly A. Mechanisms of phytonutrient modulation of cyclooxygenase-2 (COX-2) and inflammation related to cancer. Nutr Cancer. 2018;70(3):350-375.
CrossRef - Abdulkhaleq LA, Assi MA, Abdullah R, Zamri-Saad M, Taufiq-Yap YH, Hezmee MNM. The crucial roles of inflammatory mediators in inflammation: a review. Vet World. 2018;11(5):627-635.
CrossRef - Cui J, Jia J. Natural COX-2 inhibitors as promising anti-inflammatory agents: an update. Curr Med Chem. 2021;28(18):3622-3646.
CrossRef - Simon LS. Role and regulation of cyclooxygenase-2 during inflammation. Am J Med. 1999;106(5B):37S-42S.
CrossRef - Tak PP, Firestein GS. NF-kappaB: a key role in inflammatory diseases. J Clin Invest. 2001;107(1):7-11.
CrossRef - Liu T, Zhang L, Joo D, Sun SC. NF-κB signaling in inflammation. Signal Transduct Target Ther. 2017; 2:17023.
CrossRef - Sekar G, Shankar V, Rajeswari VD. Pharmacological activities of Catharanthus roseus: a perspective review. Int J Pharma Bio Sci. 2013;4(2):431-439.
- Ahmad B, Banerjee A, Tiwari H, Jana S, Bose S, Chakrabarti S. Structural and functional characterization of the vindoline biosynthesis pathway enzymes of Catharanthus roseus. J Mol Model. 2018;24(3):53.
CrossRef - Dhyani P, Quispe C, Sharma E, Bahukhandi A, Sati P, Attri DC, et al. Anticancer potential of alkaloids: a key emphasis to colchicine, vinblastine, vincristine, vindesine, vinorelbine and vincamine. Cancer Cell Int. 2022;22(1):206.
CrossRef - Kumar S, Singh B, Singh R. Catharanthus roseus (L.) G. Don: a review of its ethnobotany, phytochemistry, ethnopharmacology and toxicities. J Ethnopharmacol. 2022;284:114647.
CrossRef - Singh P, Singh DP, Patel MK, Binwal M, Kaushik A, Mall M, et al. Vindoline is a key component of Catharanthus roseus leaf juice extract prepared through an Ayurveda-based method for ameliorating insulin-resistant type 2 diabetes. Protoplasma. 2025. doi:10.1007/s00709-024-02026-w.
CrossRef - Goboza M, Meyer M, Aboua YG, Oguntibeju OO. In vitro antidiabetic and antioxidant effects of different extracts of Catharanthus roseus and its indole alkaloid, vindoline. Molecules. 2020;25(23):5546.
CrossRef - Sertel S, Fu Y, Zu Y, Rebacz B, Konkimalla B, Plinkert PK, et al. Molecular docking and pharmacogenomics of vinca alkaloids and their monomeric precursors, vindoline and catharanthine. Biochem Pharmacol. 2011;81(6):723-735.
CrossRef - RCSB Protein Data Bank. RCSB PDB: The Protein Data Bank [Internet]. https://www.rcsb.org/. Accessed January 31, 2025.
- Morita I. Distinct functions of COX-1 and COX-2. Prostaglandins Other Lipid Mediat. 2002;68-69:165-175.
CrossRef - Wang B, Wu L, Chen J, et al. Metabolism pathways of arachidonic acids: mechanisms and potential therapeutic targets. Signal Transduct Target Ther. 2021;6:94.
CrossRef - Dolcet X, Llobet D, Pallares J, Matias-Guiu X. NF-kB in development and progression of human cancer. Virchows Arch. 2005;446(5):475-482.
CrossRef - Sastry GM, Adzhigirey M, Day T, Annabhimoju R, Sherman W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput Aided Mol Des. 2013;27(3):221-234.
CrossRef - Valdes-Tresanco MS, Valdes-Tresanco ME, Valiente PA, Moreno E. AMDock: a versatile graphical tool for assisting molecular docking with Autodock Vina and Autodock4. Biol Direct. 2020;15(1):12.
CrossRef - National Center for Biotechnology Information. PubChem Compound Summary for CID 260535 (Vindoline) and CID 5018304 (Diclofenac Sodium) [Internet]. https://pubchem.ncbi.nlm.nih.gov/. Accessed January 31, 2025.
- Ai S, Lin G, Bai Y, Liu X, Piao L. QSAR classification-based virtual screening followed by molecular docking identification of potential COX-2 inhibitors in a natural product library. J Comput Biol. 2019;26(11):1296-1315.
CrossRef - Nguyen NT, Nguyen TH, Pham TNH, et al. Autodock Vina adopts more accurate binding poses but Autodock4 forms better binding affinity. J Chem Inf Model. 2020;60(1):204-211.
CrossRef - Yadav CK, KC S, Thapa S. In vitro and in silico analysis of α-amylase inhibitory activity of ethanolic extract of Adhatoda vasica Glob Adv Integr Med Health. 2024;13.
CrossRef - Trott O, Olson AJ. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem. 2010;31(2):455-461.
CrossRef - Umi B, Zahra SM, Wanda D, Fauzian GR, Muhammad Y. Molecular interaction analysis and visualization of protein-ligand docking using Biovia Discovery Studio Visualizer. Indones J Comput Biol. 2023;2(1):22-30.
CrossRef - Daina A, Michielin O, Zoete V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep. 2017;7:42717.
CrossRef - Rajapaksha HKK, Fernando MN, Nelumdeniya RM, Bandara KK, Silva RN. Evaluation of in vitro anti-inflammatory activity and in-silico pharmacokinetics and molecular docking study of Horsfieldia iryaghedhi. J Phytopharmacol. 2024;13(2):143-153.
CrossRef - Clark JH. Denaturation changes in egg albumin with urea, radiation, and heat. J Gen Physiol. 1943;27(2):101-111.
CrossRef - Goryanin I, Ovchinnikov L, Vesnin S, Ivanov Y. Monitoring protein denaturation of egg white using passive microwave radiometry (MWR). Diagnostics (Basel). 2022;12(6):1498.
CrossRef - Dharmadeva S, Galgamuwa LS, Prasadinie C, Kumarasinghe N. In vitro anti-inflammatory activity of Ficus racemosa bark using albumin denaturation method. Ayu. 2018;39(4):239-242.
CrossRef - Madhuranga HDT, Samarakoon DNAW. In vitro anti-inflammatory egg albumin denaturation assay: an enhanced approach. J Nat Ayurvedic Med. 2023;7(3):000411.
CrossRef - Godhandaraman S, Ramalingam V. In vitro anti-inflammatory activity of different parts of Pedalium murex (L.). Int J Herbal Med. 2016;4(3):31-36.







