Manuscript accepted on :26-03-2026
Published online on: 13-05-2026
Plagiarism Check: Yes
Reviewed by: Dr. Nadhim M. H.
Second Review by: Dr. Amritlal Mandal
Final Approval by: Dr. Prabhishek Singh
Department of Biological Sciences, Faculty of Science, Mutah University, Karak, Jordan
Corresponding Author E-mail: Yousef.hajaya@mutah.edu.jo
Abstract
Non-invasive prenatal testing (NIPT), a genetic test performed during pregnancy that analyzes cell-free fetal DNA (cfDNA), has become an integral component of prenatal screening for trisomy 21 (Down syndrome), trisomy 18 (Edwards syndrome), and trisomy 13 (Patau syndrome). Nevertheless, real-world data from Middle Eastern populations, particularly from Jordan, remain limited. The aim of this study is to evaluate the prevalence and distribution of trisomies 13,18, and 21 detected by NIPT in a regional prenatal screening population in Jordan. Additionally,to compare the observed distribution with contemporary international evidence reported in the current era of NIPT.This retrospective cohort analyzed 260 NIPT reports obtained from two medical laboratories in Jordan. Data extracted from laboratory records, including maternal age, gestational age at testing, fetal fraction, pregnancy type (IVF vs non-IVF), and screening outcomes for common autosomal trisomies (Trisomy 21, Trisomy 18, and Trisomy 13), and sex chromosome abnormalities. Screening outcomes were summarized using descriptive statistics. Welch’s t-test was used to compare groups. Mann-Whitney U test and Fisher’s exact test for continuous and categorical variables, respectively.Among 260 screened pregnancies, 250 (96.2%) were reported as low risk, whereas 10 (3.85%) were classified as high risk (screen-positive). Common autosomal trisomies (Trisomy 21, Trisomy 18, and Trisomy 13) accounted for nine cases (3.46%), while one case (0.38%) corresponded to a sex chromosome aneuploidy (47, XXY; Klinefelter syndrome). Of all results, trisomy 21 was the most frequently detected abnormality (1.92%) followed by trisomy 13 (1.15%) and trisomy 18 (0.38%).Mean fetal fraction was lower among screen-positive cases compared with low-risk cases (7.30% vs 10.02%, P = 0.005). However, using Mann-Whitney U test, this difference did not remain statistically significant (P = 0.082).No statistically significant association was detected between screening outcomes and IVF status or advanced maternal age.In this cohort, NIPT demonstrated screening outcomes consistent with global NIPT reports, with trisomy 21 as the most commonly identified abnormality. These results provide real-world evidence supporting the clinical integration of NIPT into prenatal screening settings in Middle Eastern populations, particularly in Jordan.
Keywords
Cell-free DNA; Jordan; NIPT; Prenatal Care;Trisomy 13; Trisomy 18; Trisomy 21; Trisomy Prevalence
| Copy the following to cite this article: Al-Hajaya Y. Prevalence of Trisomies 13,18 and 21 Detected by Cell-Free DNA-Based NIPT in Jordanian Prenatal Screening Cohort. Biomed Pharmacol J 2026;19(2). |
| Copy the following to cite this URL: Al-Hajaya Y. Prevalence of Trisomies 13,18 and 21 Detected by Cell-Free DNA-Based NIPT in Jordanian Prenatal Screening Cohort. Biomed Pharmacol J 2026;19(2). Available from: https://bit.ly/4uHv5u1 |
Introduction
Cell-free fetal DNA (cffDNA) fragments werefirstdiscovered in maternal blood circulation in 1997.1The majority of cffDNA originates from the cytotrophoblast and then moves as a result of maternal blood circulation.1-3In 2011, non-invasive prenatal testing (NIPT) analyzed circulating cffDNAfragments using massive parallel sequencing to identify the genetic variants that represent chromosomal abnormalities.4,5NIPT was introduced into clinical settings and became widely available.6,7Clinical trial data based on actual clinical experience and multiple systematic reviews have demonstrated the effectiveness of NIPT in screening for the most common fetal chromosomal abnormalitiesthatoccur at birth, including trisomies 21 (Down syndrome), 18 (Edward syndrome) and 13 (Patau syndrome).8-13Indeed, several studies have reported that NIPT canreducethe incidenceofunnecessary invasive diagnostic procedures such as amniocentesis and chorionic villus sampling andminimizethe unnecessary risk of exposing all pregnant women.5,14,15
Numerous large-scale validation studies and meta-analyses have establishedthatclinical validity of cfDNA screening. It has been demonstrated that NIPT achieves very high diagnostic performance, with reported sensitivity exceeding 99% for trisomy 21 in appropriate clinical settings and similarly high specificity (98.80%).16-19NIPT also has high performance metric for trisomy 18 and trisomy 13,withsensitivitiesof98.83% and 100 %,and specificities of 99.93% and 96.96%,respectively.13,20
Although extensive evidence on the analytical performance of NIPT is available, understanding the prevalence of trisomies detected in local screening populations remains crucial. The observed presence of chromosomal aneuploidies inthescreening cohort is affected not only by underlying biological incidence but alsobythe maternal age distribution, referral indications, gestational age at sampling, timing of testing, access to prenatal care, and clinical pathways for test utilization.14,21,22 Thus, regional epidemiological data may help inform genetic counseling, anticipate demand for confirmatory diagnostic approaches and specialized fetal medicine services, and contextualize screening local outcomes relative to international benchmarks.19,20
One of the strongest biological risk factors for nondisjunction events leading to common autosomal trisomies is maternal age. The effect of increasing maternal age on trisomies has been extensively studied. Several studies have demonstrated an increased risk of trisomies (21,13, and 18) with advanced maternal age, although clinically significant aneuploidy is not limited to older pregnant individuals but can occur across all maternal age groups.14,21,23 In parallel, implementation studies have shown that the distribution of positive results may change as NIPT becomes more widely used among average-risk populations.22,24
Modern reviews underscore that cfDNA-based NIPT, while highly accurate for detecting common autosomal trisomies, remains a non-diagnostic rather than a definitive test. Numerous biological factors may contribute to discordant NIPT results, including confined placental mosaicism, maternal mosaicism, vanishing twin syndrome, and maternal copy-number variants.25-27Such limitations emphasize the importance of confirmatory diagnostic testing after high-risk screening and the need for appropriate patient counseling and careful result interpretation.
However, although the widespread global adaptation of NIPT, published data characterizing screening outcomes and trisomy prevalence intheJordan population remain relatively limited, highlighting the need for region-specific epidemiological evidence to better inform clinical practice, refine genetic counseling and healthcare planning in local clinical setting. Therefore, the present study aimed to evaluate the prevalence and distribution of trisomies 13,18, and 21 detected by cfDNA-based NIPT in a regional prenatal screening population in Jordan. Additionally, to compare the observed distribution with contemporary international evidence reported in the current era of NIPT.22,24,28
Materials and Methods
Study design and reporting
This retrospective observational cohort study analyzed the results of non-invasive prenatal testing (NIPT) performed in routine clinical practice. NIPT reports were collected from two accredited diagnostic laboratories in Jordan: Medlabs Consultancy Group and Precision Medical Laboratory.
Study population
A total of 260 NIPT reports were included in the analysis. All the samples were obtained from women with singleton pregnancies. NIPT results for trisomies 21, 18,and 13 were eligible for inclusion.
NIPT procedure and outcomes
Maternal peripheral blood was collected in two streak cfDNA tubes and processed for cfDNA extraction.NIPT test was performed using massively parallel next-generation sequencing (NGS) method, as documented by the participating laboratories. However, the specific sequencing platform was not revealed in the laboratory NIPT outcomes.Chromosomal risk classification for trisomies 21, 18,and 13wasidentified based on the laboratory NIPT reports. NIPT test outcomes were classified as low risk or high risk.Low-risk results indicate no increased risk for any of the chromosomal conditions assessed, whereas high-risk results indicate an increased risk of a fetal chromosomal abnormality.
Variables
Different maternal variables were extracted from laboratory recordsincludingmaternal age (years), gestational age at sampling (weeks), fetal fraction (%), fetal sex, and pregnancy status (in vitro fertilization or natural), and the final reported NIPT risk classification. Advanced maternal age was defined as maternal age ≥ 35 years at the time of testing.
Statistical analysis
Continuous variables are presented as mean ± standard deviation, median, and range. Categorical variables are reported as frequencies and percentages. Prevalence of trisomies 21,18, and 13 was calculated with Wilson ninety-five percent confidence intervals (95% CI) as the proportion of high-risk findings between all screened pregnancies. Analyses were descriptive and conducted to support epidemiological interpretation and comparison with published cohorts.For non-parametric comparisons, Mann-Whitney U test was used. Statistical significance was defined as P < 0.05. All statistical analyses were performed using SPSS version 26.0 (IBM).
Results
Study Population Characteristics
A total of 260 NIPT screens performed at Medlabs consultancy group and Precision medical lab in Jordan were included in the cohort study. All samples were obtained from women with singleton gestations. The Mean maternal age was 33.5 (range, 20-48)and 41.5% (108/260) of cases were classified as advanced maternal age (≥ 35 years). The median gestational age at NIPT was 11 weeks and 3 days (Interquartile range [IQR] 11.0-13.0 weeks). The mean fetal fraction across the cohort was 9.92 ±4.78%.The majority of pregnancies (91.2%; 237/260) were conceived naturally, whereas 8.8% (23/260) were conceived through in vitro fertilization (IVF) pregnancies. Baseline demographic and clinical characteristics stratified by screening outcomes are shown in Table 1.
Distribution of NIPT Screening Outcomes
Of the 260 NIPT results, 250 (96.15%) were reported as low risk for the tested conditions and did not indicate a trisomy of chromosomes 21, 18, 13, or sex chromosome aneuploidies (SCAs), while 10 cases (3.85%) were classified ashigh risk for aneuploidy. Autosomal trisomies accounted for 9 of the 10 screen-positive results (3.46%) while one case (0.38%) corresponded to a SCAs(47, XXY; Klinefelter syndrome). The most commonly detected abnormality was trisomy 21, identified in 5 cases (1.92%), followed by trisomy 13 in 3 cases (1.15%) and trisomy 18 in one1 (0.38%). The distribution of NIPT screening outcomes is presented in Table 4, and the relative frequencies of screen-positive findings is illustrated in Figure 1.
Association between fetal fraction and screening outcome
Mean fetal fraction was lower between screen-positive cases compared with low-risk cases (7.30± 2.34% vs 10.02 ± 4.83%, respectively). Statistical comparisons via Welch’s t-test showed a significant difference between the groups (P = 0.005). However, the non-parametric analysis using the Mann-Whitney U test did not reach statistical significance (P = 0.082). suggesting that this result should be interpreted with caution.Detailed descriptive statistics for fetal fraction stratified by screening outcome are presented in Table 5, and the distribution of fetal fraction values within the groups is illustrated in Figure 2.
Subgroup analysis
Among women with advanced maternal age (≥ 35 years), the screen-positive rate was 4.63% (5/108), compared with 3.29% (5/152) among those aged < 35 years. This difference was not statistically significant (Fisher’s exact test P = 0.746).Of the 23 IVF pregnancies included in the cohort, none were screen-positive, whereas 10 of 237 non-IVF pregnancies (4.22%) yielded screen-positive results (Fisher’s exact test, P = 0.607). Results of subgroup comparisons are summarized in Table 3.
Table 1: Baseline demographic and clinical characteristics of the study populationstratified by non-invasive prenatal testing (NIPT) screening outcome.
| Characteristic | Overall (n=260) | Low risk (n=250) | High risk (n=10) | P value |
| Maternal age (years), mean ± SD | 33.53 ± 5.67 | 33.47 ± 5.67 | 35.10 ± 5.67 | 0.394 |
| Gestational age (weeks), median (IQR) | 11.3 (11.0-13.0) | 11.1 (11.0-13.0) | 14.0 (13.0-16.5) | < 0.001 |
| Fetal fraction (%), mean ± SD | 9.92 ± 4.78 | 10.01 ± 4.83 | 7.30 ± 2.34 | 0.005 |
| IVF pregnancy, n (%) | 23 (8.8) | 23 (9.2) | 0 (0.0) | 0.607 |
| Advanced maternal age (≥ 35 years), n (%) | 108 (41.5) | 103 (41.2) | 5 (50.0) | 0.746 |
| Fetal sex male, n (%) | 124 (47.7) | 117 (46.8) | 7 (70.0) | 0.201 |
Table 2: Prevalence of screen-positive findings for autosomal trisomies and sex chromosome aneuploidies (Wilson 95% CI)
| Finding | n | N | Percent (%) | 95% CI (%) | |
| Lower | Upper | ||||
| Trisomy 21 | 5 | 260 | 1.92 | 0.82 | 4.42 |
| Trisomy 18 | 1 | 260 | 0.38 | 0.07 | 2.15 |
| Trisomy 13 | 3 | 260 | 1.15 | 0.39 | 3.34 |
| Sex chromosome aneuploidy (XXY) | 1 | 260 | 0.38 | 0.07 | 2.15 |
| Any screen-positive finding | 10 | 260 | 3.85 | 2.10 | 6.93 |
| Any autosomal trisomy (21/18/13) | 9 | 260 | 3.46 | 1.83 | 6.45 |
Table 3: Subgroup analysis of screening outcomes according to maternal age category and IVF status (Wilson 95% CI).
| Subgroup | N | High risk (n) | Percent (%) | 95% CI (%) |
| maternal age≥ 35 years | 108 | 5 | 4.63 | 1.99-10.38 |
| Maternal age < 35 years | 152 | 5 | 3.29 | 1.41-7.47 |
| IVF pregnancies | 23 | 0 | 0.00 | 0.00-14.31 |
| Non-IVF pregnancies | 237 | 10 | 4.22 | 2.31-7.59 |
Table 4: Distribution of NIPT screening outcomesacrossthe study cohort.
| Outcome | n | % |
| Low risk | 250 | 96.15 |
| High risk- T21 | 5 | 1.92 |
| High risk- T18 | 1 | 0.38 |
| High risk- T13 | 3 | 1.15 |
| High risk-SCA | 1 | 0.38 |
| Total High risk | 10 | 3.85 |
| Total | 260 | 100 |
Table 5: Comparison of fetal fraction values between low-risk and screen-positive cases.
| Group | N | Mean ± SD (%) | Median (IQR) (%) | Min-Max (%) | P value (Welch t-test) | P value (Mann-Whitney U) |
| Low risk | 250 | 10.02 ± 4.83 | 9.00 (6.92-11.67) | 3.72-43.06 | ||
| High risk | 10 | 7.30 ± 2.34 | 8.05 (7.01-8.29) | 2.00-10.69 | 0.005 | 0.082 |
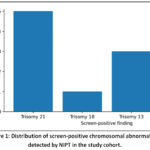 |
Figure 1: Distribution of screen-positive chromosomal abnormalities detected by NIPT in the study cohort. |
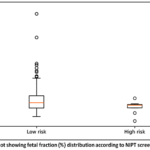 |
Figure 2: Boxplot showing fetal fraction (%) distribution according to NIPT screening outcome. |
Discussion
The current study evaluated the prevalence of trisomies 21,18, and 13 detected by cell-free DNA-based NIPT in a regional prenatal screening population in Jordan. Among the screened pregnancies, the overall screen-positive prevalence for the three trisomies was 3.46%. Trisomy 21 represented the most frequently detected chromosomal aneuploidy. These results are largely consistent with the established epidemiological distribution of fetal abnormalities reported in large-scale prenatal screening cohorts, in which trisomy 21 accounts for the majority of chromosomal abnormalities identified during pregnancy.1,10,26
The predominance of trisomy 21 in this study is consistent with well-established epidemiological observations reported in prenatal genetics. Population-based studies have consistently demonstrated that trisomy 21 is the most frequent chromosomal abnormality detected during pregnancy, whereas trisomy 13 and trisomy 18 occur at lower frequencies.19,29,30Evidence from large implementation studies of cell-free DNA screening further supports the notion that trisomy 21 typically accounts for the majority of screen-positive NIPT outcomes. The detection rate for trisomy 21 has exceeded 99% in many large-scale NIPT cohorts. Its prevalence estimates generally ranging from about 0.1% and 2% depending on the maternal age distribution of the screened population and the clinical indications for testing.28,31,32 In the current study, the relative distribution of trisomies 21,18, and 13 reflects the expected biological pattern of autosomal trisomiesin prenatal screening populations.
The prevalence pattern reported in the current study is also comparable in direction, while not identical in magnitude, to recent large-scale NIPT series. In a cohort of 118,969 pregnant women in China, Qin and co-workers reported that the positive rates of trisomy 21, trisomy 18, and trisomy 13 increased with maternal age, with trisomy 21 showing the most pronounced age-related risk, reaching 1.27% in the oldest maternal age strata.33 Likewise, in a real-world experience in Thailand, Chansriniyom et al. analyzed 17,714 NIPT results and reported an incidence of common trisomies of 0.37%, with NIPT sensitivity at 97.92%, 94.12%, 85.71% for trisomy 21, trisomy 18, and trisomy 13, respectively.34 The relatively higher proportion of screen-positive results observed in this study likely reflects variations in baseline maternal age structure, population risk composition, and referral practices rather than a fundamentally different biological distribution of common autosomal trisomies.
The prevalence estimates reported in this cohort also fall within ranges described in previous real-world NIPT studies. International clinical validation studies involving tens of thousands of pregnant women have reported overall screen-positive rates between approximately 2% and 5% when cfDNA analysis is implemented in routine clinical practice.9,18,35 Variability in reported trisomy prevalence across different studies may reflect variations in maternal age distribution, referral practices, and the underlying risk characteristics of the screened populations. In the present cohort, more than 40% of the study population was categorized as being of advanced maternal age, which may partly explain the relatively higher proportion of screen-positive findings observed in this cohort. Maternal age remains one of the most consistently recognised risk factors for fetal chromosomal nondisjunction, with the incidence of trisomy 21 increasing markedly with advancing maternal age.36,37
Interestingly, although the screen-positive rate was higher among pregnant women aged ≥ 35 years in the present cohort, no statistically significant association between advanced maternal age and screening outcome was observed in this study.This result may be attributable to the relatively small number of screen-positive cases in the study population, which, in turn, limits the statistical power to detect the significant associations in subgroup comparisons. In addition, the expanding use of NIPT between pregnancies considered to be at average risk has also altered the epidemiological profile of detected aneuploidies. As cfDNA screening becomes more widely adapted as a first-line test, clinically relevant chromosomal abnormalities are increasingly identified across all maternal age groups rather than being restricted to high-risk pregnancies.38-40 These observations reflect the evolving role of NIPT as a population-level prenatal screening tool rather than a test limited to pregnancies with advanced maternal age.
A notable finding of the current study was a lower fetal fraction in screen-positive cases compared with low-riskpregnancies. However, this difference should be interpreted with caution. Although Welch’s-test indicated statistical significance, the non-parametric Mann-Whitney U test did not reach statistical significance, which may reflect the small sample size of the high-risk group and the potential influence of outliers. Therefore, the detected difference in fetal fraction should be considered a trend rather than definitive evidence of an association.Fetal fraction, defined as the proportion of fetal-derived cfDNA in maternal plasma, is a key determinant of NIPT detection efficacy.41-44 Sufficient fetal fraction is essential to enable detection chromosomal abnormalities using massively parallel sequencing procedures, whereas low fetal fraction may compromise test sensitivity and lead to inconclusive results or reduced test accuracy.45,46
Several factors influence fetal fraction, including gestational age, maternal age, maternal body mass index, placental function, and fetal aneuploidy.40,47,48Gestational age represents a pivotal potential confounder in this retrospective analysis as the fetal fraction is positively correlated with increasing gestational age. In the current study, the significantly higher gestational age observed in the high-risk group may have partially affected fetal fraction measurements and should therefore be considered when interpreting intergroup differences.
Fetal DNA accounts approximately 3-20% of fetal circulating plasma DNA during early pregnancy, although considerable interindividual variation has been reported.42,48-50 Lower fetal fraction values have been found to be associated with an increased test failure rates or no-call results and have been associated with an increased risk of rare chromosomal abnormalities, and reduced signal-to-noise ratios in cell-free DNA sequencing-based NIPT platforms.51,52It should be noted that the presence of outliers in the low-risk group, with fetal fraction values extending to higher ranges, may have influenced mean estimates and contributed to the observed differences among groups. Such extreme values may skew parametric analyses and should therefore be considered when interpreting the findings.
The reduced fetal fraction observed between screen-positive cases in the current study may reflect biological differences in placental DNA release or underlying placental dysfunctions associated with aneuploid pregnancies.26,48 Many studies further support the central analytical importance of fetal fraction in cfDNA-based screening.In a large 2024 study of 106, 846 samples, Benn and co-workers reported that fetal fraction measurement is a key quality-control parameter and should be routinely reported as part of NIPT.53 Similarly, Xie et al. demonstrated that minimum fetal fraction levelsstrongly influence trisomy 21 detection performance in low-depth wholegenome sequencing-based NIPT methods.54 Collectively, these data support the interpretation that the lower fetal fraction observed in the screen-positive group in the preset study is biologically and analytically relevant rather than incidental.
Importantly, the vast majority of cell-free DNA present in maternal blood originates from the placental trophoblast cells rather than directly from fetal tissues.1,3 Therefore, placental biology plays a pivotal role in determining NIPT signal patterns. Biological phenomena such as confined placental mosaicism (CPM), in which chromosomal abnormalities are present in the placental tissue while the fetus’schromosomes are normal, may influence cell-free DNA test results and contribute to discordant test findings.55,56 CPM has been recognized as one of the most common biological explanations for discordant or false-positive NIPT results and highlights the importance of confirmatory diagnostic testing following high-risk screening outcomes.26,57Recent studies have also refined the interpretation of discordant screening outcomes. Studies examining mosaic signals in NIPT indicate that the relative contribution of aneuploid cfDNA fragments-often described as mosaic ratio-can influence the positive predictive value of high-risk cfDNA results, particularly for trisomies 21,18, and 13.58 These observations are clinically important because CPMrepresents a major cause of discordant or false-positive NIPT findings, especially for trisomies 18 and 13. Recognition of this limitation is essential during post-test counseling and highlights the need for confirmatory diagnostic testing following high-risk screening results.59,60
In this cohort, subgroup of pregnancies conceived through in vitro fertilization (IVF) did not detect any screen-positive cases. While this result was not statistically significant, it is generally in agreement with previous reports indicating that the clinical performance of cfDNA-based NIPT is comparable among pregnancies conceived via assisted reproductive procedures and those conceivednaturally.14Some studies have reported uncertainvariances in fetal fraction between IVF and naturally conceived pregnancies, potentially reflecting variations in placental development or maternal characteristics, however, the clinical significance of these observations remains uncertain.61 Given the relatively small number of IVF pregnancies included in the current study, further large studies would be essential to more definitively evaluate potential associations between assisted reproductive technologies and NIPT screening outcomes.
From a clinical perspective, the findings of this study support the expanding role of NIPT as an effective screening approach for common autosomal trisomies (21, 18, and13). Several professional organizations, including the American College of Obstetricians and Gynecologists (ACOG) and the Society for Maternal-Fetal Medicine (SMFM), now identify cell-free DNA screening as the most sensitive and specific non-invasive screening method for trisomies 21,18, and 13.14 Compared with combined first-trimester screening strategies, NIPT demonstrates markedly higher detection rates and substantially lower false-positive rates,18,19 leading to a significant reduction in unnecessary invasive diagnostic procedures.62,63
The current study provides valuable region-specific evidence demonstrating NIPT screening outcomes in a Jordanian cohort. While various large-scale investigations have evaluated NIPT performance in different regions such as Europe, North America, and East Asia, published data from Jordanian population remain limited. Variations in prenatal screening practices, referral pattern and maternal age distribution may influence the observed prevalence and distribution chromosomal abnormalities across populations. Consequently, region-specific studies as the present study provide important epidemiological insights that may help inform local clinical practice, support the development of evidence-basedprenatal screening strategies, and contribute to the expanding global literature on NIPT implementation.
Study limitations
Numerous limitations should be acknowledged when interpreting the findings of the current study. First, confirmatory invasive diagnostic testing (e.g. amniocentesis or chorionic villus sampling) and pregnancy outcome follow-up were not consistently available for all screen-positive cases, which prevented formal assessment of diagnostic performance measurements such as sensitivity, specificity, and positive predictive value.Second, the retrospective study design is limited the availability of certain maternal clinical variables known to influence fetal fraction, including maternal body mass index and placental-related factors. Additionally, the cohort size limits precision of estimates for low-frequency outcomes, resulting in wider Confidence intervals (Cls) for the prevalence of trisomies 13 and 18. Third, the relatively small number of screen-study population was derived from two laboratories within asingle country, which may limit the broader generalizability of the findings to other populations or healthcare settings.Accordingly, future prospective multi-center investigationsintegrating confirmatory diagnostic outcomes and comprehensive maternal clinical data are therefore necessary to further characterize the epidemiological patterns and clinical performance of NIPT within Middle Eastern populations, particularly in Jordan.
Conclusion
In a regional prenatal screening cohort of 260 singleton pregnancies, NIPT demonstrated screening outcome distributions consistent with international implementation reports, with trisomy 21 representing the majority of high-risk outcomes (5/260, 1.92%),followed by trisomy 13 (3/260, 1.15%) and trisomy 18 (1/260, 0.38%). Lower fetal fraction was significantly associated with screen-positive cases, while advanced maternal age and IVF conception were not. These findings provide valuable real-world evidence supporting the clinical implementation of cfDNA-based NIPT in modern prenatal screening programs within a Middle Eastern population. Additionally, the findings presented in this study contribute crucial population-specific data regarding the prevalence of common autosomal trisomies within a Middle Eastern population, particularly in Jordan.
Acknowledgement
The author would like to acknowledge Medlabs Consultancy Group and Precision Medical Lab, Jordan, for providing access to anonymized laboratory data used in this study. The author would also like to thank Mutah University, Jordan for facilitating the ethical approval process and academic support.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
Ethical approval for this study was obtained from the Research Ethics Committee, Mutah University, Jordan (Approval No. 120/10/751).
Informed consent statement
As this study involved retrospective analysis of fully anonymized laboratory data, individual informed consent was waived by the institutional Review Board in accordance with local regulations.
Clinical Trial Registration
This study does not involve any clinical trials.
Permission to reproduce material from other sources
Not applicable
Author Contributions
The sole author was responsible for the conceptualization, methodology, data collection, analysis, writing, and final approval of the manuscript.
References
- Lo YD, Corbetta N, Chamberlain PF, et al. Presence of fetal DNA in maternal plasma and serum. Lancet. 1997;350(9076):485-487.
CrossRef - Sifakis S, Koukou Z, Spandidos DA. Cell-free fetal DNA and pregnancy-related complications. Mol Med Rep. 2015;11(4):2367-2372.
CrossRef - Hui L, Bianchi DW. Fetal fraction and noninvasive prenatal testing: what clinicians need to know. Prenat Diagn. 2020;40(2):155-163.
CrossRef - Chiu RWK, Akolekar R, Zheng YW, et al. Non-invasive prenatal assessment of trisomy 21 by multiplexed maternal plasma DNA sequencing: large-scale validity study. BMJ. 2011;342:c7401.
CrossRef - Jayashankar SS, Nasaruddin ML, Hassan MF, et al. Non-invasive prenatal testing (NIPT): reliability, challenges, and future directions. Diagnostics (Basel). 2023;13(15):2570.
CrossRef - Chandrasekharan S, Minear MA, Hung A, Allyse M. Noninvasive prenatal testing goes global. Sci Transl Med. 2014;6(231):231fs15.
CrossRef - Hartwig TS, Ambye L, Sørensen S, Jørgensen FS. Discordant non-invasive prenatal testing (NIPT): a systematic review. Prenat Diagn. 2017;37(6):527-539.
CrossRef - Dar P, Curnow KJ, Gross SJ, et al. Clinical experience and follow-up with large-scale single-nucleotide polymorphism–based noninvasive prenatal aneuploidy testing. Am J Obstet Gynecol. 2014;211(5):527.e1-527.e17.
CrossRef - Gil MM, Quezada MS, Revello R, Akolekar R, Nicolaides KH. Analysis of cell-free DNA in maternal blood in screening for fetal aneuploidies: updated meta-analysis. Ultrasound Obstet Gynecol. 2015;45(3):249-266.
CrossRef - Norton ME, Jacobsson B, Swamy GK, et al. Cell-free DNA analysis for noninvasive examination of trisomy. N Engl J Med. 2015;372(17):1589-1597.
CrossRef - Zhang H, Gao Y, Jiang F, et al. Non-invasive prenatal testing for trisomies 21, 18 and 13: clinical experience from 146958 pregnancies. Ultrasound Obstet Gynecol. 2015;45(5):530-538.
CrossRef - Mackie FL, Hemming K, Allen S, Morris RK, Kilby MD. The accuracy of cell-free fetal DNA-based non-invasive prenatal testing in singleton pregnancies: a systematic review and bivariate meta-analysis. BJOG. 2017;124(1):32-46.
CrossRef - Rose NC, Barrie ES, Malinowski J, et al. Systematic evidence-based review: the application of noninvasive prenatal screening using cell-free DNA in general-risk pregnancies. Genet Med. 2022;24(7):1379-1391.
CrossRef - American College of Obstetricians and Gynecologists; Society for Maternal-Fetal Medicine. Screening for fetal chromosomal abnormalities: ACOG practice bulletin No. 226. Obstet Gynecol. 2020;136(4):e48-e69.
CrossRef - Cerreta V, Musti MA, Simonazzi G, et al. Screening for major aneuploidies at 10 weeks of gestation in an unselected population: comparing the combined test to the non-invasive prenatal testing with rolling-circle replication (SAPERER study). Eur J Obstet Gynecol Reprod Biol. 2026:115035.
CrossRef - Bianchi DW, Parker RL, Wentworth J, et al. DNA sequencing versus standard prenatal aneuploidy screening. N Engl J Med. 2014;370(9):799-808.
CrossRef - Norton ME, MacPherson C, Demko Z, et al. Obstetrical, perinatal, and genetic outcomes associated with nonreportable prenatal cell-free DNA screening results. Obstet Gynecol Surv. 2024;79(3):146-148.
CrossRef - Taylor-Phillips S, Freeman K, Geppert J, et al. Accuracy of non-invasive prenatal testing using cell-free DNA for detection of Down, Edwards and Patau syndromes: a systematic review and meta-analysis. BMJ Open. 2016;6(1):e010002.
CrossRef - Gil MM, Accurti V, Santacruz B, Plana MN, Nicolaides KH. Analysis of cell-free DNA in maternal blood in screening for aneuploidies: updated meta-analysis. Ultrasound Obstet Gynecol. 2017;50(3):302-314.
CrossRef - Hui L, Ellis K, Mayen D, et al. Position statement from the International Society for Prenatal Diagnosis on the use of non-invasive prenatal testing for the detection of fetal chromosomal conditions in singleton pregnancies. Prenat Diagn. 2023;43(7):814-828.
CrossRef - Savva GM, Walker K, Morris JK. The maternal age-specific live birth prevalence of trisomies 13 and 18 compared to trisomy 21 (Down syndrome). Prenat Diagn. 2010;30(1):57-64.
CrossRef - van der Meij KR, Henneman L, Sistermans EA. Non-invasive prenatal testing for everybody or contingent screening. Prenat Diagn. 2023;43(4):443-447.
CrossRef - Nicolaides KH. Screening for fetal aneuploidies at 11 to 13 weeks. Prenat Diagn. 2011;31(1):7-15.
CrossRef - Sebire E, Rodrigo CH, Bhattacharya S, Black M, Wood R, Vieira R. The implementation and impact of non-invasive prenatal testing (NIPT) for Down’s syndrome into antenatal screening programmes: a systematic review and meta-analysis. PLoS One. 2024;19(5):e0298643.
CrossRef - Benn P, Borrell A, Chiu RWK, et al. Position statement from the Chromosome Abnormality Screening Committee on behalf of the Board of the International Society for Prenatal Diagnosis. Prenat Diagn. 2015;35(8):725-734.
CrossRef - Bianchi DW, Chiu RWK. Sequencing of circulating cell-free DNA during pregnancy. N Engl J Med. 2018;379(5):464-473.
CrossRef - Samura O, Okamoto A. Causes of aberrant non-invasive prenatal testing for aneuploidy: a systematic review. Taiwan J Obstet Gynecol. 2020;59(1):16-20.
CrossRef - Dar P, Jacobsson B, MacPherson C, et al. Cell-free DNA screening for trisomies 21, 18, and 13 in pregnancies at low and high risk for aneuploidy with genetic confirmation. Am J Obstet Gynecol. 2022;227(2):259.e1-259.e14.
CrossRef - Bevilacqua E, Gil MM, Nicolaides KH, et al. Performance of screening for aneuploidies by cell-free DNA analysis of maternal blood in twin pregnancies. Ultrasound Obstet Gynecol. 2015;45(1):61-66.
CrossRef - Gregg AR, Skotko BG, Benkendorf JL, et al. Noninvasive prenatal screening for fetal aneuploidy, 2016 update: a position statement of the American College of Medical Genetics and Genomics. Genet Med. 2016;18(10):1056-1065.
CrossRef - Liang D, Cram DS, Tan H, et al. Clinical utility of noninvasive prenatal screening for expanded chromosome disease syndromes. Genet Med. 2019;21(9):1998-2006.
CrossRef - Li Y, Yang X, Zhang Y, et al. The detection efficacy of noninvasive prenatal genetic testing (NIPT) for sex chromosome abnormalities and copy number variation and its differentiation in pregnant women of different ages. Heliyon. 2024;10(2).
CrossRef - Qin S, Zhao Y, Deng F, et al. Performance evaluation of noninvasive prenatal testing on 24 chromosomes in a cohort of 118969 pregnant women in Sichuan, China. J Int Med Res. 2024;52(9):03000605241274584.
CrossRef - Chansriniyom N, Panburana P, Chantratita W, et al. Clinical performance of expanded non-invasive prenatal testing for aneuploidies: a real-world experience in Thailand. BMC Pregnancy Childbirth. 2025.
CrossRef - De Graaf G, Buckley F, Skotko BG. Estimation of the number of people with Down syndrome in the United States. Genet Med. 2017;19(4):439-447.
CrossRef - Palomaki GE, Kloza EM, Lambert-Messerlian GM, et al. DNA sequencing of maternal plasma to detect Down syndrome: an international clinical validation study. Genet Med. 2011;13(11):913-920.
CrossRef - Ashoor G, Syngelaki A, Poon LCY, Rezende JC, Nicolaides KH. Fetal fraction in maternal plasma cell-free DNA at 11–13 weeks’ gestation: relation to maternal and fetal characteristics. Ultrasound Obstet Gynecol. 2013;41(1):26-32.
CrossRef - Wang E, Batey A, Struble C, Musci TJ, Song K, Oliphant A. Gestational age and maternal weight effects on fetal cell-free DNA in maternal plasma. Prenat Diagn. 2013;33(7):662-666.
CrossRef - Rava RP, Srinivasan A, Sehnert AJ, Bianchi DW. Circulating fetal cell-free DNA fractions differ in autosomal aneuploidies and monosomy X. Clin Chem. 2014;60(1):243-250.
CrossRef - Hou Y, Yang J, Qi Y, et al. Factors affecting cell-free DNA fetal fraction: statistical analysis of 13661 maternal plasmas for non-invasive prenatal screening. Hum Genomics. 2019;13(1):62.
CrossRef - Chiu RWK, Chan KCA, Gao Y, et al. Noninvasive prenatal diagnosis of fetal chromosomal aneuploidy by massively parallel genomic sequencing of DNA in maternal plasma. Proc Natl Acad Sci U S A. 2008;105(51):20458-20463.
CrossRef - Canick JA, Palomaki GE, Kloza EM, Lambert-Messerlian GM, Haddow JE. The impact of maternal plasma DNA fetal fraction on next generation sequencing tests for common fetal aneuploidies. Prenat Diagn. 2013;33(7):667-674.
CrossRef - Deng C, Liu S. Factors affecting the fetal fraction in noninvasive prenatal screening: a review. Front Pediatr. 2022;10:812781.
CrossRef - Martínez FJF, Mira MMG, González CG, Bajo IM, Ionescu RO, Alonso CO. NIPT of maternal plasma-originated cfDNA: applications and guide for the implementation. Appl Clin Genet. 2025;18:41.
CrossRef - Shen S, Qi H, Yuan X, Gan J, Chen J, Huang J. The performance evaluation of NIPT for fetal chromosome microdeletion/microduplication detection: a retrospective analysis of 68588 Chinese cases. Front Genet. 2024;15:1390539.
CrossRef - Garg RK, Kumar Y, Niwas R, Singh J. Non-invasive prenatal testing (NIPT): a paradigm shift in prenatal care. Int J Prev Med. 2025;16:86.
CrossRef - Zhou Y, Zhu Z, Gao Y, et al. Effects of maternal and fetal characteristics on cell-free fetal DNA fraction in maternal plasma. Reprod Sci. 2015;22(11):1429-1435.
CrossRef - Deng C, Liu J, Liu S, et al. Maternal and fetal factors influencing fetal fraction: a retrospective analysis of 153306 pregnant women undergoing noninvasive prenatal screening. Front Pediatr. 2023;11:1066178.
CrossRef - Lo YD, Chan KCA, Sun H, et al. Maternal plasma DNA sequencing reveals the genome-wide genetic and mutational profile of the fetus. Sci Transl Med. 2010;2(61):61ra91.
CrossRef - Ashoor G, Syngelaki A, Wagner M, Birdir C, Nicolaides KH. Chromosome-selective sequencing of maternal plasma cell-free DNA for first-trimester detection of trisomy 21 and trisomy 18. Am J Obstet Gynecol. 2012;206(4):322.e1-322.e5.
CrossRef - Norton ME, Baer RJ, Wapner RJ, Kuppermann M, Jelliffe-Pawlowski LL, Currier RJ. Cell-free DNA vs sequential screening for the detection of fetal chromosomal abnormalities. Am J Obstet Gynecol. 2016;214(6):727.e1-727.e6.
CrossRef - Ren Y, Hao N, Chang J, et al. Clinical implications of noninvasive prenatal testing failures due to low fetal fraction: associations with adverse maternal and fetal outcomes. Prenat Diagn. 2025;45(11):1417-1425.
CrossRef - Benn P, Zhang J, Lyons D, Xu W, Leonard S, Demko Z. Accuracy of fetal fraction measurements in a single-nucleotide polymorphism-based noninvasive prenatal test. Prenat Diagn. 2024;44(10):1218-1224.
CrossRef - Xie X, Yin W, Li F, Xuan S, Ouyang Y. Analysis of fetal fraction in non-invasive prenatal testing with low-depth whole genome sequencing. Heliyon. 2025;11(1).
CrossRef - Lee TJ, Rolnik DL, Menezes MA, McLennan AC, da Silva Costa F. Cell-free fetal DNA testing in singleton IVF conceptions. Hum Reprod. 2018;33(4):572-578.
CrossRef - Gil MM, Galeva S, Jani J, et al. Screening for trisomies by cfDNA testing of maternal blood in twin pregnancy: update of the fetal medicine foundation results and meta-analysis. Ultrasound Obstet Gynecol. 2019;53(6):734-742.
CrossRef - Grati FR, Malvestiti F, Ferreira JC, et al. Fetoplacental mosaicism: potential implications for false-positive and false-negative noninvasive prenatal screening results. Genet Med. 2014;16(8):620-624.
CrossRef - Flowers NJ, Love CJ, Scarff KL, et al. Assessment of placental chromosomal mosaicism during prenatal cell-free DNA screening refines positive predictive values for fetal trisomy. Clin Chem. 2025;71(10):1036-1046.
CrossRef - Reilly K, Doyle S, Hamilton SJ, Kilby MD, Mone F. Pitfalls of prenatal diagnosis associated with mosaicism. Obstet Gynaecol. 2022.
CrossRef - Eggenhuizen GM, Go AT, Hoffer MJ, et al. Confined placental mosaicism detected with non-invasive prenatal testing: is there an association between mosaic ratio and pregnancy outcome? Prenat Diagn. 2024;44(12):1462-1469.
CrossRef - Society for Maternal-Fetal Medicine, Rink BD, Dugoff L, et al. Society for maternal-fetal medicine consult series #74: cell-free DNA screening for aneuploidies: updated guidance. Pregnancy. 2025;1(6):e70139.
CrossRef - Hui L, Bianchi DW. Noninvasive prenatal DNA testing: the vanguard of genomic medicine. Annu Rev Med. 2017;68:459-472.
CrossRef - van Prooyen Schuurman L, de Koning HJ, Meier E, et al. Clinical and economic impact of genome-wide non-invasive prenatal testing as a first-tier screening method compared to targeted NIPT and first-trimester combined testing: a modeling study. PLoS Med. 2025;22(11):e1004790.
CrossRef








