Manuscript accepted on :17-04-2025
Published online on: 01-05-2025
Plagiarism Check: Yes
Reviewed by: Dr. Durgeshranjan Kar
Second Review by: Dr. Naha Ananya
Final Approval by: Dr. Eman Refaat Youness
Premalatha Ravi1 , Jayanthi Krishnasamy Balasundaram1*
, Jayanthi Krishnasamy Balasundaram1* , Rajasekaran Chinnappan1
, Rajasekaran Chinnappan1 and Sureshkumar Ramasamy2
and Sureshkumar Ramasamy2
1Department of ECE, K.S.Rangasamy College of Technology, Namakkal, India
2Radiation Oncologist, Erode Cancer Centre, Erode, India.
Corresponding Author E-mail: jayanthikb@gmail.com
DOI : https://dx.doi.org/10.13005/bpj/3172
Abstract
(Breast cancer represents the most prevalent variant of malignancy observed in the female population) 2.3 million were diagnosed with breast cancer in 2022. Early detection enhances the quality of life for breast cancer patients, and one promising approach to achieving this is through the analysis of histopathological images using pre-trained models of convolutional neural networks (CNNs) architectures, namely ResNet152, InceptionV3, and MobileNetV2, all initially trained on the ImageNet dataset. This paper presents an analysis of these architectures applied to the breast cancer dataset, comparing their robustness and effectiveness in detecting breast cancer. The results demonstrate that models pre-trained on ImageNet perform significantly better compared to the same architectures trained from scratch on the breast cancer dataset. This difference in performance highlights the importance of transfer learning in analyzing medical images. It shows that using models already trained on large and varied datasets like ImageNet can greatly improve the ability to identify features in histopathological images. The results help to decide the robustness of the architectures for the given dataset. The results will support researchers working in this domain to understand which architecture yields better results in breast cancer diagnosis.
Keywords
Breast Cancer; CNN; Histopathological Images; Pre-trained Models; Transfer Learning
Download this article as:| Copy the following to cite this article: Ravi P, Balasundaram J. K, Chinnappan R, Ramasamy S. Comparative Analysis of Transfer Learning Models for Breast Cancer Detection: Leveraging Pre-Trained Networks for Enhanced Diagnostic Accuracy. Biomed Pharmacol J 2025;18(2). |
| Copy the following to cite this URL: Ravi P, Balasundaram J. K, Chinnappan R, Ramasamy S. Comparative Analysis of Transfer Learning Models for Breast Cancer Detection: Leveraging Pre-Trained Networks for Enhanced Diagnostic Accuracy. Biomed Pharmacol J 2025;18(2). Available from: https://bit.ly/3GCdpfF |
Introduction
Incorporating the applicable criteria that follow. Breast cancer has led to 670000 deaths globally in 2022 (The burden of breast cancer is high), (Especially in nations exhibiting elevated levels of the Human Development Index (HDI)). Breast cancer treatments are more effective when started early. Histopathological images are crucial in diagnosing breast cancer, providing detailed views of tissue samples to identify cancerous cells accurately. These images play a vital role in detecting breast cancer at an early stage, which significantly improves the effectiveness of subsequent treatments.1
Breast cancer is the primary cause of mortality for women in developing nations,2,3,4 With the ratio of health professionals to the population being low in these countries, it is necessary for an automated diagnosis and classification of the disease.5,6
With the recent developments in machine learning, and deep learning techniques, researchers have started to use various Conventional Neural Network (CNN) architectures for diagnosis of various diseases as well as classification . The limitation in this is the size of the dataset.7,8,9Machine learning techniques require larger datasets for better accuracy . In medical image analysis, the size of the dataset is always small. This leads to overfitting which reduces the test accuracy of any network.10
Pre-trained models are proposed by the researchers to address this problem. ImageNet is the largest dataset available and is used for training the CNNs.11,12 Though the dataset has very few medical images, pre-trained models with this dataset are seen to provide results with improved accuracy of classification. The learning transferred from this dataset increases the accuracy of classification.13,14
This paper proposes to analyze breast cancer detection with three state–of–the–art pre-trained models. ResNet152, Mobile NetV2, and InceptionV3 networks. These networks, pre-trained on the ImageNet dataset, are employed to analyze histopathological images of breast cancer. Both the scratch and the pre-trained models are used and are tested with the same dataset for analysis. Pre-trained models yield better results when compared to scratch models highlighting the significance of transfer learning in medical image analysis. The results indicate that researchers can work further in this area to select the optimal network for the specific dataset.
Literature survey
Several deep-learning models are being proposed by researchers for disease identification and classification. Deep Breast Cancer Net 15 proposes a framework that is validated with standard publicity available dataset with an accuracy claimed of 99.63%. ResNet and InceptionV3 architectures are used as default models 16and with transfer learning applied from pre-trained models.17 In this pre-trained model with the dataset, ResNet50 gives the best results considering all the metrics. Feature selection is very important in convolution Neural Networks. Researchers on this to identify the significant features. Nearest Neighbour, Random Forest (RF) and support vector machine are taken for analysis with the RSNA (Radiological Society of North America) dataset.18The results are better with MIAS (Mammographic Image Analysis Society) and DDSM (Digital Database for Screening Mammography) datasets than RSNA. LeNet with modified ReLU (Rectified Linear Unit) gives an improved performance with batch normalization.19Mobile Net-based architecture gives results with less than 90% accuracy.20,21
For breast histopathology classification, a study comparing CNN and DenseNet121 based on transfer learning revealed that DenseNet121 obtained 86.6% accuracy at 100X magnification with 128×128 scaling, whereas training accuracy was improved by 16.4% with transfer learning.22FA-VGG16, a deep network for classifying images of breast histopathology that is based on Forward Attention. It greatly outperformed standard VGG16, with 97.7% accuracy for binary classification and 92.4% accuracy for quaternary classification.23Deep Learning (DL) improves cancer diagnosis in histopathology by more accurately and quickly analysing whole slide images (WSI) of stained tissues. DL compares its performance to human experts, highlighting challenges and opportunities in breast cancer detection and medical image analysis.24Pretrained models 25,26are applied with optimization algorithms like Advanced AI-Birum Earth Radius to boost the classification performance. Additional statistical tests are also conducted to evaluate the results and the model is robust.27
Inception V3, a sophisticated deep convolutional neural network architecture, for automatic breast cancer detection through thermography with average classification accuracies obtained (at epochs 3, 5, and 6) were (98.104%, 98.712%, and 97.816%) respectively.28A concatenated transfer learning model designed for breast cancer histopathology utilizes pre-trained convolutional neural networks such as VGG-16, MobileNetV2, ResNet50, and DenseNet121 to derive deep features, which are then combined into a unified feature vector for classification purposes.29By leveraging pre-trained models and advanced feature selection methods, researchers can enhance the accuracy and robustness of diagnostic tools, ultimately contributing to better early detection and treatment outcomes for breast cancer patients.30,31
Xception achieved the best accuracy of 90.86%, surpassing prior state-of-the-art results, and transfer learning using InceptionV3 and Xception on the BreakHis dataset surpassed training from scratch.32Using the BreakHis dataset of histological images, transfer learning model of DenseNet121 achieved the highest breast cancer classification accuracy of 0.965 compared to other transfer learning models like (DenseNet201, VGG16, VGG19, InceptionV3, MobileNetV2).33The BreakHis dataset was used to test seven transfer learning models (LeNet, VGG16, DarkNet53, DarkNet19, ResNet50, Inception, and Xception) without any preprocessing or augmentation. On the unbalanced dataset, DarkNet53 had the best Balanced Accuracy (87.17%), whereas Xception had the highest accuracy (83.07%).34A learnable adaptive resizer for breast cancer classification using the BreakHis dataset to address data loss resulting from downsizing high-resolution histopathology images. Using a variety of convolutional neural networks, including DenseNet201, the method produced a high accuracy of 98.96% at 448×448 resolution.35
Hybrid approaches combining hand-crafted features like HOG (Histogram of Oriented Gradients) and LBP (Local Binary Pattern) with deep learning features from a CNN, breast cancer classification using mammograms outperforms the state-of-the-art techniques for breast cancer classification on the CBIS-DDSM dataset.36CNNs are the most accurate and widely used models for breast cancer diagnosis, evaluated mainly using accuracy metrics. The Wisconsin Breast Cancer Dataset (WBCD) was used, with 273 out of 569 samples for testing and the remaining for training and validation.37Two methods based on ResNet are used to categorise histopathology images of breast cancer: one uses self-supervised contrastive learning with limited labeled data, while the other combines ResNet50 with Inception modules to create an effective architecture. The model achieved 98% accuracy at 40X and 200X magnifications, and 94% at 100X and 400X, with a significantly reduced parameter count of 3.6 million.38Fusion of hybrid deep features (FHDF) using multiple CNNs, such as VGG16, VGG19, ResNet50, and DenseNet121, specifically tackling the issues of multilabel classification and unequal dataset distributions. The method detects different severities of breast cancer with high classification accuracies of 98.7%, 97.7%, and 98.8%, respectively, when applied to three public datasets (MIAS, CBIS-DDSM, and INbreast).39
Recent research has focused on enhancing deep-learning frameworks for the classification of histopathological images pertaining to breast cancer. Deep transfer learning models like ResNet50 are used for breast cancer detection using histopathology images, with ResNet50 outperforming other models with 90.2% accuracy, 90.0% AUC, 94.7% recall rates, and a marginal loss of 3%.40,41The breast cancer images from the ICIAR (International Conference on Image Analysis and Recognition) 2018 histopathological dataset are used, utilizing the ResNet-152v2 CNN architecture to extract features for distinguishing various types of breast cancer (normal, benign, in situ carcinoma, and invasive carcinoma) from histopathological images.42Deep transfer learning models for breast cancer histopathology classification that use ResNet50 and DenseNet121, optimised with ImageNet pre- trained weights and data augmentation. The models outperformed earlier CAD (Computer Aided Diagnosis) systems, achieving 100% accuracy in binary classification and 98% accuracy in multiclass classification.43The CNN model achieved a higher accuracy (92%) and F1-score (92%) on a dataset consisting of 57 patients in thermal breast imaging, outperforming state-of-the-art architectures like ResNet50, SeResNet50, and Inception.44CNN commonly employed to develop an efficient breast cancer classification model.45
Transfer learning
Transfer learning is a machine learning technique in which a model trained on one task is repurposed as the foundation for a model on a related task. This method is especially valuable when the dataset for the second task is limited, as it builds on the knowledge gained from the first task. Nowadays, it is very hard to see people training whole convolutional neural networks from scratch. Instead, it is typical to use a pre-trained model, often trained on diverse images for similar tasks such as models trained on ImageNet (1.2 million images across 1,000 categories) and leverage their features to address new tasks.
Pre-trained Model Representation
Assume there is a pre-trained model denoted as Mpre, which has been trained on a source task with a dataset Dsource. This model can be represented as a function that maps input data Xsource to predicted labels Ysource, denoted in equation (1).
![]()
In neural networks, this can be represented as shown in equation (2).
![]()
where θpre represents the pre-trained model’s parameters.
Feature Extraction
To transfer knowledge from Mpre, to a new task with a dataset Dtarget, are extracted features from the pre-trained model. Let the extracted features be denoted as mentioned in equations (3) and (4).

where Xtarget is the input data for the target task.
In neural networks, this is represented as shown in equations (5) and (6).

where gpre represents the feature extraction function.
Materials and Methods
Dataset
The histopathological images of breast cancer are collected from the Kaggle database. 3928 images are taken from the Kaggle database out of which 2,627 images utilized for training the network, 656 images set aside for validation, and 645 images allocated for testing. The data is collected from the given link: https://www.kaggle.com/datasets/obulisainaren/multi-cancer.
Data Pre-Processing
The Breast Cancer dataset collected from the Kaggle database is initially pre-processed. Collected images are resized to 224 X 224 pixels and augmented and used for training.
CNN Architectures
In deep learning, pre-trained models serve as invaluable resources for tasks such as image classification, where extensive computational resources and labeled datasets are required for training. This paper conducted a comprehensive comparative study of three state-of-the-art pre-trained models: ResNet152, InceptionV3, and MobileNetV2, initialized with weights trained on the ImageNet dataset. Each model explores their unique characteristics and advantages. Furthermore, evaluating the performance of these models on ImageNet, a widely recognized benchmark dataset for image classification tasks.
ResNet152
ResNet-152 introduces the idea of residual connections, often referred to as skip connections or shortcut connections. These connections enable the network to bypass certain layers and propagate the input directly to deeper layers. By implementing this approach, ResNet-152 is able to learn residual mappings, which capture the difference between the input and the target output. This effectively addresses the vanishing gradient problem often faced when training very deep neural networks, enabling better optimization and training of the model.
Compared to shallower architectures, such as ResNet-50 or VGG-16, ResNet-152 has a larger capacity to capture intricate patterns and details in the data.
InceptionV3
InceptionV3 introduces a series of inception modules, which are convolutional modules with several parallel branches. Each branch performs convolutions of varying sizes (1×1, 3×3, and 5×5) to capture features across various spatial scales. These parallel branches are concatenated together to form the output of the inception module, enabling the network to capture both local and global features efficiently.
InceptionV3 employs factorized convolutions to reduce computational complexity and memory requirements. Instead of performing a single convolution with a large kernel size, factorized convolutions split the operation into smaller convolutions, such as 1×3 and 3×1 convolutions, which are less computationally expensive. This allows the network to achieve better performance.
MobileNetV2
MobileNetV2 is a convolutional neural network architecture designed for mobile and embedded vision applications. MobileNetV2 introduces two hyperparameters, width multiplier, and resolution multiplier, which allow for flexibility in controlling the model’s size and computational complexity. MobileNetV2 replaces the standard ReLU activation function with ReLU6 in the bottleneck layers. It helps to mitigate the problem of vanishing gradients and exploding activations, especially in deeper networks.
Proposed Method
The proposed block diagram for breast cancer prediction is shown in Figure 1. Pre-trained models such as ResNet152, InceptionV3, and MobileNetV2 are initially chosen. These models have undergone pre-training on the ImageNet dataset, which contains many images across a wide range of categories. The features learned by these models during training on ImageNet can be valuable for extracting meaningful features from breast cancer images.
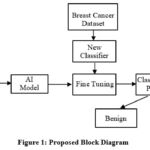 |
Figure 1: Proposed Block DiagramClick here to view Figure |
The pre-trained models serve as feature extractors. The convolutional layers of these models act as feature extractors that can effectively capture meaningful patterns and features from breast cancer images. The images are passed through the pre-trained models, and the output of the last convolutional layer is extracted as features. After feature extraction, fine-tuning is done by adding a new layer on top of the pre-trained model for the classification process. The fine-tuned models are then trained on the breast cancer dataset. The dataset is typically split into training, validation, and testing sets. The dataset is customarily partitioned into training, validation, and testing subsets. The model undergoes training utilizing the training subset, while its efficacy is assessed on the validation subset to oversee potential overfitting and ascertain the appropriate cessation of training. Finally, the trained model’s performance is evaluated on the testing set to find the ability and accuracy in classifying breast cancer images.
Results
The pre-trained models are trained for 25 epochs with a learning rate of 0.0001 using a Adam optimizer and binary cross entropy loss function. The training dataset consists of 2,627 images. Once the model is trained the same model is validated with 656 images. A batch size of 32 is used for training, and the activation function is sigmoid. The training and validation process is carefully monitored throughout all 25 epochs, ensuring the model’s performance is continually assessed and optimized. The models are trained using an NVIDIA GeForce RTX 4090 GPU with 24 RAM, ensuring efficient computation. This iterative process involves training the selected pre-trained models such as ResNet152, InceptionV3, and MobileNetV2 on the histopathological images sourced from Kaggle. The same hyperparameter values are given to train the scratch model of the same architecture.
After the completion of each epoch, the accuracy curves are plotted for both the training and validation datasets. Graphs [1-2] shows the comparison of training and validation accuracy plots of ResNet152 while Graphs [3-4] depicts the same for Inception V3 and Graphs [5-6] for MobileNetV2.
 |
Graph 1: Training Accuracy Plot-ResNet152Click here to view Graph |
 |
Graph 2: Validation Accuracy Plot- ResNet152Click here to view Graph |
 |
Graph 3: Training Accuracy Plot-InceptionV3Click here to view Graph |
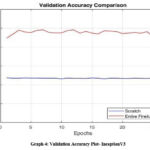 |
Graph 4: Validation Accuracy Plot- InceptionV3Click here to view Graph |
 |
Graph 5: Training Accuracy Plot-MobileNetV2Click here to view Graph |
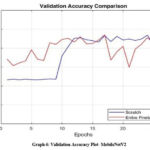 |
Graph 6: Validation Accuracy Plot- MobileNetV2 Click here to view Graph |
Discussion
Once the training process is completed, the resultant trained models are subjected to testing using a separate set of 656 test images. These images are unseen by the model during training and validation, serving as an independent evaluation of its performance. Table 1 gives the metrics of accuracy, F1-score, Precision, and Recall for the scratch model. The scratch models’ performance indicates that ResNet152V2 and InceptionV3 perform similarly, with training accuracies of 47.13% and 47.05% and validation accuracies of 47.19% and 47.50%, respectively. An F1-score of 0.74 and a testing accuracy of 59.22% are attained by both models, suggesting balanced precision (0.58) and recall (1.0). According to this, they may over predict positive cases but are skilled at spotting them. MobileNetV2, on the other hand, exhibits a substantially lower testing accuracy of 75.50% but a significantly better training accuracy of 94.03% and validation accuracy of 93.59%). Together with a precision of 0.59 and recall of 0.37, the model’s F1-score of 0.45 indicates that it has trouble generalizing and does not detect all relevant positive examples during testing.
Table 1: Performance Analysis for Scratch Model
| CNN | Training | Validation | Testing | F1-Score | Precision | Recall |
| RESNET152V2 | 47.13 | 47.19 | 59.22 | 0.74 | 0.58 | 1.0 |
| INCEPTIONV3 | 47.05 | 47.50 | 59.22 | 0.74 | 0.58 | 1.0 |
| MOBILENETV2 | 94.03 | 93.59 | 75.50 | 0.45 | 0.59 | 0.37 |
The Pre-trained model performance gives the metrics of accuracy, F1-score, Precision, and Recall, as shown in Table 2. InceptionV3 stands out as the leading model with the highest testing accuracy of 96.74%, alongside a remarkable F1-score of 0.96, a precision of 0.99, and a recall of 0.93, signifying its strong ability to both recognize positive cases and reduce false positives. ResNet152V2 also shows commendable results, achieving a training accuracy of 99.38%, a validation accuracy of 93.75%, and a testing accuracy of 83.10%. Its F1-score stands at 0.83, with a precision of 0.95 and a recall of 0.75, demonstrating a reasonable balance with a slight compromise in recall. MobileNetV2 records a testing accuracy of 90.07%, with an F1-score of 0.92, a precision of 0.86, and an ideal recall of 1.0, indicating its exceptional ability to capture all relevant positive cases. InceptionV3 excels in performance, followed by MobileNetV2 and then ResNet152V2.
Table 2: Performance Analysis For Pre-Trained Model
| CNN | Training | Validation | Testing | F1-Score | Precision | Recall |
| RESNET152V2 | 99.38 | 93.75 | 83.10 | 0.83 | 0.95 | 0.75 |
| INCEPTIONV3 | 99.81 | 96.88 | 96.74 | 0.96 | 0.99 | 0.93 |
| MOBILENETV2 | 99.61 | 94.06 | 90.07 | 0.92 | 0.86 | 1.0 |
When compared to the scratch model, the Pre-trained model utilizing ImageNet demonstrates superior performance across all metrics, emphasizing the importance of transfer learning in the field of medical image analysis. Within this Pre-trained model, InceptionV3 shows strong results for breast cancer histopathological images, attaining a testing accuracy of 96.74% and an AUC of 98.97%.
Conclusion
When compared to models trained from scratch, pre-trained models on the ImageNet dataset demonstrate significantly superior performance, underscoring the value of transfer learning in medical image analysis. Specifically, the InceptionV3 model, when pre-trained on ImageNet and fine-tuned for breast cancer histopathological images, exhibits remarkable results. This model achieves a testing accuracy of 96.74% and an Area Under the Curve (AUC) of 98.97%. Further, planned to utilize this model to be deployed in the Jetson Nano Development Kit. The Jetson Nano Development Kit is specifically designed for edge computing applications, offering a balance between computational power and energy efficiency. By deploying the InceptionV3 model on this platform, researchers can take advantage of its computational capabilities while minimizing resource consumption, making it well- suited for edge deployment scenarios where computational resources are limited. Furthermore, the InceptionV3 model’s ability to accurately classify breast cancer images on the Jetson Nano underscores its suitability for real-world applications in healthcare and medical imaging. The combination of computational efficiency and accuracy ensures that the model can provide timely and accurate diagnoses, ultimately improving patient care and outcomes.
Acknowledgment
The authors would like to express their sincere gratitude to Science and Engineering Research Board (SERB), India for providing financial support to this project through Core Research Grant (CRG).
Funding Source
Science and Engineering Research Board (SERB), India for providing financial support to this project through Core Research Grant (CRG). File No: CRG/2022/008063
Conflict of Interest
The author(s) declares no conflict of interest.
Data Availability Statement
The dataset used in this research was obtained from Kaggle and is publicly available. All data preprocessing, analysis, and model development were conducted based on this dataset. The specific dataset can be accessed at https://www.kaggle.com/datasets/obulisainaren/multi-cancer
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to Reproduce Material From Other Sources
Not Applicable
Author Contributions
- Jayanthi K.B : Conceptualization, Study design.
- Premalatha R : Methodology development, Literature review
- Rajasekaran C : Interpretation of results, manuscript review.
- Sureshkumar R: Data collection, Data analysis.
References
- Kaur A, Kaushal C, Sandhu JK, Damaševičius R, Thakur N. Histopathological image diagnosis for breast cancer diagnosis based on deep mutual learning. Diagnostics (Basel). 2024;14(1):95.
CrossRef - Choudhary S, Singh P, Mittal M, Singh G. Automated breast cancer diagnosis based on machine learning algorithms. Int J Adv Netw Appl. 2024;15(6):6229-6238.
CrossRef - Sha R, Kong Xm, Li Xy, et Global burden of breast cancer and attributable risk factors in 204 countries and territories, from 1990 to 2021: results from the Global Burden of Disease Study 2021. Biomark Research. 2024;12:87.
CrossRef - Afaya A, Ramazanu S, Bolarinwa OA, et al. Health system barriers influencing timely breast cancer diagnosis and treatment among women in low and middle-income Asian countries: evidence from a mixed-methods systematic BMC Health Services Research. 2022;22:1601.
CrossRef - Hekal AA, Elnakib A, Moustafa HED. Automated early breast cancer detection and classification system. Signal, Image & Video Processing. 2021;15:1497-1505.
CrossRef - Zhang M, Li S, Xue M, Zhu Q. Two-stage classification strategy for breast cancer diagnosis using ultrasound-guided diffuse optical tomography and deep learning. J Biomed Opt. 2023;28(8):086002-1.
CrossRef - Alzubaidi L, Zhang J, Humaidi Review of deep learning: concepts, CNN architectures, challenges, applications, future directions. J Big Data. 2021;8(53).
CrossRef
- Shekhar M, Khetavath S. Implementing lung cancer diagnosis framework in early stages using segmentation procedures and adaptive recurrent convolution neural network with region attention for classification. Sens Imaging. 2025;26:43
CrossRef - Alabduljabbar A, Khan SU, Alsuhaibani A, Almarshad F, Altherwy YN. Medical imaging datasets, preparation, and availability for artificial intelligence in medical imaging. J Alzheimer’s Dis Rep. 2024;8(1):1471-1483.
CrossRef - Kumar RR, Shankar SV, Jaiswal Advances in deep learning for medical image analysis: a comprehensive investigation. J Stat Theory Pract. 2025;19(9).
CrossRef - Raptis C, Karavasilis E, Anastasopoulos G, Adamopoulos A. Comparative analysis of conventional CNN vs. ImageNet pretrained ResNet in medical image classification. 2024;15(12):806.
CrossRef - Zhao,Z. (2024). A comparative study of large-scale and lightweight convolutional neural networks for ImageNet Applied and Computational Engineering,47,101-110.
CrossRef - Kim, E., Cosa-Linan, A., Santhanam, N. et al. Transfer learning for medical image classification: a literature review. BMC Med Imaging 22, 69 (2022).
CrossRef - Kumar R, Kumbharkar P, Vanam S, et al. Medical images classification using deep learning: a survey. Multimed Tools Appl. 2024;83:19683–19728. doi:10.1007/s11042-023-15576-7
CrossRef - Raza A, Ullah N, Khan JA, Assam M, Guzzo A, Aljuaid H. Deep Breast Cancer Net: A Novel Deep Learning Model for Breast Cancer Detection Using Ultrasound Images. Appl Sci. 2023;13(4):2082.
CrossRef - Ereken ÖF, Tarhan C. Breast Cancer Detection using Convolutional Neural Networks. In: 2022 International Symposium on Multidisciplinary Studies and Innovative Technologies (ISMSIT); 2022:597-601.
CrossRef - Guevara-Ponce V, Roque-Paredes O, Zerga-Morales C. Detection of Breast Cancer using Convolutional Neural Networks with Learning Transfer Mechanisms. Int J Adv Computer Sci Appl. 2023;14(6).
CrossRef - Jafari Z, Karami E. Breast Cancer Detection in Mammography Images: A CNN-Based Approach with Feature Selection. Information. 2023;14(7):410.
CrossRef - Balasubramaniam S, Velmurugan Y, Jaganathan D, Dhanasekaran S. A Modified LeNet CNN for Breast Cancer Diagnosis in Ultrasound Diagnostics (Basel). 2023;13(17):2746.
CrossRef - Ansar W, Shahid AR, Raza B, Dar AH. Breast Cancer Detection and Localization Using MobileNet Based Transfer Learning for Mammograms. In: Brito-Loeza C, Espinosa-Romero A, Martin-Gonzalez A, Safi A, eds. Intelligent Computing Systems. ISICS 2020. Communications in Computer and Information Science, vol 1187. Springer, Cham.
CrossRef - Joshi A, Gaud N. Breast Cancer Detection using MobileNetV2 and Inceptionv3 Deep Learning Techniques. Int J Eng Res Technol. 2022;11(9).
- Muntean CH, Chowkkar Breast Cancer Detection from Histopathological Images using Deep Learning and Transfer Learning. In: Proceedings of the 7th International Conference on Machine Learning Technologies (ICMLT 2022). 2022:164-169.
- Roy S, Jain PK, Tadepalli K, et al. Forward attention-based deep network for classification of breast histopathology Multimed Tools Appl. 2024;83:88039–88068.
CrossRef - Duggento A, Conti A, Mauriello A, Guerrisi M, Toschi N. Deep computational pathology in breast cancer. Semin Cancer Biol. 2020; 72: 226-237.
CrossRef - Guan S, Loew Breast Cancer Detection Using Transfer Learning in Convolutional Neural Networks. In: 2017 IEEE Applied Imagery Pattern Recognition Workshop (AIPR); 2017:1-8.
CrossRef - Mednikov Y, Nehemia S, Zheng B, Benzaquen O, Lederman Transfer Representation Learning using Inception-V3 for the Detection of Masses in Mammography. In: 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC); 2018:2587-2590.
CrossRef - Alhussan AA, Abdelhamid AA, Towfek SK. Classification of Breast Cancer Using Transfer Learning and Advanced Al-Biruni Earth Radius Optimization. 2023; 8(3):270.
CrossRef - Al Husaini MAS, Habaebi MH, Gunawan TS, Islam MR, Hameed SA. Automatic Breast Cancer Detection Using Inception V3 in Thermography. In: 2021 8th International Conference on Computer and Communication Engineering (ICCCE); 2021:255-258.
CrossRef - Dhayanithi J, Balasubramaniam S, Sureshkumar V, Dhanasekaran S. Revolutionizing Breast Cancer Diagnosis: A Concatenated Precision through Transfer Learning in Histopathological Data Analysis. Diagnostics (Basel). 2024;14(4):422.
CrossRef - Ahmad, N, Asghar, & Gillani, S.A. Transfer learning-assisted multi-resolution breast cancer histopathological images classification. Vis Computer. 2022;38: 2751–2770.
CrossRef
- Chakravarthy S, Nagarajan B, Kumar. Breast Tumor Classification with Enhanced Transfer Learning Features and Selection Using Chaotic Map-Based Optimization. Int J Computer Intelligent Syst. 2024;17:18.
CrossRef - Rai R, Sisodia Real-Time Data Augmentation Based Transfer Learning Model for Breast Cancer Diagnosis Using Histopathological Images. In Advances in Biomedical Engineering and Technology: Select Proceedings of ICBEST. 2018 ;473-488.
CrossRef - Ramasamy MP, Subburaj T, Krishnasamy V, Mannarsamy Performance analysis of breast cancer histopathology image classification using transfer learning models. Int J Electr Comput Eng. 2024;14(5).
CrossRef - Rana M, Bhushan Classifying breast cancer using transfer learning models based on histopathological images. Neural Comput Appl. 2023;35:14243–14257.
CrossRef - Duzyel O, Catal MS, Kayan CE, et Adaptive resizer-based transfer learning framework for the diagnosis of breast cancer using histopathology images. Signal, Image & Video Processing. 2023;17:4561–4570.
CrossRef - Sajid U, Khan RA, Shah SM, Arif Breast cancer classification using deep learned features boosted with handcrafted features. Biomed Signal Process Control. 2023; 86:105353
CrossRef - Akhtar N, Pant H, Dwivedi A, Jain V, Perwej A breast cancer diagnosis framework based on machine learning. Int J Sci Res Sci Eng Technol. 2023;10(3):118-132.
CrossRef - Ashraf FB, Alam SM, Sakib Enhancing breast cancer classification via histopathological image analysis: Leveraging self- supervised contrastive learning and transfer learning. Heliyon. 2024;10(2):e24094.
CrossRef - Chakravarthy, , Bharanidharan, N., Khan, S.B. et al. Multi-class Breast Cancer Classification Using CNN Features Hybridization. Int J Comput Intell Syst. 2024;17:191
CrossRef - Soumik MFI, Aziz AZB, Hossain Improved Transfer Learning Based Deep Learning Model For Breast Cancer Histopathological Image Classification. In: 2021 International Conference on Automation, Control and Mechatronics for Industry 4.0 (ACMI); 2021;1-4.
CrossRef - Mahmud MI, Mamun M, Abdelgawad A Deep Analysis of Transfer Learning Based Breast Cancer Detection Using Histopathology Images.
- Sumitha A, Isaac Transfer Learning-based CNN Model for the Classification of Breast Cancer from Histopathological Images. Int J Adv Comput Sci Appl. 2024;15(4).
CrossRef - Yari Y, Nguyen TL, Nguyen HT. Deep Learning Applied for Histological Diagnosis of Breast Cancer. IEEE Access. 2020;8:162432-
CrossRef - Zuluaga-Gomez J, Al Masry Z, Benaggoune K, Meraghni S, Zerhouni A CNN-based methodology for breast cancer diagnosis using thermal images. Comput Methods Biomech Biomed Eng Imaging Vis. 2020;9(2):131-145.
CrossRef - Murtaza G, Shuib L, Wahab AWA, Mujtaba G, Mujtaba G, Nweke HF, Al-garadi MA, Zulfiqar F, Raza G, Azmi NA. Deep learning- based breast cancer classification through medical imaging modalities: state of the art and research challenges. Artif Intell Rev. 2020;53:1655-1720.
CrossRef







