Manuscript accepted on :25-03-2026
Published online on: 12-05-2026
Plagiarism Check: Yes
Reviewed by: Dr. Karthikeyan and Dr. Jothiraj S
Second Review by: Dr. Sudhanshu Kumar Bharti
Final Approval by: Dr. Prabhishek Singh
Meenu Bala1 , Ayush Dogra2*
, Ayush Dogra2* , Priya Sadana1
, Priya Sadana1 and Bhawna Goyal3
and Bhawna Goyal3
1Department of Computer Science and Engineering, Chitkara University Institute of Engineering and Technology Chitkara University, Punjab, India,.
2Department of Electronics and Communication Engineering, Chitkara University Institute of Engineering and Technology,Chitkara University, Punjab, India.
3Department of Engineering, Marwadi University Research Centre, Marwadi University, Rajkot, Gujarat, India.
Corresponding Author’s Email : ayush123456789@gmail.com
Abstract
Multimodal medical image fusion (MMIF) has emerged as a prominentparadigm in clinical research and therapeutic practice. However, a single imaging modality has its diagnostic accuracy limitation, since different modalities reflect different levels of anatomical or functional details. While MMIF integrates complementary information from modalities like MRI, CT, PET, and SPECT to improve portrayal of the pathological features, it can provide more precise and informed clinical decision support. The process of merging the complementary spatial and functional features derived from diverse medical imaging sources is referred to as multimodal medical image fusion. The central goal of image fusion is to unify spatial and functional information to enhance the disease visualization and support reliable disease evaluation, facilitating interpretation for clinicians and automated systems. This article discusses the key aspects and methodological framework required for effective multimodal medical image fusion, including comparison of key contributions of existing review papers, radiographic and tomographic imaging methods, medical image fusion framework, medical image databases, fusion techniques, performance metrics, and challenges. The experiment and comparisons of some existing methods are carried out in this paper. FDCT method outperformed standard algorithms in most fusion metrics. A comparative evaluation of state-of-the-art fusion approaches, including CNP-NSST, ADCPNN, GoD, NSST-AGPCNN, and FDCT, is carried out on two sets of multimodal medical images, MRI/CT and T-1 MRI/CT. The comparisons demonstrate notable performance divergencies among the methods, examined through standard quantitative metrics comprising SSIM, EN, SD, QAB/F, and Mutual Information (MI).The reviewpresents a structured analysis of MMIF techniques, facilitating the directions for future breakthroughs in medical imaging and clinical decision making.Although advanced technologies are introduced, the issues of contrast enhancement, inadequate retention of fine-grained details, and spectral distortion remain to challenge the overall performance. The advantages and limitations of the existing methods are outlined, providing information to guide future progress in various clinical processes.
Keywords
Deep Learning Domain; Frequency Domain; Fusion Quality Metrics; Hybrid Domain; Pixel-Level Domain; Radiographic Modalities; Sparse Representation
| Copy the following to cite this article: Bala M, Dogra A, Sadana P, Goyal B. Multimodal Medical Image Fusion: A Review of Imaging Modalities, Databases, Fusion Strategies and Performance Validation. Biomed Pharmacol J 2026;19(2). |
| Copy the following to cite this URL: Bala M, Dogra A, Sadana P, Goyal B. Multimodal Medical Image Fusion: A Review of Imaging Modalities, Databases, Fusion Strategies and Performance Validation. Biomed Pharmacol J 2026;19(2). Available from: https://bit.ly/3R2WVCI |
Introduction
Image processing, which emerged in the 1960s, incorporates the digitization of visual data and the utilization of computational techniques designed to enhance and extract valuable details from images. Numerous techniques have been applied, including medicine, the arts, and TV image enhancement. Among its diverse applications, image processing is an important aspect of medical imaging, enabling disease detection as well as the development of an effective treatment strategies.
Medical imaging involves generating a visual representation of anatomical components for clinical examination, medical intervention, and visualizing the operation of organs or tissues. With the aid of digital image analysis methods in the medical domain, allows radiologists to swiftly identify abnormalities. In 1895, X-rays were discovered for the first practical medical imaging application. After X-rays, many other technologies have emerged, including ultrasound, CT scans in the 1970s, MRI in the 1980s, PET, and SPECT. Each modality performs a specific function in a particular application to detect bone fractures and dental checks. In contrast, PET and SPECT detect functional and metabolic activity, while ultrasound, CT and MRI imaging techniques are employed to identify the internal anatomy of organs.1, 2 Therapeutic procedures typically involve different anatomical regions within the human anatomy that are often difficult to diagnose using a single modality. The doctor needs complementary information about the patient’s lesion for a more accurate diagnosis. Therefore, information acquired by multiple imaging sources can be integrated for generating a new image, which helps doctors achieve better results and informed decision-making. The primary objective of image fusion in healthcare is to combine multimodality images to enhance image clarity, decrease noise, and provide better visibility of significant features to generate a high-quality image.3 The fused image should adhere to the outlined standards: preservation of all the original information of the source images is indispensable.4,5 The introduction of fabricated details should be avoided. Noise and misregistration should be avoided to ensure image fidelity.5 An increasing volume of research publications in periodicals, books, and magazines reflects the significant interest and relevance of multimodal image integration.
Fig. 1 depicts the annual distribution of publications related to multimodal medical image fusion, retrieved from Google Scholar, a prominent indexing platform for scholarly publications.
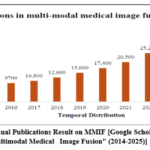 |
Figure 1: Annual Publications Result on MMIF [Google Scholar: Keyword “Multimodal Medical Image Fusion” (2014-2025)]
|
Table 1: Summary of key attributes of review works in multimodal medical image fusion
|
Author, Year |
Imaging Sources |
MMIF Approach |
Fusion Process |
Objective Assessment |
Subjective Assessment |
Database Discussions |
Comparison of Fusion Methods |
|
Osama s. Faragallah 6 |
✔ |
✔ |
🗶 |
✔ |
✔ |
🗶 |
✔ |
|
Huang, B 7 |
🗶 |
✔ |
🗶 |
✔ |
✔ |
🗶 |
✔ |
|
Tirupal 8 |
✔ |
✔ |
✔ |
✔ |
✔ |
🗶 |
✔ |
|
Adeel Azam 9 |
✔ |
✔ |
✔ |
✔ |
✔ |
✔ |
✔ |
|
Sajid Ulla Kha 10 |
✔ |
✔ |
✔ |
✔ |
✔ |
✔ |
✔ |
|
Haribabu 11 |
✔ |
✔ |
✔ |
✔ |
✔ |
✔ |
✔ |
|
Shatabadi Basu 12 |
✔ |
✔ |
🗶 |
✔ |
✔ |
✔ |
🗶 |
|
Our Work |
✔ |
✔ |
✔ |
✔ |
✔ |
✔ |
✔ |
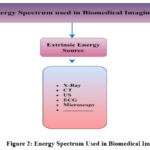 |
Figure 2: Energy Spectrum Used in Biomedical Imaging
|
The fig. 2 illustrates a typology of the energy spectrum used in biomedical imaging, divided into three main categories: intrinsic, extrinsic, and mixed intrinsic/extrinsic sources of energy.9,10
Intrinsic sources are imaging methods that enable clinicians to detect biological and chemically generated signals within the body using injected tracers, as observed in diagnostic imaging platforms such as Positron Emission Tomography (PET), Single Photon Emission Computed Tomography (SPECT), and thermography.
Extrinsic energy sources are employed externally to interact with soft tissue using non-invasive methods, such as X-rays, ultrasound (US), and Computed Tomography (CT), to produce images of anatomical structures. Hybrid imaging modalities (e.g., PET/CT, PET/MRI, and SPECT/CT) comprise the third category, as they synergistically combine internal emission with external energy input to enhance anatomical and functional visualization capabilities.
The objective of this article includes the following key aspects
Analyze various medical imaging modalities and the energy spectrum used in biomedical imaging, as illustrated in Fig. 2.
To provide an overview of open accessible multimodal medical imaging datasets and their applications in recent medical studies, as summarized in Table 3.
To present the general framework utilized in multimodal medical image integration.
To elaborate the sequential steps involved in the multimodal medical image fusion process.
To evaluate and illustrate the commonly used quality assessment metrics for measuring fusion performance.
To identify the current issues and suggest prospective avenues for future research in the domain of multimodal medical image fusion.
Table 1 illustrates a synthesized comparative analysis of significant review articles in the multimodal medical image fusion domain. This table involves the spectrum sources of image modalities, image fusion approaches, fusion process, objective and subjective assessment scope, and comparative analysis of existing fusion approaches. However, some of the reviews mentioned do not describe these attributes. In contrast to earlier studies, our review article addresses all the key parameters listed in Table 1.
The subsequent sections are laid out in the following structure: Section 2 presents a granular analysis of radiographic and tomographic imaging modalities. The clinical applications of medical image fusion are briefly discussed in Section 3. Section 4 discusses the free access online multimodal medical databases. The framework for multi-modal medical image fusion is described in Section 5. Section 6 has explored the multi-modal medical image fusion techniques. Section 7 covers the eleven performance evaluation metrics, encompassing quantitative and qualitative metrics. The review is wrapped with remarks in Section 8.
An in-depth understanding of multimodal medical image fusion requires a comprehensive knowledge of medical imaging modalities. The subsequent section demonstrates various radiographic and tomographic imaging techniques along with their diagnostic capabilities and existing challenges.
Radiographic and Tomographic Imaging Modalities
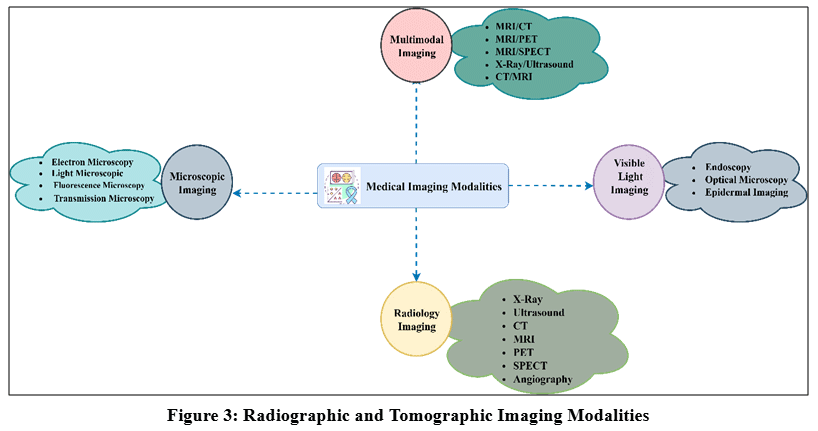 |
Figure 3: Radiographic and Tomographic Imaging Modalities
|
The healthcare sector is swiftly transforming, and by incorporating cutting-edge technologies, medical imaging now employs a diverse range of modalities for image acquisition. Each modality has its distinctive properties which facilitate physiological system analysis, disease diagnosis, treatment monitoring, and multi modality image fusion to get a more accurate decision for the specific lesion. Biomedical imaging is organized into four categories: Visible light imaging, microscopy imaging, radiology, and multimodal imaging. Visible light imaging employs wavelengths to capture the anatomical and structural properties. The key applications of visible light imaging are Endoscopy, Optical microscopy, and epidermal imaging, which facilitate high spatial resolution imaging of superficial anatomical structures and subcutaneous tissues. Microscopic imaging uses various microscopies such as electron microscopy, light microscopy, fluorescence microscopy, and Transmission microscopy to visualize the cellular and molecular architecture. 8,12 Multimodal imaging encompasses the synthesization of anatomical and functional imaging to obtain complementary information for accurate interpretation of the disease. Radiology utilizes diverse forms of radiations and magnetic fields, including Magnetic Resonance Imaging, X-Rays, Computed Tomography (CT), Ultrasound, Digital Subtraction Angiography (DSA), Single Photon Emission Computed Tomography (SPECT).8 Table 2 depicts the different radiology modalities, including their applications and challenges.
Table 2: Diagnostic Imaging Methods: A Comparative Snapshot
|
Modalities |
Imaging Mechanism |
Diagnostic Nature |
Application |
Challenges |
|
X-Rays (X-Radiations) 4 |
Electromagnetic radiation, for bones and certain soft tissues |
Non-Invasive |
Bone fractures, tumor detection Dental Treatment, Pneumonia |
Limited Image Resolution, Radiation Dose Risk, and Overlapping Structure leads to Diagnostic Error |
|
US (Ultrasound) 11 |
Sound Waves, for Structural and Functional |
Non-Invasive |
Pregnancy Monitoring, Gall Bladder Disease, Breast Lumps, Prostate Issue, Blood Flow Problems |
Low Contrast, High Spatial Resolution, limited Penetration |
|
CT (Computed Tomography) 9–11 |
X-Ray radiations, for Bones and hard tissues |
Non- Invasive |
Tumor detection, Bone Tumor Surgery, Deep Brain Simulation |
Image Noise, Noise due to motion, Low contrast resolution, Radiation dose |
|
MRI (Magnetic Resonance Imaging) 6,8,12,13 |
Use radio waves to generate slices of human body part, for soft tissue, blood vessels, breast, ligament tears |
Non- Invasive |
Cancer Assessment and Diagnosis, Surgery Planning, Lung Disease Detection, Liver Disease Detection |
Low Signal to Noise Ratio, Patient Movement, metallic implants, Image distortion |
|
PET (Positron Emission Tomography) 6,8,12,13 |
Used to inject medicines into patient’s vein to make detail, use ionize and molecular modality, For tissues and organs |
Invasive |
Gross tumor volume detection, cancer treatment |
Patient movement, limited spatial resolution, Accuracy |
|
SPECT /Gamma Camera (Single Photon Emission Tomography) 6,8,12,13 |
Cross sectional radiotracer images within human body are generated by ionizing, for detection of blood circulation |
Invasive |
Neck, head, cancer detection, tumor detection, fusion of different modality images |
Technical Limitation, allergic complications, Image artifacts |
|
DSA (Digital Subtraction Angiography) 6,8,12,13 |
A catheter is used to inject the contrast agent after that X-Rays are taken |
Invasive |
Dural Fistulas, Stenosis, Abnormal Anatomy of Vessels |
Invasiveness, Long Examination Time |
A detailed knowledge of radiographic and tomographic modalities establishes the foundation for recognizing appropriate datasets employed in fusion studies. Adequate multimodal medical imaging datasets are crucial for the effective implementation and validation of fusion algorithms. The next section delineates publicly accessible datasets, their imaging origins, and their significance to multimodal medical image fusion research.
Materials and Methods
Multimodal medical imaging databases
Before venturing into multimodal medical image fusion (MMIF) methods, one needs to know the nature and extent of the publicly accessible multimodal medical imaging databases. The databases are made up of multi-directional medical scans taken of the single subject using multisensor imaging techniques. While various databases exist, this research considers six commonly used and freely available repositories, each containing thousands of images collected using different imaging methods. The chosen databases are: The Cancer Imaging Archive, Open Access Series of Imaging Studies, Alzheimer’s Disease Neuroimaging Initiative, Whole Brain Atlas, Michigan Institute of Data Science, and the Digital Database for Screening Mammography (DDSM).
(https://github.com/sfikas/medical-imaging- datasets?tab=readme-ov-file).
Apart from these, another publicly accessible database with images of various anatomical areas presenting a variety of pathological conditions also exists. Furthermore, it should be noted that medical images can be presented in different formats depending on the acquisition system and modality employed.
TCIA (The Cancer Imaging Archive)
TCIA offers open access to systematically arranged cancer imaging datasets for medical imaging research and academic research. The dataset is structured into ‘collections’ aligned around particular diseases, imaging modality or research domain. The TCIA repository utilizes the standard file format DICOM for radiology images, also including patient follow-ups, therapeutic data, genomic and expert analysis.
OASIS (Open Access Series of Imaging Studies)
OASIS provides unrestricted access to brain neuroimaging data for biological research. OASIS1 consists of non-longitudinal MRI scans from 416 people, aged 18 to 96 years, encompassing young, older adults, and middle-aged and across various cognitive states. In each scanning session, every participant undergoes 3 to 4 T1-weighted MRI sequences. The cohort comprises right-handed individuals. The cohort of 100 participants over the age of 60 was diagnosed with Alzheimer’s disease, exhibiting severity varying mild to medium stage. OASIS 2 comprises of Non-longitudinal MR and Longitudinal MRI Data of non- cognitive impairment and cognitive impairment Older Adults, Young, middle-aged individuals. This dataset consists of 150 participants age between 60 to 96 years. Each participant conducted imaging during at least two examinations, having a minimum interval of one year, for 373 imaging sessions. Each individual was scanned on two or more visits, separated by at least one year, for 373 imaging sessions. This dataset has 3 or 4 individual T1-weighted MRI scans which are acquired in single session for each individual, also having subjects which are right- handed individuals. The cohort consist of 72 individuals, clinically characterized as non-cognitive impairment throughout the study, 64 individuals were clinically identified with non-cognitive impairment during their screening visits and persisted during later scans, and 51 individuals with subtle to moderate stage Alzheimer’s disease. Another 14 subjects were clinically diagnosed as cognitively normal at the time of their screening visit and were consistently characterized as cognitively impaired at their follow-up visit. OASIS 3 consists of Longitudinal Multimodal Neuroimaging, Clinical, and demented datasets for Normal Ageing and Alzheimer’s Disease. OASIS-3 is a collection of historical clinical data of 1378 participants collected across various ongoing projects through the WUSTL Knight ADRC over 30 years. The dataset comprises 755 cognitively normal adults and 622 individuals exhibiting varying degrees of cognitive deterioration, with ages spanning from 42 to 95 years. PET imaging has been acquired from various tracers, PIB, AV45, and FDG. Additionally, 451 Tau PET sessions and post-processed PUP are currently available for OASIS-3 subjects in a sub-project ‘OASIS-3_AV1451’. Oasis 4 consists of a collection of 663 subjects aged 21 to 94.
ADNI (Alzheimer’s Disease Neuroimaging Inititaive)
This database comprises publicly accessible medical images of Alzheimer’s disease. Positron Emission Tomography (PET) and Magnetic Resonance Imaging (MRI) of numerous human organs are accessible, and researchers have utilized these images to assess and validate the Alzheimer’s Disease Neuroimaging Initiative (ADNI).
MIAS (Mammography Image Analysis Society)
The MIAS dataset contains 330 images of mammography images of 1024×1024 pixels size of all classes. This is unsuitable for training, but it was used for exploratory data analysis and as a supplementary test dataset.
IDC (Imaging Data Commons)
IDC is a cloud-based repository of publicly available cancer imaging data. IDC is a node within the broader NCI Cancer Research Data Commons (CRDC)opens a new tab infrastructure that provides secure access to a large, comprehensive, and expanding collection of cancer research data. All data hosted by IDC will be available publicly and contains the following types of images, including Clinical and preclinical imaging, Radiological, Digital pathology, Multispectral microscopy images, Image annotations, Parametric maps, and Measurements derived from the images, and Expert assessments of the image findings.
Table 3: Multimodal Medical Image Databases
|
Database/Digitally Accessible |
Diagnostic Modality |
Anatomical Structure |
Imaging Categories |
Image Format |
|
TCIA/2011 |
CT, MRI, Mammography, PET, Histopathology, Colon |
Lungs, Breast, Brain, Heart, Chest, Kidney |
23 collections, 3 million images |
DICOM |
|
IDC/2017 |
CT, PET, Mammography, Radiotherapy Imaging, MRI |
Breast, Brain, liver, lungs |
130 tables with 70 different collections |
DICOM |
|
MIAS/1994 |
Mammography |
Breast |
330 Scans |
PGM |
|
DDSM/ 1997 |
Mammography |
Breast |
2630 Images |
PGM |
|
AANLIB/1995 |
CT, MRI, PET, SPECT |
Brain |
2D,3D Images |
GIF |
|
ADNI/2004 |
PET, MRI, FMRI, Chemical Biomarker, Genetic |
Brain |
Cognitively Normal (CN): 2,665 examinations Mild Cognitive Impairment (MCI): 3,924 examinations Alzheimer’s Disease (AD): 1,731 examinations |
DICOM, NIFTI |
|
OASIS/2007 |
Alzheimer’s Disease, Biomarker, MR, Neuroimaging |
Brain |
OASIS 1: 416 Subjects OASIS 2: 150 Subjects OASIS 3: 1378 Subjects OASIS 4: 663 Subjects |
NIFTI |
The founding of reliable multimodal imaging databases also supports algorithmic development and facilitates various clinical studies. The following section examines practical contexts in which medical image fusion has been successfully implemented, highlighting its diagnostic and therapeutic significance across various specializations.
Medical Image Fusion Applications
Medical imaging is a crucial element in the contemporary healthcare sector and facilitates non-surgical visualization of anatomical structures to diagnose and treat a wide range of lesions. A diverse range of techniques is used to detect and monitor the diseases. Various applications of medical imaging are demonstrated as follows:
CT-PET fusion for lung cancer treatment
Medical image fusion is essential for lung cancer diagnosis, integrating CT and PET input images. The fusion of these images aids in visualizing the lungs’ anatomical structure and functional activities. In addition, the fused image facilitates the ability to detect and categorize the lung lesions. 14
MRI-ULTRASOUND fusion for diagnosis of vascular blood flow
MRI facilitates high-resolution images of blood vessels, visualization of vessel morphology, branching patterns, and structural abnormalities. Ultrasound provides real-time visualization and confirms vessel patency. Fusion of MRI-Ultrasound images aids in detecting and categorizing vascular pathologies like atherosclerosis, aneurysms, thrombosis and vascular abnormalities. In addition, it helps to select the appropriate treatment planning method. 15
CT-MRI fusion for skull-based surgery
With the help of CT and MRI image fusion, surgeons can draw a 3-Dimensional model of the patient’s skull alongside structures, which helps to identify critical structures like blood vessels, nerves, and tumours more accurately. Surgeons can plan the surgical approach more precisely to reduce the risk of damage to surrounding structures. It helps to detect the tumour size as well as the location of the tumour. In addition, it provides guidance during surgery. 16
PET-MRI fusion for intracranial tumour diagnosis
PET-MRI fusion is a powerful tool that permits the visualization of metabolic activity, anatomical structures, and functional parameters. The doctors can obtain in depth information of the disease location and size of abnormalities which helps to boost the accuracy. 17
OSSEOUS-VASCULAR fusion for treatment of vascular related abnormalities
Osseous-Vascular fusion helps to detect the vascular related malfunctioning. This technique improves the procedure accuracy, eliminate procedural risks as well as treatment results. It provides accurate location of abnormalities. In addition, this fusion facilitates the blood vessel visualization with respect to adjacent bones. 16
Table 4: Multimodal Medical Images Applications
|
Fusion Type |
Application Area |
Modalities Involved |
Purpose & Benefits |
|
CT–PET Fusion [14] |
Lung Cancer Diagnosis |
CT and PET |
Visualizes lung anatomy and metabolic activity; improves detection and categorization of lung lesions. |
|
MRI– Ultrasound Fusion [15] |
Vascular Blood Flow Analysis |
MRI and Ultrasound |
Detects vascular Pathologies (e.g., atherosclerosis, thrombosis); combines structure with real-time imaging. |
|
CT–MRI Fusion [16] |
Skull-Based Surgical Planning |
CT and MRI |
Enables 3d skull modelling; identifies tumors, nerves, and vessels for precise and safe surgical intervention. |
|
PET–MRI Fusion [17] |
Intracranial Tumor Diagnosis |
PET and MRI |
Provides combined metabolic and structural imaging for improved tumor localization and diagnostic accuracy. |
|
Osseous– Vascular Fusion [16] |
Vascular Abnormality Treatment |
Bone and Vascular Imaging |
Enhances vessel visualization near bones, improves targeting, reduces risks, and guides treatment. |
The numerous healthcare applications of medical image integration emphasize the need for an effective analytical architecture that can incorporate multimodal input. The ensuing discussion illustrates the architecture and workflow of a multimodal medical image fusion framework, outlining its crucial elements, phases and fundamental ideas while considering the physiological consequences of fusion.
Multi-modal Medical Image Fusion Framework
Medical image fusion is a phenomenon of the integration of two modalities into a single image. Medical image fusion typically involves consists of modalities such as Magnetic Resonance Imaging (MRI), Positron Emission Tomography (PET), Computed Tomography (CT), and Single Photon Emission Computed Tomography (SPECT).
PET and SPECT pseudo-colored scans with superior resolution are employed to synthesize the functional information to acquire the responsiveness of biological functions, including cellular metabolism activity, circulatory functions to tissue, and blood flow. Alternatively, medical imaging modalities such as MRI, CT, and Ultrasound (US) are grey-scale images characterised by limited spatial resolution and are mostly employed for anatomical features. The integration of functional and structural modalities through multimodal image fusion enhances the complementary strengths of each, thereby providing more comprehensive diagnostic information and supporting clinicians in the accurate detection and evaluation of pathological conditions.11, 18
The process of medical image fusion is depicted in Fig. 4. Source images of medical imaging, such as MRI and CT, are decomposed into wavelet coefficients using the wavelet decomposition technique. Then, the appropriate fusion rule is applied to the decomposed coefficients to combine the relative information. Furthermore, the inverse of the transformation is applied to reconstruct the fused image. The fused image is required to adhere to the criteria: (a) the fused image should preserve all the original information of the source images. (b) The fused image should not generate any articulate information. (c) Noise and misregistration should be avoided.18 The Fig. 4 shows the sequence of operations for medical image fusion:
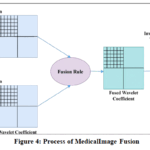 |
Figure 4: Process of Medical Image Fusion
|
In healthcare imaging, image fusion is segmented across different layers of abstraction, including pixel- level, feature-level, and decision levels to enhance the diagnostic utility.
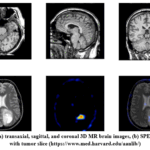 |
Figure 5: (a) transaxial, sagittal, and coronal 3D MR brain images, (b) SPECT Images with tumor slice (https://www.med.harvard.edu/aanlib/)
|
In pixel-level fusion, every intensity value of the source image is computed through basic mathematical operations to get the final image. A diverse blend of multiscale decomposition techniques can produce the final fused image. This fusion technique is employed in spatial transformation. Within the hierarchy of image fusion, at the intermediate stage, the feature level is applied to derive the detailed characteristics of an image, including intensity, edges, and smoothness of the input image. 13,19
After this, the extracted features are integrated to generate the fused image. Region-based fusion is the best example of feature-level fusion. At the apex of the image fusion abstraction level, the results of multiple algorithms are integrated to achieve a final fused decision. Decision-level image fusion is used in fingerprints and also in biometrics. 13,19
Image fusion techniques utilize various fusion rules to enhance the quality and detail of fused images, crucial for applications like medical imaging. The Activity Level Fusion Rule quantifies the quality of a particular segment of the source image by utilizing window-based measurement, coefficient-based measures, and region-based measures, with methods like the weighted average approach and the rank filter method ensuring accurate activity level evaluation. 11,20
The Coefficient Grouping Fusion Rule effectively improves image fusion results, especially when integrated with sophisticated multi-scale transforms, by utilizing relationships among coefficients to attain enhanced detail retention and visual quality of the fused image. The Cross-Scale Fusion Rule (CS) was developed to transmit information across and within each decomposition level, enabling inter-scale and intra-scale uniformity to preserve the maximum amount of detailed information while minimizing artifacts. Lastly, the Coefficient Combining Rule includes maximum selection, weighted averaging, energy-based, saliency-based, morphological consistency, and adaptive rules. 9,11 Some of the fusion rule equations are given below:
Where L (i, j) and M (i, j) denote the pixel intensity values of the two input images.
Maximum Rule
![]()
𝐼𝑖 (i, j) signifies the intensity value at the spatial coordinates (x, y) in the ith input image. Max is used to select the maximum pixel intensity value among all input images.
Weifgt Rule
![]()
𝐼𝑖 (i, j) signifies the intensity value at the spatial coordinates (x, y) in the ith input image. Wi represents the weight assigned to the ith input image. The sum of assigned weights must be 1.
N denotes the total number of images used for the fusion process.
Average Rule
![]()
If x,y signifies the average of two input images.
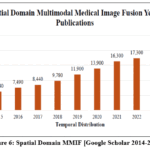 |
Figure. 6: Spatial Domain MMIF [Google Scholar 2014-2025]
|
A concise fusion framework serves as a foundation for integrating details from multiple imaging modalities. The ensuing discussion focuses on various fusion strategies that accomplish every phase of the framework, as determined by this conceptual framework. These methods, ranging from conventional transform-based models to sophisticated deep learning and hybrid methods, demonstrate the conversion of theoretical design into tangible fusion performance.
Image Fusion Techniques
This section delivers a comprehensive discussion of multi-modal image fusion approaches. Multi-modal image fusion approaches are categorized into an array of methods, depicted in Fig. 7. 21,22 In the current era, image fusion has gradually developed in the healthcare sector.
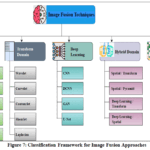 |
Figure 7: Classification Framework for Image Fusion Approaches
|
Spatial Domain
The spatial domain image fusion method deals directly with pixel values and their spatial relationships in an image. This approach relies on local features such as spatial frequency, gradient and standard deviation of input images. Various algorithms, such as the weighted average fusion method, IHS colour space fusion method, neural network fusion methods, Principal component analysis, minimum fusion method, Brovey method and maximum fusion method, have been introduced. 9,10 This method has good real-time performance, low computational complexity and high signal-to-noise ratio.
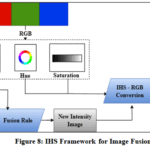 |
Figure 8: IHS Framework for Image Fusion
|
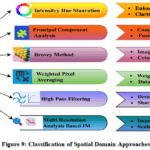 |
Figure 9: Classification of Spatial Domain Approaches
|
The spatial Domain has persistence issues, such as spectral and spatial distortion, leading to less research interest. The spatial domain literature is systematically reviewed in Table 5.
Comparative Insights
Spatial fusion provides operational efficiency and minimal latency, making it suitable for real-time or resource-limited environments. Nonetheless, direct pixel operations are prone to spectral distortion, edge blurring, and misregistration sensitivity, which might diminish edge-focused indices despite an appropriate information quantity such as an Entropy (EN) and Standard Deviation (SD). In general, spatial methods offer clear baselines but are usually surpassed in structural fidelity including information transfer rate (QAB/F) and structural similarity information (SSIM) by transform and hybrid techniques.1,23-31
To address the constraints of spatial domain approaches, the research has shifted towards the frequency domain due to its structural integrity and robustness. The subsequent section depicts the detailed analysis of frequency domain approaches.
Table 5: Literature Review of Spatial Domain Image Fusion
|
Author |
Dataset Image Type |
Techniques |
Parameters used |
Compared With |
Issue Addressed/ Identified |
|
Jingyue Chen et al.23 |
MRI, CT |
Rolling Guidance Filter, Laplacian Pyramid, Sum-modified Laplacian |
MI, SD, 𝑄𝐹 |
WT, CVT, NSCT, LP, GFF, SR |
Blurred Edges and Reduced Contrast |
|
WEISHENG Li et al.24 |
MRI-PET, MRI-SPECT |
Guided Image Filter, Spectral Residual, Graph-Based Visual Saliency, Intensity Hue Saturation |
EI, MI, 𝑄𝐹, EN, AG, QAC, SF, NIQE, TMQI |
GIHS, PCA, NSCT, GFF, LES, DSIFT, PSF, LES, CNN, DCNN |
Low Contrast Detail |
|
K. Padmavathi et al. 25 |
MRI PET |
TV-L1, Adaptive weighing scheme |
AG, MI, SD, SC, PSNR, SSIM, DIS, OP |
LP, RP, wavelet, DTCWT, CURVELET, NSCT, GFF, MMIF, LPSR |
Ringing Artifacts |
|
S. Osama et al.26 |
CT, MRI |
PCA, SVD |
AG, PSNR, STD, EN, MI, UIQI, 𝑄𝐹, LC, ES, SSIM, FSIM |
PCA, DWT, DT-CWT, Curvelet |
Processing Time, Memory Requirements |
|
Li et al.27 |
CT, MRI, PET, SPECT |
Joint bilateral filter, gradient energy |
MI, QM, QS, QCB, QAB/F, QNICE |
Zhu, LP-CNN, LLF-IOI, LRD, NSST, NSCT |
Color Distortion, Contrast Reduction |
|
Zhu et al.28 |
CT, MRI, PET, SPECT |
Adaptive Co-occurrence Filter-Based Decomposition and Fusion |
PSNR, SSIM, MI, VIF. Edge Strength |
CBF, CSMCA, CSR, LE, MBEC, LPSR, NSCTSR, SAIF, TBME, DTNP, |
Fixed Filter Scale, Low Contrast, Information Loss |
|
Lin et al.29 |
MRI/CT, MRI/SPECT, MRI/PET |
IRGF, Wiener Filter, VSM-based weighted average, WLS optimization |
SF, SSIM, ESF, CF, VIF, AG |
CBF, CNN |
Inadequate Edge Preservation Fusion Artifacts and Color Distortions. |
|
Jie et al.30 |
CT/MR-T2, MR-T1/MR-T2, MR-Gad/MR-T2, MR/PET, MR/SPECT |
XDoG, SFE, and |
QMI, QNICE, QS, QSSIM, QG, QCB |
ASR, LLP, NSSTM, NSCTM, LARD, PCNNM, MLEP, LMF, DDcGAN, SDNet, U2Fusion, Swin Fusion, CFL |
Fail to retain both significant edge and energy information. |
|
Zhang et al.31 |
MR2, SPECT |
Quasi-cross bilateral filtering (QCBF) |
PSNR, SSIM, Edge Strength |
PCA, WT, NSCT, ST, CBF, CNN |
Blurred edges, artifacts, or loss of important clinical details |
|
Kaur et al.13 |
CT, MRI, MRT1, MRT2 |
Two-scale Image Decomposition, Saliency Detection |
PSNR, SSIM, MI, VIF. Edge Strength |
LP, DWT, DT- CWT, CVT, GFS, GBFMIF, ASR, S-ADE, NSST- MSMG- PCNN |
Artifacts, inadequate lighting. |
Transform Domain
In transform domain fusion, the spatial domain images are changed into the transform domain by various approaches. This approach facilitates the ability to perform selective processing on specific frequency components. The frequency domain method outperforms spatial domain methods by decomposing images into transform layer and coefficient layer. It provides enhanced images as well as reduces computational complexity. 23,32
The spatial domain techniques suffer from numerous anomalies, such as reducing the contrast of fused images, blocking effect, and spectral degradation. To overcome these anomalies, transform domain methods are introduced.
Numerous algorithms are applied in transform domain fusion for better performance, such as the Laplacian pyramid fusion algorithm, the discrete transform fusion method, the discrete cosine transform, the discrete wavelet transform method, the stationary wavelet transform and the curvelet transform method.
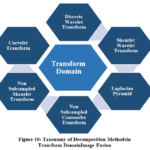 |
Figure 10: Taxonomy of Decomposition Methods in Transform Domain Image Fusion
|
As illustrated in Fig.10, the transform domain encompasses numerous multiscale decomposition methods, including Curvelet transform, Contourlet transform, Laplacian pyramid, Non- subsampled Shearlet transform, Shearlet wavelet transform, and Non-subsampled Contourlet transform, which offer smooth curves, directional features and reduce the artifacts.
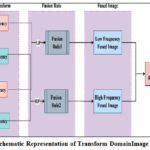 |
Figure 11: Schematic Representation of Transform Domain Image Fusion Process
|
Fig. 11 illustrates a generalized framework for image fusion for multimodality medical images such as CT, PET, MRI and ultrasound based on frequency decomposition. The transform domain is generally structured around three analytical phases: Composite Decomposition, Fusion Rule, and Image Reconstruction. Each phase contributes to ensuring the retention of vital information, such as anatomical features, metabolic function, or pathological features from each modality, and synergistically synthesizes to obtain the diagnostically enriched unified image. The process commences with the decomposition of two different modality images into approximation and detail coefficients utilizing a feasible multiscale transformation such as Contourlet transform, Discrete wavelet transforms, and non-subsampled contourlet transform. Low Frequency 33Components also known as coarse-scale features capture global intensity gradients and encode general anatomical characteristics, like the overall shape and the overall brightness.
Conversely, High Frequency 33, whose components are defined as detail coefficients, highlight localized spatial variations, guaranteeing the fine-grained anatomical information such as contours, edges, and texture information to make effective clinical decisions. Following image decomposition, specific fusion rules are used on each frequency band to combine the complementary properties of source images.
To maintain morphological information, neuroimaging fusion strategy typically uses averaging methods and weighted averaging strategies to combine low frequency bands, and use spectral frequency, regional intensity variations, saliency scoring function to enhance the contrast of the significant parts, structural contours and vascular information. Once the coefficient integration has been completed the inverse transform is used to produce the final fused image in the spatial domain.
The resultant image is then applied to a performance evaluation criterion, in which there are a great number of quality and quantity performance parameters, applied to evaluate the effectiveness and fidelity of the fusion process.
This framework is structurally versatile and can accommodate a variety of decomposition methods such as wavelet transforms, contourlets, and other multiscale techniques, making it suitable for diverse fields, especially health care imaging, surveillance, and machine learning. The significance of the transform domain is reinforced by steady growth in this domain. As depicted in Fig. 12, the publication results of multimodal medical image fusion are in the transform domain. The results were acquired from Google Scholar.
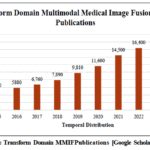 |
Figure 12: Transform Domain MMIF Publications [Google Scholar (2014-2025)]
|
Table 6: Literature Review of Transform Domain Image Fusion
|
Author |
Dataset Image Type |
Techniques |
Parameters used |
Compared With |
Issue Addressed/Identified |
|
Rao et al. 34 |
CT and MRI |
DWT, DCT Variance, DCT Variance + CV, SWT |
IQI, MIM, FF, FS, FI, RMSE, PSNR, Entropy, CC, SF |
Among themselves (DWT, DCT, CV- DCT, SWT) |
Limited retention of anatomical and functional information. |
|
Prakash et al.35 |
CT, MRI, T-1 MR, MRA |
Lifting Scheme-based biorthogonal Wavelet Transform |
Fusion Factor, Fusion Symmetry, Entropy, Standard Deviation, Edge Strength |
CLT, NSCLT, LWT, MWT, SWT. |
Noisy images |
|
Ganasala & Parsad 36 |
CT and MRI |
Stationary Wavelet Transform, Laws of Texture Energy Measures + Morphological Operations |
VIFF, RSF error, QAB/F, SSIM, Execution Time |
SWT, PCA, Max Contourlet NSCT, Phase NSST, PC NN Spatial, Fr ei-Chen NSST, NMSF |
Poor contrast, distortion, or high computation. |
|
Li et al.37 |
CT, MRI |
NSCT |
MI, IE, AvG, CC |
CL, CWT, MSGA |
low contrast, low resolution, Blurred features, image misregistration |
|
Zhou et al.38 |
PET & MRI |
IST, NSST, YUV |
AG, SF, FD, EDGE, SD, VIFF |
Wavelet, INS, NSST- BM- PCNN, MSDnet, Zero- learning, NSCT, DDcGAN |
Low brightness, edge/detail loss |
|
S.U. Khan et al.39 |
PET- MRI, SPE CT- MRI |
HIS, NSST, Structural Info, PCA |
EN, SD, SF, 𝑄AB/F MI, FMI |
NSST- PAPCNN, CNN, CSMCA, IFNN |
Scale decomposition and fusion strategy constraints |
|
Ramaraj et al. 40 |
MRI, CT |
DWT, Inverse DWT |
PSNR, Precision, SSIM, Score, Recall, Support |
CNN, NSCT, RP NN, NSST, PA PCNN, NSCT, LE |
Low resolution MRI, high contrast CT, artifacts improved |
|
Jamwal et al.41 |
CT, MRI |
Curvelet transform, Ridgelet transform, Empirical coefficient selection |
PSNR, SSIM, MI, Visual Inspection |
DWT (Discrete Wavelet Transform) PCA Brovey IHS NSCT Shearlet Deep learning methods |
Poor edge/texture retention Limited Directionality, Ineffective high-frequency component preservation |
|
Zhang et al.42 |
LSUN, STL-10, CELEBAHQ, CIFAR-10 |
Contour Wavelet Diffusion |
FID, Recall, Time |
DDGAN, DDPM, Score SDE, Fast DDPM, Recovery EBM, DDPM Distillation, CNN, NVAE, VAEBM |
Slow training and inference speed, image quality |
|
Jampani Ravi et al.43 |
Multi-focus images |
ONSST, FIPOA |
EN, SD, AG, FMI, QCB, Spatial Frequency, MEFB |
DOX, AOA, POA, DCT, DTCWT, FDCuT- FE, NSST- PAPCNN, DTCWT- WAF |
Redundancy, inconsistency, Directional Imitations Poor detail retention |
Table 6 depicts a benchmarking study of the frequency domain approaches, demonstrating their strength to preserve the anatomical and texture features involving diverse diagnosis imaging frameworks.
Comparative Insights
These methods enable the selective processing of frequency components by converting spatial domain images into the transform domain, thereby enhancing image quality and reducing computational complexity. The analyzed literature illustrates the efficacy of diverse transform domain techniques, including NSCT, wavelet transforms, and curvelet transforms, in maintaining vital details such as edges, textures, and structural information crucial for medical diagnosis more effectively than spatial fusion, as indicated by improved QAB/F and SSIM scores and enhanced MI resulting from superior cross-modal transfer. Transform domain techniques are not immune to critical limitations and challenges, such as trade-offs involving increased computation and periodic ringing in homogeneous areas when fusion rules are not optimal. In practice, transformation approaches provide a consistent benchmark for clinical-level structural preservation when execution time is feasible.34-43
To mitigate the constraints of these approaches, a recent study has shifted towards data-oriented paradigms, such as deep learning based and hybrid approaches. The following section has elaborated on deep learning methods.
Deep Learning Based Fusion
Deep learning procedures are used to enhance decision-making and performance of various tasks through the integration of prediction and parameters through various deep learning models. Multi-layer hierarchical networks are used by deep learning methods to create high-level features of images through selecting the low-level features of lines and edges using supervised and unsupervised learning methods.22 The earlier research in deep learning includes the convolutional neural network, the convolutional sparse representation, the deep Boltzmann machine and the stacked auto encoders. In 2017, Convolutional neural network was evolved for image fusion.44
Besides the Convolutional neural network, in 2019, GAN (Generative Adversarial Network) was developed for image data fusion.45 Deep learning has proven to be a powerful paradigm in medical image fusion, owing to its exceptional ability to learn intricate patterns and extract contextual details from heterogeneous medical data. Various deep learning architectures have been explored to enhance fusion quality, including convolutional neural networks (CNNs), deep belief networks (DBNs), autoencoders, residual and attention-based networks, as well as transformer-integrated hybrid models. For instance, CNNs have been effectively applied to extract spatial features for fusing multi-modality medical images with high detail preservation. DBNs have shown promise in modeling the underlying distribution of medical data.
Recent trends have seen attention-guided frameworks, lightweight MobileNet-based designs, dual-branch networks with transformer modules, and invertible architectures to losslessly transform features. Moreover, contrast learning and zero-learned models have also been proposed to enhance fusion that does not use large labelled datasets. All these developments point to the multi-purpose nature of deep learning in dealing with such critical issues as the problem of inter-modality misalignment, noise suppression, and the enhancement of semantic details.
In parallel with architectural advancements, deep learning approaches in medical image fusion are increasingly distinguished by their learning paradigms—namely, supervised, unsupervised, and semi-supervised methods. Supervised techniques rely on annotated data to train models in an end-to-end manner using task-specific targets or fused image labels. CNN-based approaches leverage supervised learning to extract the optimal fusion rule that is aligned with annotated training datasets.
The absence of a definitive reference image in multimodal image fusion hinders the reliability of supervised learning frameworks in the medical image fusion domain. An unsupervised learning framework has been proposed to improve visual fidelity by maximizing energy or structural similarities in the absence of annotated ground truth. Deep belief networks (DBN) and autoencoder-based models facilitate learning representations by transforming high-dimensional data into hidden spaces.46 Liu et al. 47 designed an unsupervised bilevel optimization framework that employs learning-based weight maps and data-driven priors to regulate the fusion process without the need of supervised information.
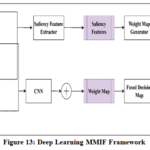 |
Figure 13: Deep Learning MMIF Framework
|
Fig. 13 depicts the framework of CNN for multimodal medical image fusion. The process begins with the acquisition of two input images acquired from various imaging sources such as CT, MRI, or PET. These images are first decomposed into coarse and detail coefficients using a multilevel decomposition technique. The coarse coefficients capture low-frequency information representing the global structure and smooth regions of the image, while the detail coefficients contain high-frequency information, such as edges and textures critical for diagnostic clarity. Different decomposition methods can be used in Deep learning, including Pyramid Decomposition, Gradient decomposition, wavelet decomposition and contourlet decomposition.
Following decomposition, two parallel processing paths are used. The low-frequency components (coarse coefficients) are fed into a traditional feature extraction module, which derives key structural descriptors. Meanwhile, the high-frequency components (detail coefficients) are processed by a trained convolutional neural network (CNN) to extract deep features that represent the most salient details 48. The results of these two streams are then added together, increasing the semantic and structural richness of the fused representation.
This is followed by a mechanism of fusion decision, which involves saliency feature calculation and weight map production. Saliency features aid in determining the most informative regions of each input image and the weight map is used to determine the contribution of each modality to the final output. These elements inform the combination of the processed features.
These fused images are then sent to the image reconstruction step, in which an inverse transform is utilized to recreate the final fused image.48 This combines the structural clarity afforded by modalities, such as MRI or CT, with the functional information afforded by modalities, such as PET or SPECT. Lastly, the resultant fuse is evaluated by its performance through qualitative visual inspection and quantitative measures.
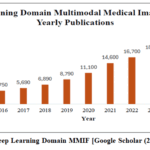 |
Figure 14: Deep Learning Domain MMIF [Google Scholar (2014-2025)]
|
Deep learning has become a revolutionary technique in biomedical imaging over the past few years, especially in disease diagnosis, segmentation, and multimodal image fusion. Fig. 14 demonstrates that the number of publications dealing with deep learning-related issues has increased dramatically, with the largest number of 21,000 articles expected in 2024, as compared to 3,790 articles in 2014.
Table 7: Literature Review of Deep Learning Domain
|
Author |
Dataset Image Type |
Techniques |
Parameters used |
Compared With |
Issue Addressed/Identified |
|
He, D. et al49 |
MRI, PET, SPECT |
Invertible Blocks, Discrete Wavelet |
SD, AG, CC, SCD, SF, VIFF, MS-SSIM, PSNR |
TL-SR, Cloud, EMFusion, MATR, CDD Fuse, M4FNet, GeSeNet EMMA, MM-Net |
Information loss, Lack of color preservation, |
|
Li, W. et al 50 |
MRI, MRI/PET, MRI/SPECT |
CNN (Res2Net) Vision Transformer |
SSIM, QMI, FMI, NCIE, QAB/F,QCB, QCV |
LES, NSCT, SVT, CSR, NSST-PACNN, IFCNN, MSENet, DSAGAN, EMFusion |
Limited Global Context in CNN, Poor retention of texture and intensity details |
|
Alnaggar et al. 51 |
CT, X-ray, PET, MRI, Ultrasound |
U-Net, CNN, SVM, ANN, |
PSNR, SSIM, MSE, Accuracy, Precision, Recall, F-measure, mIoU |
Transfer Learning Models |
Lack of efficiency analysis, Absence of standardized evaluation metrics Computational complexity and interpretability |
|
Yang et al.52
|
MRI, CT, PET, SPECT |
AGF, Res2Net (CNN) |
SF, AG, PSNR, QAB/F, FMI, MSSSIM |
CNN, CSMCA, DTCWT-SR, NSCT, NSST-PAPCNN, Union LAP, IFCNN |
Lack of annotated training data, Overfitting and convergence Parameter selection |
|
Di et al.53 |
MRI, CT, PET |
Multiscale convolution, |
AG, EN, SF, MI, SD, CC, QAB/F |
LRD, MLEPF, NSST, LEGFF, CNN, PA-PCNN, SDNet, CFL |
Loss of texture and contour detail Limited feature extraction in existing methods, High complexity |
|
Huang et al. 54 |
CT and MR-T2 |
Asymmetric dual |
PSNR, HVS ChenHmrtric, QAB/F, FMI, SSIM |
GFF, LPCNN, PAPCNN, RPCNN, FW Net, IFCNN, PMGI, U2Fusion, DDcGAN |
Brightness stacking Poor extraction of features, Limited Generalizability, High complexity |
|
Li et al. 46 |
CT, MRI, SPECT |
Deep Belief Network (DBM) |
PSNR, SSIM, MAE(HU) |
CGAN, U-Net, CycGAN, DDPM, |
Incomplete information |
|
Liu et al. 47 |
Infrared & Visible, CT, MRI & SPECT |
Bilevel optimization |
VIF, AG, EN, SF, SCD, Qab/f |
MSVD, CBF, GTF, RP, DRTV, JSRSD, WAVE, NSCT, CURLET, CSR, CSMCA, DenseFuse, Fusion GAN, DDcGAN, PAPCNN, SR, ASR |
Artifact and noise suppression, Poor performance |
As summarized in Table 7, learning fusion-based approaches have significant advancements to improve structural clarity, adaptability and semantic retention, often yielding higher MI/EN and competitive SSIM/QAB/F. The mathematical frameworks, incorporating CNNs, residual networks, attention modules, and transformers, have substantially strengthened the effectiveness and reliability of multidimensional image integration. Although these advantages exist, deep learning strategies suffer impediments. Deep learning-based fusion strategies have been limited by substantial computational aspects specifications, reliance on richly classified datasets, and a lack of interpretability elements for clinical applications, which conflicts with the accurate accountability and dependability standards of real-time healthcare applications. Such restrictions have prompted studies on hybrid approaches that aim to amalgamate the positive aspects of learning-based methods and other domains. The ensuing section addresses hybrid domain techniques developed to enhance computing effectiveness, interpretability, and overall fusion accuracy for various clinical imaging applications.49-54
Hybrid Domain
Hybrid techniques integrate spatial domain and transform domain approaches to surpass deep learning-based approaches in varied environments, attaining interpretability, accuracy, and robustness. They outperformed in retaining high-resolution details, have a lower dependency on large datasets, and have less computational complexity. These approaches were particularly superior in restricted resource environments or when data availability is limited, as they have less efficacy for overfitting and facilitate better noise resilience. Fig. 15 depicts the number of research publications in the hybrid domain multimodal medical image fusion methods per year from 2014-2025, derived through Google Scholar publications. The data demonstrate the sustained growth in publication volume and continuous technological advancements.
The paper 55 suggested a hybrid medical image fusion approach utilizing NSCT (Non-Subsampled Contourlet Transform) and DTCWT (Dual-tree Complex Wavelet Transform) techniques. This proposed approach utilized the theory of CNN for producing a weight map to combine the pixel information from dual or multimodal medical images. NSCT was used for contour and texture features, and DTCWT was used to enhance the directionality and frequency components. The hybrid fusion technique effectively maintains essential details, including edges, textures, and structural information, while minimizing noise and artefacts, rendering it especially beneficial in medical imaging modalities, including CT, MRI, and PET. The findings demonstrated that the hybrid method surpasses conventional fusion approaches, providing substantial enhancements in the quality of fused images, which is essential for precise diagnosis and clinical decision-making. an improved model 56 based on a hybrid approach consisting of non-subsampled contourlet transform (NSCT) and stationary Wavelet transform (SWT). The detailed coefficient of non- subsampled contourlet transform (NSCT) was merged using the locally weighted sum of squared coefficients with integrity validation, while the weighted sum modified Laplacian and entropy of square of coefficient fusion rules were employed for SWT. The suggested approach has better performance than the existing traditional approaches. In recent years, researchers have focused on hybrid fusion approaches yielding superior fused image quality. Figure 15 presents the annual publication trends in the domain under investigation from 2014 to 2025, where a gradual increase is observed.
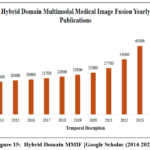 |
Figure 15: Hybrid Domain MMIF [Google Scholar (2014-2025)]
|
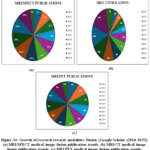 |
Figure 16: Growth of research towards modalities Fusion [Google Scholar (2014-2025)] (a) MRI/SPECT medical image fusion publication trends, (b) MRI/CT medical image fusion publication trends, (c) MRI/PET medical image fusion publication trends.
|
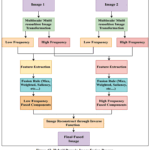 |
Figure 17: Hybrid Domain Image Fusion Process
|
Fig.17 illustrates a typical multi-resolution image fusion framework based on frequency decomposition and feature-level integration. Two source images undergo an image transformation process to decompose them into low-frequency (LF) and high-frequency (HF) components. Feature extraction is then independently applied to each frequency band of both images. The corresponding LF and HF features are subsequently combined using dedicated fusion rules to generate fused LF and HF representations. Finally, the fused components are recombined via an inverse image transformation to reconstruct the final fused image.57 This hierarchical approach enables the integration of complementary structural and textural information from multiple modalities.
Table 8: Literature Review of Hybrid Domain
|
Author |
Dataset Image Type |
Algorithm |
Parameters |
Compared With |
Issues Addressed |
|
Shukla et al.58 |
CT,MRI |
Guided Filtering, Laplacian Pyramid |
PSNR, SSIM, MSE, VIF |
NSCT-PCNN, GFF, MCNN, GFS, MCA, DWP, DPL |
Blurring, Information Loss, Loss of complementary Features |
|
Jie et al. 59 |
MRI/PET, MRI/SPECT, CT/MR-T2, MR-GAD, MR-T1/MR-T2
|
Multi-dictionary Learning, Truncated Huber Filtering |
PSNR,SSIM, RMSE, VIF |
MSMG, MLMG,LFD,CFL, LMF, DDcGAN, SDNet, IFCNN, MATR, CDD, EMFusion, U2Fusion, LRD |
Poor detail preservation and low noise robustness |
|
Zhao et al. 60 |
CT/MR-T2, MR-T1/MR-T2, MRI/PET, MRI/SPECT |
Three-Scale Decomposition, Structure Similarity Balanced Optimization |
SSIM, CC, QNICE, QMI, QG, Nab/f |
CNN, PCNN, CSMA, ConCFL, LEGFF, CNP-MIF, NSCT, NSST, XDOG, ZERO, ENfusion, DTNP, LRD |
Loss of fine details, Blurring features |
|
Moghtaderi et al.61 |
MRI, TTTc, SOECT, PET |
Adaptive Image Decomposition, Multilevel Guided Filtering |
EN, FMI, SCD, MSSIM, Piella SS, Qab/f, Qy |
MLMG-PCNN, JBF, ACOF, FGF, PCLLE, MDHU, Local extreme mapGF |
Low Adaptability, Blurred Edges, Weak Structure- Texture Sepration |
|
Venkatesan, B et al.62
|
CT, MRI |
DWT with Daubechies (db) or Symlet8 wavelets
|
SD, PSNR, MI, EPY, VAR, QAB/F
|
Pixel Avg., RSSF, Region based DT-CWT |
Blur Detail, Loss Frequency Information |
|
Sa.I.Ibrahim et al.63 |
CT, MRI |
NSC, PCNN |
PSNR, EN, MI, QAB/F, SD, AG, QNICE
|
DWT, CSR, NSCT, CSMCA, GFS, NSCT-SR, NSCT-PCLP, m-PCNN |
Information Loss, Low Contrast, Blurred Edges |
|
Xiaosong Li et.64
|
CT, Gad, PET, SPECT |
Sparse representation and neighbor energy, activity operator
|
QCB, QNICE, QG, QM, QS, QMI
|
ASR, CSMCA, CSR, KIM, KSVD, KBS, TIP, LRD, LP_SR, CNN_MLCF, INF |
Incomplete Detail Preservation, Low Contrast, Loss of Information |
|
Zhu et al,65
|
MRI/PET, CT/MRI, MRI/SPECT, MR-T1/MR-T2 |
Rolling Weighted Guided Image Filter (RWGIF), Multi-Scale Shearing Rolling Weighted Guided Image Filter (MSRWGIF)
|
MI, VIF, QCB, QAB/F, NFIM
|
CVT, DTCWT, NSCT, NSST, LLF, RGF, MSRWGIF |
Redundant Artifacts, Blurred Details, Low Contrast |
|
S. Liu et al,48
|
CT, MR-T2 |
Iterative Joint Bilateral Filter, CNN
|
MI, SSIM, Q, UIQI,VIFF, Qw, Qe, Qab/f
|
GFF, CNN, CSMCAPCNN, PCLLE, PAPCNN, RGF |
Loss of Information, Blocking Artifacts |
Comparative Insights
Table 8 explores the different hybrid approaches. Hybrid methods (NSCT, SWT, Shearlet) offer improved edge retention and contrast. Decomposition methods enhanced contrast and feature retention, resulting in sharper edges and improved contrast. Structural preservation techniques maintain spatial consistency in fused images. Hybrid methods, such as the Wavelet Transform, NSCT, and PCA, excel in medical image fusion by preserving both anatomical structures (low-frequency details) and functional information (high-frequency details), as compared to other approaches. Despite these advantages, hybrid methods still require improvements in balancing contrast enhancement and maintaining high-frequency components, as well as increasing the QAB/F and lower down the loss of information and emergence of artifacts and noise in fused result. The subsequent section discusses the sparse representation, which provides a compact, noise-resistant solution for medical image fusion, particularly in data-limited or real-time diagnostic applications.58-65
Sparse Representation
Sparse representation (SR) has become a prominent paradigm in multimodal medical image fusion, offering a principal approach for integrating complementary information from diverse imaging modalities. SR allows efficient encoding of salient structures with high fidelity and noise resistance by mapping source images (e.g., CT, MRI, or PET) to a sparse domain by projecting and encoding these images using overcomplete dictionaries. Such fusion is normally done by measuring the activity of sparse coefficients and using selective fusion strategies like the maximum coefficient rule. The process aids in the maintenance of critical anatomical and functional features and inhibition of redundant or modality-specific noise. In comparison with traditional fusion algorithms in the transform or spatial domain, SR-based algorithms have been shown to perform better in terms of preserved edges, contrast, and clinically meaningful features.66 More recently, the additions of adaptive or convolutional sparse coding (CSC) have further increased the ability of SR systems to capture complex structures and utilize data-based optimization, thereby enhancing the interpretability and clinical relevance of the resulting fused images. This work demonstrates the importance of sparse representation in medical image fusion which improves diagnostic accuracy. Fig. 18 shows that annual growth in annual publications is on an upward trend.
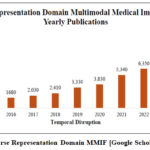 |
Figure 18: Sparse Representation Domain MMIF [Google Scholar (2014-2025)]
|
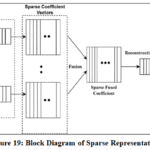 |
Figure 19: Block Diagram of Sparse Representation
|
Table 9: Literature Review of Sparse Domain
|
Author |
Dataset Image Type |
Techniques |
Parameters |
Compared with |
Issues addressed |
|
Chen et al.67
|
CT, PET, SPECT, MR-T1, MR-T2 |
CSR, SWT Dictionary learning and patch-based fusion |
QMI, QE,QCB, QG |
SR, NSST, CSR, CSMCA, CNN, IFCNN, EMFusion, IFT, Transfusion, CDDFuse, DDFM |
Lack of directionality |
|
Kittusamy et al.68 |
CT, MRI |
NSCT |
AG, EN, Sharpness SD MSE, MI PSNR |
DCT, DWT, CT |
Lack of preserving anatomical and soft tissue details, halo artifacts and color distortion |
|
Li et al.64 |
CT/T2, Gad/T2, PET/T1, PET/T2, SPECT/T2 |
SR, Neighbor Energy Activity KSVD |
QMI, QNCIE, QG, QM,QS, QCB |
CSR, CSMCA, ASR, LP-SR, KSVD, KIM, KBS, LRD, TIP |
Low retention, Color distortion and low contrastBlock artifacts and high CC |
|
Qin et al.69
|
Multi-focus microscopic images |
Improved SR, K-SVD, DWT |
EN, SF, QAB/F , |
DWT-based fusion |
Long computation time, Limited contrast, detail retention, poor high-frequency preservation,Overfitting |
|
Xu et al.70
|
MRI-CT, MRI-PET, MRI-SPECT, T1-T2, T2-PD |
Fractal Component-wiseModelling with patch-based decomposition |
AG,CC,EN,MI,SF,MSE,RMSE |
NSST-PAPCNN |
Over smoothing and noise, Redundancy Poor preservation of structural and detailed features. |
|
Liu et al.66
|
CT, MRI (T1, T2), PET, SPECT |
Local and Single-Component SR |
QLMI, QG, QE, QCB |
SR, CSR |
Patch redundancy |
Comparative Insights
Table 9 illustrates that sparse representation-based fusion methods efficiently capture vital characteristics and decrease redundancies through implementing sparsity criteria during the integration phase. Sparse frameworks enable streamlined, noise-resistant algorithms that preserve significant edges and textures with minimal data. The efficiency of sparse approaches in integration often becomes sensitive to the quality of the dictionary and the optimization factors, consequently limiting scalability across various imaging environments and potentially shifting variance or block artifacts.66-70
In the subsequent section, Table 10 presents a comparison study that provides a framework for selecting effective fusion methods aligned with specific diagnostic demands, therapeutic targets, and computational constraints.
Table 10: Advantages and limitations of various domains
|
Domain |
Advantages |
Limitations |
|
Spatial Domain 71 |
Simple implementation Computationally efficient Direct pixel-level manipulation |
Sensitive to noise and misregistration Lacks frequency or structural context Poor edge and detail preservation |
|
Transform Domain 71 |
Preserves frequency and structural features Well-suited for multi-scale analysis (e.g., wavelets, NSCT) |
A fixed basis may not adapt well to complex features Requires inverse transform High computational cost in high-resolution scenarios |
|
Sparse Representation 72
|
Compact representation of salient features Robust to noise and occlusion, Preserves edge and texture information |
Computationally intensivedictionary learning Performance depends on dictionary quality |
|
Deep Learning Domain12
|
Learns hierarchical, data-driven features Superior performance in preserving semantics and fine details, Highly adaptable |
Requires large annotated dataset High training complexity and resource requirements |
|
Hybrid Domain55 |
Combines strengths of multiple domains (e.g., spatial and transform, or SR and deep learning) |
Increased model complexity Requires careful integration and tuning Potentially higher inference time |
The recent analysis involved an in-depth overview of medical image fusion approaches that cover various methodology areas, highlighting the main principles, advantages, and limitations as outlined in the literature. Even though these approaches demonstrate promising results in the incorporation of more healthcare data, their effectiveness can be proved only by the rigorous evaluation. Therefore, a systematic and objective measurement of the quality of fusions is necessary to determine the dependability and diagnostic effectiveness of each method.
Comparative Insights of Fusion Domain
To summarise, spatial techniques are computationally efficient and simple to use, but they are not good at maintaining fine edges and other complex details. Transform-based algorithms enhance edge preservation, structural coherence, which creates an accurate clinical level standard of fusion accuracy. Deep learning models enable semantic and task-based integration with improved information transfer, and hybrid approaches are always able to reach the best balance between quantitative performance (MI, QAB/F, SSIM) and perceptual consistency of multimodal combinations. Sparse representation methods are highly adaptable and resilient when faced with noisy scenarios or sparse data, and can be effectively used in conjunction with transform-based decomposition to enhance structural stability and interpretability. Transform domain methods provide a good foundation, hybrid methods guarantee good fusion capability, deep learning methods are good at region-specific or pathology-centric tasks, and sparse representation is useful where computational simplicity and longevity is important.
The next section describes the performance evaluation parameters that are typically applied to measure the structural preservation, contrast enhancement, information content, and the overall visual quality of fused medical images.
Performance Metrics
Fusion is a complex and multifaceted process which entails not only the process of integrating multimodal information but also requires proper visualization of anatomy. In this process, artifact introduction can greatly affect the fidelity of the fused image resulting in the misinterpretation of important areas such as the presence of normal anatomical variations, which may be interpreted as pathological. To overcome these issues, different measurement criteria have been established to measure the output of image fusion algorithms in a quantitative and qualitative manner.
Generally, there are two major types of evaluation methodologies: subjective and objective evaluation. Subjective assessment is based on the expertise visualization, most often by radiologists, of the perceptual quality and diagnostic value of the fused image. Objective evaluation, on the other hand, involves computational measures of image quality.73,74 Objective methods can also be divided into two subtypes:
Reference-based evaluations. Reference-based approaches are based on a ground truth or standard image to determine the attribute of the algorithm, including clarity, feature preservation, and the overall quality and used as a reference to verify the fusion algorithm.
Non-reference-based methods, on the other hand, take the fused image by itself and use intrinsic image features, such as information content, sharpness, and spatial consistency, without a standard set. The classification of these evaluation methodologies is summarized in the table below:
 |
Table 11: Performance Metrics
|
The above metrics offer a unified framework of assessing the quality and accuracy of fused medical images. These measurements are objective measures of progress in data retention, structural integrity, and visual enhancement of different fusion methods. Based on the evaluation exercise, the following section outlines the experimental data produced by the comparable fusion methods. This is followed by a detailed discussion to clarify these results in terms of diagnostic accuracy, visual effectiveness and consistency with the accepted assessment criteria.
Experimental Results and Discussion
This section explores cutting-edge techniques for both subjective and objective analysis, utilizing a system equipped with a 12th Gen Intel Core i5-1235U processor (1.30 GHz) and a 64-bit Windows operating system, with simulations implemented in MATLAB 2018. The test was conducted using publicly available dataset available at: https://github.com/Imagingscience/Image-Fusion-Image-Denoising-Image Enhancement/commits?author=Imagingscience
Qualitative Evaluation
Fig. 20–23 provides qualitative comparisons of five multimodal medical image fusion approaches, such as CNP-NSST, ADCPCNN, GoD, NSST-AGPCNN and FDCT, with four datasets and imaging conditions. Each figure displays the respective source CT/MRI inputs, followed by fused outputs from the competing algorithms.
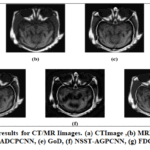 |
Figure 20: Qualitative results for CT/MRI images. (a) CT Image, (b) MRI Image, (c) CNP-NSST, (d) ADCPCNN, (e) GoD, (f) NSST-AGPCNN, (g) FDCT
|
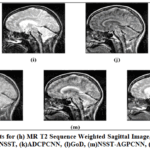 |
Figure 21: Qualitative Results for (h) MR T2 Sequence Weighted Sagittal Image, (i) MR T1 Contrast Image, (j) CNP-NSST, (k) ADCPCNN, (l) GoD, (m) NSST-AGPCNN, (n) FDCT
|
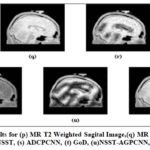 |
Figure 22: Qualitative Results for (p) MR T2 Weighted Sagital Image, (q) MR T1 Weighted Sagital Image, (r) CNP-NSST, (s) ADCPCNN, (t) GoD, (u) NSST-AGPCNN, (v) FDCT
|
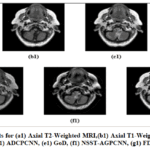 |
Figure 23: Qualitative Results for (a1) Axial T2-Weighted MRI, (b1) Axial T1-Weighted MRI, (c1) CNP-NSST, (d1) ADCPCNN, (e1) GoD, (f1) NSST-AGPCNN, (g1) FDCT
|
Fig.20(a) and Fig. 20 (b) illustrate a pair of CT and MRI images from Dataset 1. Dataset 1 comprises co-registered CT and MRI images of the upper cranial region, encompassing the skull, orbits, paranasal sinuses, and cerebral hemispheres. The CT images in this dataset provide high-resolution visualization of calcified structures, including the skull and orbital bones, with excellent bone-air contrast. However, they exhibit limited soft tissue differentiation, particularly within the cerebral parenchyma. Conversely, the respective MRI images have better soft tissue contrast, allowing good differentiation between gray and white matter. Major neuroanatomical structures like corpus callosum, basal ganglia, and brainstem are clearly defined. The lateral and third ventricles are hypointense, which is in line with cerebrospinal fluid signal features, and fall within normal morphological ranges. This data is specifically helpful in assessing intracranial anatomical integrity and subtle parenchymal characteristics, which points to the complementary advantages offered by CT and MRI modalities in imaging the brain.
In dataset 2, Fig. 21(a) shows an MR T2 sequence-weighted sagittal brain image that contains the anatomy of the brain, including the Pons verolli, silla turcica, lateral section of the cerebrum and cerebellum, as well as displays the thalamus. The cerebral cortex, corpus callosum, and ventricular system can be easily visualised as the cerebrospinal fluid (CSF) appears hypointense (dark) and fat and white-matter structures are relatively hyperintense. An MR T1 contrast-enhanced figure is shown in Fig. 21(b): giving good anatomy and a clear definition of gray and white matter borders hyperintense (bright) CSF and greater contrast in edema or fluid-rich tissues. This contrast helps to promote the view of pathological alterations and fluid collections and provides the general structural context. The complementary contrasts of T1- and T2-weighted images can be used together to demonstrate in-depth anatomical and pathological data that can be used in multimodal fusion analysis.
In the case of Dataset 3, Fig. 22(p) and Fig. 22(q) illustrates sagittal sections of the brain MRI dataset, which was obtained after two independent magnetic resonance imaging scans. Fig. 22(p) is the T2-weighted MRI sequence, which is characterised by a hyperintense cerebrospinal fluid (CSF) signal and a comparatively darker white matter signal and therefore better at detecting oedema, lesions, or abnormal fluid build-up. T2-weighted imaging is made very useful by the contrast profile to determine pathological variations, such as tumors, inflammation, or demyelination. Fig. 22(q) is a representation of the T1-weighted MRI sequence, where the cerebrospinal fluid (CSF) has been hypointense, the white matter is hyperintense, and the gray matter is intermediate. T1-weighted images depict the structural and anatomical features that will help distinguish cortical and subcortical regions more effectively and provide a superior level of spatial resolution to perform morphological analysis.
Moreover, dataset 4 that comprises Fig. 23(a1) depicts an axial T2-weighted MRI section at the posterior fossa level. The cerebral-spinal fluid (CSF) in the fourth ventricle is bright (hyperintense), and it clearly defines the ventricular space. The cerebellar hemispheres are visible with their folia, where gray–white matter differentiation can be faintly appreciated. The brainstem at the medulla–pons transition shows an intermediate signal. On either side, the cerebellopontine angle cisterns are filled with bright CSF. Anteriorly, the clivus forms part of the skull base, while posteriorly, the occipital bone is seen. Laterally and posteriorly, the paraspinal muscles are identified as dark soft tissue structures. Fig. 23 (b1) depicts an axial T1-weighted MRI at a similar level. Here, the CSF within the fourth ventricle is dark (hypointense). The cerebellar hemispheres are well outlined, with the gray matter appearing slightly darker and the white matter relatively lighter. The brainstem structures, including the pons and medulla, are seen, although their internal gray–white distinction is less sharp compared with the cerebrum. The cerebellopontine angle region is visible bilaterally. The petrous parts of the temporal bones and inner ear structures are faintly seen laterally. The surrounding muscles appear hypointense compared to fat, while the narrow spaces of the skull base demonstrate bright signal intensity due to the presence of fat on T1 imaging.
Quantitative Evaluation
After the qualitative analysis of the fused images, a rigorous quantitative analysis has been carried out to objectively compare the performance of each fusion method on four Datasets. The analysis used standard fusion quality measures, such as Entropy (EN), Standard Deviation (SD), Mutual Information (MI), Edge Preservation Index (QAB/F), and Structural Similarity Index (SSIM), summarized in Tables 12, 13, 14 and 15. All these measures are used to evaluate the information richness, contrast distribution, sharpness of edges, and structural consistency between the fused and source images. Increased EN, SD, MI, QAB/F, and SSIM tend to reflect better preservation of diagnostic information and high quality fusion. The respective quantitative findings are shown in Tables 12, 13, 14, and 15, which are a comprehensive summary of the performance of all fusion methods in the selected quality metrics.
Table 12: Performance evaluation of benchmark techniques for dataset 1
|
Methods |
Dataset 1 |
|||||
|
EN |
SD |
MI |
QAB/F |
SSIM |
Run Time (Sec) |
|
|
CNP-NSST 29 |
6.798457 |
57.60181 |
3.220991 |
0.746964 |
0.44923 |
2.49 |
|
ADCPNN 75 |
6.801396 |
57.90403 |
3.261044 |
0.743749 |
0.454734 |
2.44 |
|
GoD 76 |
6.253933 |
51.1434 |
4.359352 |
0.748024 |
0.458508 |
0.014 |
|
NSST-AGPCNN 77 |
6.296421 |
49.40166 |
1.489107 |
0.600106 |
0.294822 |
1.5 |
|
FDCT 78 |
6.10776 |
59.19149 |
5.911846 |
0.805777 |
0.487633 |
0.042 |
Table 13: Performance evaluation of benchmark techniques for dataset 2
|
Methods |
Dataset2 |
|||||
|
EN |
SD |
MI |
QAB/F |
SSIM |
Run Time (Sec) |
|
|
CNP-NSST 29 |
7.474153 |
67.31873 |
3.081312 |
0.528681 |
0.543477 |
2.33 |
|
ADCPNN 75 |
7.685867 |
87.17019 |
3.562025 |
0.529319 |
0.575824 |
2.53 |
|
GoD 76 |
7.661474 |
69.21756 |
3.746545 |
0.509759 |
0.580556 |
0.009 |
|
NSST-AGPCNN 77 |
7.506872 |
75.74939 |
3.051711 |
0.511884 |
0.54092 |
1.24 |
|
FDCT 78 |
7.760999 |
85.84002 |
7.108417 |
0.595645 |
0.532107 |
0.091 |
Table 14: Performance evaluation of benchmark techniques for dataset 3
|
Methods |
Dataset3 |
|||||
|
EN |
SD |
MI |
QAB/F |
SSIM |
Run Time (Sec) |
|
|
CNP-NSST 29 |
5.17388185 |
95.4589508 |
3.3248094 |
0.57086758 |
0.49815325 |
2.23 |
|
ADCPNN 75 |
4.95591636 |
101.045307 |
3.32710473 |
0.60027444 |
0.5067816 |
2.47 |
|
GoD 76 |
4.38766025 |
65.1905808 |
3.22506216 |
0.54132689 |
0.57356779 |
0.011 |
|
NSST-AGPCNN 77 |
5.75347537 |
77.7178027 |
2.84189841 |
0.49260704 |
0.33770025 |
1.24 |
|
FDCT 78 |
4.67577967 |
100.828061 |
5.85332333 |
0.68163775 |
0.65606163 |
0.042 |
Table 15: Performance evaluation of benchmark techniques for dataset 4
|
Methods |
Dataset 4 |
|||||
|
EN |
SD |
MI |
QAB/F |
SSIM |
Run Time (Sec) |
|
|
CNP-NSST 29 |
5.33141485 |
73.7418962 |
3.78400713 |
0.57017837 |
0.58499207 |
2.26 |
|
ADCPNN 75 |
5.44040254 |
73.7364874 |
3.71730704 |
0.55140942 |
0.55739757 |
2.42 |
|
GoD 76 |
5.212483 |
53.3555281 |
3.6112803 |
0.50726629 |
0.59501046 |
0.008 |
|
NSST-AGPCNN 77 |
5.48413632 |
58.0089557 |
2.66925343 |
0.46268038 |
0.42774432 |
1.3 |
|
FDCT 78 |
5.38284273 |
73.8454002 |
5.94656469 |
0.67401723 |
0.59275537 |
0.067 |
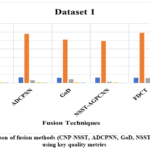 |
Figure 24: Comparison of fusion methods (CNP-NSST, ADCPNN, GoD, NSST-AGPCNN, FDCT) using key quality metrics
|
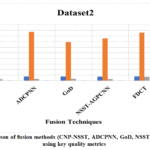 |
Figure 25: Comparison of fusion methods (CNP-NSST, ADCPNN, GoD, NSST-AGPCNN, FDCT) using key quality metrics
|
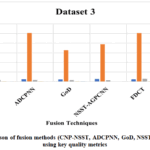 |
Figure 26: Comparison of fusion methods (CNP-NSST, ADCPNN, GoD, NSST-AGPCNN, FDCT) using key quality metrics
|
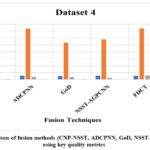 |
Figure 27: Comparison of fusion methods (CNP-NSST, ADCPNN, GoD, NSST-AGPCNN, FDCT) using key quality metrics
|
In order to compare the visual fidelity and structural preservation properties of the fusion mechanisms of Dataset 1, the outcomes of each of the techniques are presented in Figs. 21(c) to 22(g) and comparatively discussed in this section. The output image of the CNP-NSST, Fig. 21 (c), shows a good preservation of the large-scale anatomical structures and preserves overall spatial coherence. It however demonstrates poor capacity to preserve fine grained detail and demonstrates somewhat blurred tissue boundaries making it less apt to preserve high frequency textures. Conversely, Fig. 21(d) illustrates the result of ADCPCNN, which provides better edge definition and more uniform texture mapping of both the cortical and subcortical areas. A dual-channel CNN provides better modality-specific feature extraction, at the cost of a small loss of intensity balance, especially in the periventricular regions. GoD method, illustrated in Fig. 21(e) is a gradient-based methodology, which works well with respect to simple structural improvement and the ease of implementation. Nevertheless, the fused image does not contain soft-tissue contrast and is not as noisy as deep learning-based methods, which limits its clinical use, where subtle pathology must be visualized. NSST-AGPCNN, illustrated in Fig. 21(f), is efficient in integrating multiscale decomposition with attention, which leads to better saliency conservation and better emphasis of soft-tissues. Although its visual output is clearer than CNP-NSST and GoD, some inconsistencies in edge continuity and distribution of contrast are observed. FDCT, Fig. 21 (g) is the most visually harmonious and informative result of all. It has a higher quality of clarity in low-frequency anatomy background and high-frequency anatomy such as cortical margins and deep gray matter structures. The resultant fused image is more natural and sharper, and does not contain typical artifacts like blurring or modality dominance. This indicates that FDCT is better at maintaining complementary elements and yields higher diagnostically meaningful fusion results.
The effectiveness of the five fusion approaches has tested on a second independent dataset of T2 and T1-weighted MRI brain images with the same evaluation criteria used as in Dataset 1. Fig.22 (j) generated with CNP-NSST maintains the overall anatomy and spatial regions, but exhibits a slight degree of detail improvement. The resultant image is a little smoothed but finer cortical and subcortical textures are not as clearly indicated. Fig. 22(k), is generated using the ADCPCNN approach, has enhanced visual sharpness and is more effective in the retention of both CT and MRI specific information. Nevertheless, certain uneven brightness and slight imbalance fusion can be noted in areas around the ventricles and the cortical surface. The GoD technique, with Fig. 22 (l), highlights edges and boundaries, but fails to show soft-tissue. The image has a greater amount of noise and has no smooth transition of contrasts, which reduces its clinical interpretability in comparison to the other methods. As shown in Fig. 22(m), NSST-AGPCNN improves the visualization of structural tissue, as well as soft tissue, giving a clear definition of the cortex, skull, and deep brain areas. Nevertheless, some areas have slight edge artifacts and mild over-enhancement, indicating an excessive enhancement of high-gradient zones. Fig. 22 (n) obtained using the FDCT approach, reveals a superior fusion output that is effective in merging the spatial resolution of CT with the soft-tissue image of MRI. The image is of good contrast, sharpness, and anatomical accuracy especially in midline structures and brain parenchyma. It demonstrates the most harmonious combination of the complementary features with minimum distortion, and, therefore, is the most visually and diagnostically powerful of the compared methods.
The respective fusion results of Dataset 3 have been provided in Fig. 23 (p–v) and the numerical measurements have been summarized in Table 14. As compared to other methodologies, CNP-NSST (Fig. 23 r) has preserved significant anatomical borders with minor blurring and poor textural contrast. ADCPCNN (Fig. 23 s) has enhanced the continuity of edges and clarity of features by using the dual-channel convolutional learning, but localized brightness imbalance is observed around the ventricular regions. GoD (Fig. 23 t) has offered high edge-enhancement but with an appreciable noise and loss of soft-tissue appearance, and NSST-AGPCNN (Fig. 23 u) has focused on salient features but with a slight over-enhancement in high-gradient areas. Conversely, FDCT (Fig. 23 v) has produced the most visually harmonious and diagnostically consistent result, maintaining global contrast as well as fine anatomical detail with only a few artifacts.
Moreover, the results of the fusion of Dataset 4 are calculated in different ways and presented in Fig. 24 (c1-g1). The method CNP-NSST (Fig. c1) retains the global structure but has some slight blurring and moderate loss of high-frequency texture. ADCPCNN (Fig. d1) yields a better sharpness of edges and outlines of cortices using dual-channel convolutional extraction, but still has slight brightness non-uniformity in deep-grey areas. The GoD method (Fig. e1) is an efficient method to increase contrast, but it adds a little noise and over-sharp boundaries, making homogeneous tissues less smooth. NSST-AGPCNN (Fig. f1) offers acceptable soft-tissue enhancement but over-enhances bright areas resulting in some saturation of contrast. Comparatively, the best fused image, with the highest visual balance and the most diagnostic coherence, fine structure, natural contrast, and artifact-free boundaries is obtained with FDCT (Fig. g1).
In regard to both objective and subjective assessment, the findings show that both methods have their own shortcomings, in addition to their strengths, such as transform domain approaches that create artefacts at smoothing areas. Nevertheless, the hybrid techniques like FDCT have better sharpness and information preservation. Enhancing the incorporation of lightweight transformer-based structures with adaptive fusion policies to better edge preservation, artifact reduction, and real-time clinical applicability in multispectral medical imaging fusion remains achievable.. Our current study, while comprehensive in algorithmic evaluation, does not include clinical or radiologist-based validation. We therefore acknowledge this as a key limitation. Future work should integrate observer studies, clinical case-based assessments, and multicenter validation trials to evaluate how fused images influence diagnostic accuracy, inter-reader variability, and therapeutic decision outcomes. Establishing standardized clinical validation frameworks will be essential for transitioning multimodal medical image fusion from computational research to routine clinical practice.
Data Privacy
The study strictly follows internationally recognized data protection and ethical practices, including the provisions of the General Data Protection Regulation (GDPR) in the European Union and the Health Insurance Portability and Accountability Act (HIPAA) in the United States. All the datasets used in this study are publicly available and fully de-identified so that no personally identifiable information or protected health information (PHI) was accessed or processed at any point. HIPAA states that properly de-identified data are not subject to the privacy rule, whereas GDPR is concerned with transparency, rights of individuals and the safe treatment of personal health data. The two frameworks are also committed to the reduction of identities, documentation of safeguards and putting in place breach-notification protocols wherever necessary. The techniques adhere to the best practices of radiology research and largely reduce the risk of re-identification. The research used only open, de-identified datasets and did not involve direct contact with human participants, hence, institutional ethical approval and informed consent were not necessary. All research was conducted only with academic, non-commercial purposes, in accordance with modern best-practice requirements on the privacy and safety of medical imaging data.
Limitations
This study provides an in-depth discussion of the current and leading multimodal medical image fusion (MMIF) methods. MMIF is integral to present-day biomedical research, as it integrates information from various imaging modalities to enhance diagnostic accuracy and inform clinical decision-making. The core purpose of MMIF is to enhance image quality by preserving significant anatomical and functional information while minimizing artifacts and color distortions.
This paper provides an extensive overview of the various fusion approaches, fusion levels, and performance metrics. This paper describes the latest innovations and emerging directions in the field of multimodal image fusion, suggesting possible avenues for improvement and future research in medical image analysis. Each domain has distinct advantages, as well as constraints. For instance, Spatial domain integration techniques have acknowledged spatial distortions in the resultant fused image, which may degrade the quality and interpretability of the image. Spatial domain approaches often fail to effectively manage frequency information, resulting in the loss of important features across different frequency bands. According to the prior literature, the subsequent prospective areas that require further research are outlined below:
major challenge arises when input image parameters vary, as this can lead to overfitting and degrade fusion performance, underscoring the need for adaptive optimization strategies.
Deep Learning approaches introduce artifacts that may compromise diagnostic accuracy, and transformer-based methods sometimes struggle to retain edge details effectively.
Transform techniques may not effectively capture directional features, leads to omission of critical anatomical structure in the resulting image.
Hybrid methods still need improvements in balancing contrast enhancement and high-frequency component maintenance.
Future Avenue
Future research should focus on developing adaptive fusion models that can dynamically balance information retention and perceptual quality across diverse medical imaging modalities. Transformation techniques may not effectively capture directional features, potentially losing important structural information in the fused image. Certain transform methods, such as wavelet transforms, can produce artifacts like ringing effects. Deep learning-based medical image fusion has introduced significant advancements in feature extraction, detail retention, and adaptive learning. Despite these advantages, deep learning-based fusion methods also face notable limitations. Some approaches introduce artifacts that may compromise diagnostic accuracy, and transformer-based methods sometimes struggle to retain edge details effectively. Hybrid methods (NSCT, SWT, Shearlet) offer improved edge retention and contrast with reduced computational cost. The sparse representation fusion method significantly improved multi-focus image fusion, particularly in enhancing clarity, computational efficiency, and feature retention. However, computational complexity, sensitivity to noise, and limited generalization to broader medical imaging modalities remain challenges. Future studies will also focus on enhancing the fusion framework through adaptive weighting and saliency-driven approaches to emphasize regions that are diagnostically significant.79 In addition, future studies should address the challenge of modality imbalance, where high-contrast anatomical modalities (e.g., CT/MRI) often dominate and suppress metabolically informative but lower-contrast signals (e.g., PET). Developing adaptive, contrast-aware fusion strategies can ensure balanced feature integration and yield diagnostically optimal fused representations.80
Conclusion
This study presents a detailed review of multimodal medical image fusion (MMIF), emphasizing its critical role in unifying anatomical and functional information from multiple imaging modalities to enhance diagnostic accuracy. The paper has outlined several publicly available medical image databases and integration process with image fusion framework. In this review, a description of different medical imaging modalities with their imaging mechanism and their nature, applications and challenges is presented. Medical image fusion applications are also discussed, followed by an in-depth discussion of MMIF methods, spatial domain, transform domain, Deep learning domain, hybrid domain, and sparse representation. The performance metrics, Quantitative and qualitative metrics are described. Furthermore, five multimodal medical image fusion techniques are compared using the criteria of these quality assessment metrics to highlight their relative strength. At the end of this article, the current problems of multimodal image fusion methods and prospective advancements are addressed. This article analyzes that hybrid domain methods with FDCT outperforms in multiple scenarios. Despite advancements, ongoing challenges in MMIF are spatial misalignment and imaging artifacts. Such issues can be addressed by an in-depth understanding of multimodal medical image fusion techniques and the continued evolution of high-fidelity fusion algorithms.
Acknowledgement
The authors would like to thank Chitkara University, Rajpura, Punjab, India, for providing institutional support and necessary facilities during the course of this research work. The authors also extend their heartfelt appreciation to Mr. Mohit Sharma, Assistant Professor in the Department of Medical Imaging Technology, Allied Health Science, Chitkara University, Rajpura, Punjab, India, for his valuable insights and support. Additionally, the authors are deeply grateful to Nidhi Goswami, Assistant Professor at Chitkara University, Rajpura, Punjab, India, for sharing her expertise in medical radiology, drawing from her extensive 20 years of experience at PGIMER, Chandigarh, in the department of Radiodiagnosis.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials
Permission to reproduce material from other sources
Not Applicable
Author Contributions
Meenu Bala: Conceptualization, Methodology, Writing-Original Draft.
Ayush Dogra: Data Collection, Analysis, Writing – Review & Editing.
Priya Sadana: Visualization, Supervision.
Bhawna Goyal: Formal Analysis and Proof Read.
References
- James AP, Dasarathy B V. Medical image fusion: A survey of the state of the art. Information Fusion. 2014;19(1):4-19. doi:10.1016/j.inffus.2013.12.002
CrossRef - Xu F, Liu J, Song Y, Sun H, Wang X. Multi-Exposure Image Fusion Techniques: A Comprehensive Review. Remote Sens (Basel). 2022;14(3). doi:10.3390/rs14030771
CrossRef - Pavani K, Babu SBGT, Satyanarayana V. A Review on Multi-Model Medical Image Fusion Methods. AIP Conf Proc. 2022;2555:554-558. doi:10.1063/5.0108876
CrossRef - Kaur H, Koundal D, in VKA of computational methods, 2021 undefined. Image fusion techniques: a survey. Springer.
CrossRef - Bhataria KC, Shah BK. A review of image fusion techniques. In: 2018 Second International Conference on Computing Methodologies and Communication (ICCMC). IEEE; 2018:114-123.
CrossRef - Faragallah OS, El-Hoseny H, El-Shafai W, et al. A comprehensive survey analysis for present solutions of medical image fusion and future directions. IEEE Access. 2021;9:11358-11371. doi:10.1109/ACCESS.2020.3048315
CrossRef - Huang B, Yang F, Yin M, Mo X, Zhong C. A Review of Multimodal Medical Image Fusion Techniques. Comput Math Methods Med. 2020;2020. doi:10.1155/2020/8279342
CrossRef - Tirupal T, Mohan BC, Kumar SS. Multimodal Medical Image Fusion Techniques – A Review. Curr Signal Transduct Ther. 2020;16(2):142-163. doi:10.2174/1574362415666200226103116
CrossRef - Azam MA, Khan KB, Salahuddin S, et al. A review on multimodal medical image fusion: Compendious analysis of medical modalities, multimodal databases, fusion techniques and quality metrics. Comput Biol Med. 2022;144(December 2021):105253. doi:10.1016/j.compbiomed.2022.105253
CrossRef - Ullah Khan S, Ahmad Khan M, Azhar M, Khan F, Lee Y, Javed M. Multimodal medical image fusion towards future research: A review. Journal of King Saud University – Computer and Information Sciences. 2023;35(8):101733. doi:10.1016/j.jksuci.2023.101733
CrossRef - Haribabu M, Guruviah V, Yogarajah P. Recent Advancements in Multimodal Medical Image Fusion Techniques for Better Diagnosis: An Overview. Curr Med Imaging Rev. 2022;19(7):673-694. doi:10.2174/1573405618666220606161137
CrossRef - Basu S, Singhal S, Singh D. A systematic literature review on multimodal medical image fusion. Multimed Tools Appl. 2024;83(6):15845-15913.
CrossRef - Kaur H, Vig R, Kumar N, Sharma A, Dogra A, Goyal B. Multimodal Medical Image Fusion Utilizing Two-scale Image Decomposition via Saliency Detection. Curr Med Imaging Rev. 2023;20:1-13. doi:10.2174/0115734056260083230924154700
CrossRef - Kumar A, Fulham M, Feng D, Kim J. Co-learning feature fusion maps from PET-CT images of lung cancer. IEEE Trans Med Imaging. 2019;39(1):204-217.
CrossRef - Dogra A, Goyal B, Agrawal S, Ahuja CK. Efficient fusion of osseous and vascular details in wavelet domain. Pattern Recognit Lett. 2017;94:189-193. doi:10.1016/j.patrec.2017.03.002
CrossRef - Fang D, Yang C, Zhou H, et al. Effects of CT/MRI Image Fusion on Cerebrovascular Protection, Postoperative Complications, and Limb Functional Recovery in Patients with Anterior and Middle Skull Base Tumors: Based on a Retrospective Cohort Study. Contrast Media Mol Imaging. 2022;2022.
CrossRef - Odusami M, Maskeliūnas R, Damaševičius R, Misra S. Explainable deep-learning-based diagnosis of Alzheimer’s disease using multimodal input fusion of PET and MRI Images. J Med Biol Eng. 2023;43(3):291-302.
CrossRef - Dogra A, Goyal B, Agrawal S. From multi-scale decomposition to non-multi-scale decomposition methods: A comprehensive survey of image fusion techniques and its applications. IEEE Access. 2017;5:16040-16067. doi:10.1109/ACCESS.2017.2735865
CrossRef - Galande A, Patil R. The art of medical image fusion: A survey. Proceedings of the 2013 International Conference on Advances in Computing, Communications and Informatics, ICACCI 2013. Published online 2013:400-405. doi:10.1109/ICACCI.2013.6637205
CrossRef - Li S, Kang X, Fang L, Hu J, Yin H. Pixel-level image fusion: A survey of the state of the art. Information Fusion. 2017;33:100-112. doi:10.1016/j.inffus.2016.05.004
CrossRef - Kaur H, Koundal D, Kadyan V. Image Fusion Techniques: A Survey. Archives of Computational Methods in Engineering. 2021;28(7):4425-4447. doi:10.1007/S11831-021-09540-7
CrossRef - Wei M, Xi M, Li Y, Liang M, Wang G. Multimodal Medical Image Fusion: The Perspective of Deep Learning. Academic Journal of Science and Technology. 2023;5(3):202-208. doi:10.54097/ajst.v5i3.8013
CrossRef - Chen J, Zhang L, Lu L, Li Q, Hu M, Yang X. A novel medical image fusion method based on rolling guidance filtering. Internet of Things. 2021;14:100172.doi: 10.1016/j.iot.2020.100172
CrossRef - Li W, Jia L, Du J. Multi-Modal Sensor Medical Image Fusion Based on Multiple Salient Features with Guided Image Filter. IEEE Access. 2019;7:173019-173033. doi:10.1109/ACCESS.2019.2953786
CrossRef - Padmavathi K, Asha CS, Maya VK. A novel medical image fusion by combining TV-L1 decomposed textures based on adaptive weighting scheme. Engineering Science and Technology, an International Journal. 2020;23(1):225-239. doi:10.1016/j.jestch.2019.03.008
CrossRef - Faragallah OS, Muhammed AN, Taha TS, Geweid GGN. PCA based SVD fusion for MRI and CT medical images. Journal of Intelligent and Fuzzy Systems. 2021;41(2):4021-4033. doi:10.3233/JIFS-202884
CrossRef - Li X, Zhou F, Tan H, Zhang W, Zhao C. Multimodal medical image fusion based on joint bilateral filter and local gradient energy. Inf Sci (N Y). 2021;569:302-325. doi:10.1016/j.ins.2021.04.052
CrossRef - Zhu R, Li X, Huang S, Zhang X. Multimodal medical image fusion using adaptive co-occurrence filter-based decomposition optimization model. Bioinformatics. 2022;38(3):818-826. doi:10.1093/bioinformatics/btab721
CrossRef - Lin H, Song Y, Wang H, Xie L, Li D, Yang G. Multimodal Brain Image Fusion Based on Improved Rolling Guidance Filter and Wiener Filter. Comput Math Methods Med. 2022;2022. doi:10.1155/2022/5691099
CrossRef - Jie Y, Li X, wang M, Zhou F, Tan H. Medical image fusion based on extended difference-of-Gaussians and edge-preserving. Expert Syst Appl. 2023;227. doi:10.1016/j.eswa.2023.120301
CrossRef - Zhang Y, Wang M, Xia X, et al. Medical image fusion based on quasi-cross bilateral filtering. Biomed Signal Process Control. 2023;80(P1):104259. doi:10.1016/j.bspc.2022.104259
CrossRef - Mankar R, Daimiwal N. Multimodal medical image fusion under nonsubsampled contourlet transform domain. 2015 International Conference on Communication and Signal Processing, ICCSP 2015. 2015;(June):592-596. doi:10.1109/ICCSP.2015.7322555
CrossRef - C. Saranya, S. Shoba. Comparison of Image Fusion Technique by Various Transform based Methods. International Journal of Engineering Research and. 2015;V4(09):258-262. doi:10.17577/ijertv4is090321
CrossRef - Srinivasa Rao D, Ramesh Babu C, Nagendra Kumar YJ, Rajasekhar N, Ravi T. Medical image fusion using transform based fusion techniques. International Journal of Recent Technology and Engineering. 2019;8(2):309-317. doi:10.35940/ijrte.A9180.078219
CrossRef - Prakash O, Park CM, Khare A, Jeon M, Gwak J. Multiscale fusion of multimodal medical images using lifting scheme based biorthogonal wavelet transform. Optik (Stuttg). 2019;182(July 2018):995-1014. doi:10.1016/j.ijleo.2018.12.028
CrossRef - Ganasala P, Prasad AD. Medical image fusion based on laws of texture energy measures in stationary wavelet transform domain. Int J Imaging Syst Technol. 2020;30(3):544-557. doi:10.1002/ima.22393
CrossRef - Li X, Zhao J. A novel multi-modal medical image fusion algorithm. J Ambient Intell Humaniz Comput. 2021;12(2):1995-2002. doi:10.1007/s12652-020-02293-4
CrossRef - Zhou J, Xing X, Yan M, et al. A fusion algorithm based on composite decomposition for PET and MRI medical images. Biomed Signal Process Control. 2022;76(April):103717. doi:10.1016/j.bspc.2022.103717
CrossRef - Khan SU, Khan F, Ullah S, YoungmoonLee, Sami ulQudoos, Lee B. Multimodal medical image fusion in NSST domain with structural and spectral features enhancement. Heliyon. 2023;9(6). doi:10.1016/j.heliyon.2023.e17334
CrossRef - Ramaraj V, Swamy MVA, Sankar MK. Medical Image Fusion for Brain Tumor Diagnosis Using Effective Discrete Wavelet Transform Methods. Journal of Information Systems Engineering and Business Intelligence. 2024;10(1):70-80. doi:10.20473/jisebi.10.1.70-80
CrossRef - Jamwal A, Jain S. Empirical Curvelet-ridgelet Wavelet Transform for Multimodal Fusion of Brain Images. Curr Med Imaging Rev. 2024;20. doi:10.2174/0115734056269529231205101519
CrossRef - Zhang D, Li JY, Chen Z, Zou Y. Efficient image generation with Contour Wavelet Diffusion. Computers and Graphics (Pergamon). 2024;124(October 2023):104087. doi:10.1016/j.cag.2024.104087
CrossRef - Ravi J, Subbayamma B V., Kumar PV, et al. Multi-image fusion: optimal decomposition strategy with heuristic-assisted non-subsampled shearlet transform for multimodal image fusion. Signal Image Video Process. 2024;18(3):2297-2307. doi:10.1007/s11760-023-02906-3
CrossRef - Liu Y, Chen X, Peng H, Wang Z. Multi-focus image fusion with a deep convolutional neural network. Information Fusion. 2017;36:191-207.
CrossRef - Ma J, Yu W, Liang P, Li C, Jiang J. FusionGAN: A generative adversarial network for infrared and visible image fusion. Information fusion. 2019;48:11-26.
CrossRef - Kaur M, Singh D. Fusion of medical images using deep belief networks. Cluster Comput. 2020;23(2):1439-1453. doi:10.1007/s10586-019-02999-x
CrossRef - Liu R, Liu J, Jiang Z, Fan X, Luo Z. A Bilevel Integrated Model with Data-Driven Layer Ensemble for Multi-Modality Image Fusion. IEEE Transactions on Image Processing. 2021;30:1261-1274. doi:10.1109/TIP.2020.3043125
CrossRef - Liu S, Wang M, Yin L, Sun X, Zhang YD, Zhao J. Two-Scale Multimodal Medical Image Fusion Based on Structure Preservation. Front Comput Neurosci. 2022;15(January):1-14. doi:10.3389/fncom.2021.803724
CrossRef - He D, Li W, Wang G, Huang Y, Liu S. MMIF-INet: Multimodal medical image fusion by invertible network. Information Fusion. 2025;114(August 2024):102666. doi:10.1016/j.inffus.2024.102666
CrossRef - Li W, Zhang Y, Wang G, Huang Y, Li R. DFENet: A dual-branch feature enhanced network integrating transformers and convolutional feature learning for multimodal medical image fusion. Biomed Signal Process Control. 2023;80(P2):104402. doi:10.1016/j.bspc.2022.104402
CrossRef - Alnaggar OAMF, Jagadale BN, Saif MAN, et al. Efficient Artificial Intelligence Approaches for Medical Image Processing in Healthcare: Comprehensive Review, Taxonomy, and Analysis. Vol 57. Springer Netherlands; 2024. doi:10.1007/s10462-024-10814-2
CrossRef - Yang F, Jia M, Lu L, Yin M. Adaptive zero-learning medical image fusion. Biomed Signal Process Control. 2023;84(May):105008. doi:10.1016/j.bspc.2023.105008
CrossRef - Di J, Guo W, Liu J, Ren L, Lian J. AMMNet: A multimodal medical image fusion method based on an attention mechanism and MobileNetV3. Biomed Signal Process Control. 2024;96(PB):106561. doi:10.1016/j.bspc.2024.106561
CrossRef - Huang W, Zhang H, Guo H, Li W, Quan X, Zhang Y. ADDNS: An asymmetric dual deep network with sharing mechanism for medical image fusion of CT and MR-T2. Comput Biol Med. 2023;166(May):107531. doi:10.1016/j.compbiomed.2023.107531
CrossRef - Alseelawi N, Hazim HT, ALRikabi HTS. A Novel Method of Multimodal Medical Image Fusion Based on Hybrid Approach of NSCT and DTCWT. International journal of online and biomedical engineering. 2022;18(3):114-133. doi:10.3991/ijoe.v18i03.28011
CrossRef - Subbiah Parvathy V, Pothiraj S, Sampson J. A novel approach in multimodality medical image fusion using optimal shearlet and deep learning. Int J Imaging Syst Technol. 2020;30(4):847-859. doi:10.1002/ima.22436
CrossRef - Venkatesan B, Ragupathy US, Natarajan I. A review on multimodal medical image fusion towards future research. Multimed Tools Appl. 2023;82(5):7361-7382.
CrossRef - Shukla N, Sood M, Kumar A, Choudhary G. Adaptive decomposition with guided filtering and Laplacian pyramid-based image fusion method for medical applications. Discover Applied Sciences. 2024;6(8). doi:10.1007/s42452-024-06111-w
CrossRef - Jie Y, Li X, Tan H, Zhou F, Wang G. Multi-modal medical image fusion via multi-dictionary and truncated Huber filtering. Biomed Signal Process Control. 2024;88(PB):105671. doi:10.1016/j.bspc.2023.105671
CrossRef - Zhao H, Zhang X, Wang Z, Yang F, Zhu R. Multi-modal medical image fusion via three-scale decomposition and structure similarity balanced optimization. Biomed Signal Process Control. 2024;95(PA):106358. doi:10.1016/j.bspc.2024.106358
CrossRef - Moghtaderi S, Einlou M, Wahid KA, Lukong KE. Advancing multimodal medical image fusion: an adaptive image decomposition approach based on multilevel Guided filtering. R Soc Open Sci. 2024;11(4). doi:10.1098/rsos.231762
CrossRef - Venkatesan B, Ragupathy US. An Investigation on Multimodal Brain Image Fusion in the Time–Frequency Domain using Wavelet Transforms. IETE J Res. 2024;70(6):5681-5690. doi:10.1080/03772063.2023.2280670
CrossRef - Ibrahim SI, Makhlouf MA, El-Tawel GS. Multimodal medical image fusion algorithm based on pulse coupled neural networks and nonsubsampled contourlet transform. Med Biol Eng Comput. 2023;61(1):155-177. doi:10.1007/s11517-022-02697-8
CrossRef - Li X, Wan W, Zhou F, Cheng X, Jie Y, Tan H. Medical image fusion based on sparse representation and neighbor energy activity. Biomed Signal Process Control. 2023;80:104353. doi:10.1016/J.BSPC.2022.104353
CrossRef - Zhu F, Liu W. A novel medical image fusion method based on multi-scale shearing rolling weighted guided image filter. Mathematical Biosciences and Engineering. 2023;20(8):15374-15406. doi:10.3934/mbe.2023687
CrossRef - Liu Y, Chen X, Liu A, Ward RK, Wang ZJ. Recent Advances in Sparse Representation Based Medical Image Fusion. IEEE Instrum Meas Mag. 2021;24(2):45-53. doi:10.1109/MIM.2021.9400960
CrossRef - Chen Y, Liu A, Liu Y, He Z, Liu C, Chen X. Multi-Dimensional Medical Image Fusion With Complex Sparse Representation. IEEE Trans Biomed Eng. 2024;71(9):2728-2739. doi:10.1109/TBME.2024.3391314
CrossRef - Kittusamy K, Kumar LSVS. Non Sub-Sampled Contourlet with Joint Sparse Representation Based Medical Image Fusion. Computer Systems Science and Engineering. 2023;44(3):1989-2005. doi:10.32604/csse.2023.026501
CrossRef - Qin X, Ban Y, Wu P, et al. Improved Image Fusion Method Based on Sparse Decomposition. Electronics (Switzerland). 2022;11(15):1-11. doi:10.3390/electronics11152321
CrossRef - Xu G, Deng X, Zhou X, Pedersen M, Cimmino L, Wang H. FCFusion: Fractal Componentwise Modeling With Group Sparsity for Medical Image Fusion. IEEE Trans Industr Inform. 2022;18(12):9141-9150. doi:10.1109/TII.2022.3185050
CrossRef - Polinati S, Bavirisetti DP, Rajesh KNVPS, Naik GR, Dhuli R. The fusion of mri and ct medical images using variational mode decomposition. Applied Sciences (Switzerland). 2021;11(22). doi:10.3390/app112210975
CrossRef - Ghandour C, El-Shafai W, El-Rabaie S, Abdelsalam N. Comprehensive Performance Analysis of Different Medical Image Fusion Techniques for Accurate Healthcare Diagnosis Applications. Vol 83. Springer US; 2024. doi:10.1007/s11042-023-16334-5
CrossRef - Dogra A, Goyal B, Agrawal S. Medical image fusion: A brief introduction. Biomedical & Pharmacology Journal. 2018;11(3):1209.
CrossRef - Patil U, Mudengudi U. Image fusion using hierarchical PCA. ICIIP 2011 – Proceedings: 2011 International Conference on Image Information Processing. 2011;(Iciip):1-6. doi:10.1109/ICIIP.2011.6108966
CrossRef - Agrawal C, Yadav SK, Singh SP, Panigrahy C. A simplified parameter adaptive DCPCNN based medical image fusion. In: Proceedings of International Conference on Communication and Artificial Intelligence: ICCAI 2021. Springer; 2022:489-501.
CrossRef - Kurban R. Gaussian of Differences: A Simple and Efficient General Image Fusion Method. Entropy. 2023;25(8). doi:10.3390/e25081215.
CrossRef - Vajpayee P, Panigrahy C, Kumar A. Medical image fusion by adaptive Gaussian PCNN and improved Roberts operator. Signal Image Video Process. 2023;17(7):3565-3573.
CrossRef - Wang M, Shang X. A Fast Image Fusion with Discrete Cosine Transform. IEEE Signal Process Lett. 2020;27(8):990-994. doi:10.1109/LSP.2020.2999788.
CrossRef - Sharma S, Rani S, Dogra A, Shabaz M. Structure-aware medical image fusion via mean curvature enhancement in the contourlet domain. PLoS One. 2025;20(9):e0332869. doi:10.1371/journal.pone.0332869.
CrossRef - Yadav SP, Yadav S. Image fusion using hybrid methods in multimodality medical images. Med Biol Eng Comput. 2020;58(4):669-687. doi:10.1007/s11517-019-02063-9.
CrossRef
Abbreviations
Adaptive Dual-Channel Pulse Coupled Neural Network (ADCPNN), Computed Tomography (CT), Convolutional Neural Network (CNN), Coupled Neural P Systems in Nonsubsampled Shearlet Transform (CNP-NSST), Discrete Cosine Transform (DCT), Entropy (EN),Fast Discrete Cosine Transform(FDCT), Gaussian of Differences (GoD), Magnetic Resonance Imaging (MRI), Mutual Information (MI),Nonsubsampled Shearlet Transform- Adaptive Gaussian Pulse Coded Neural Network(NSST-AGPCNN), Positron Emission Tomography (PET), Single-Photon Emission Computed Tomography (SPECT), Standard Deviation (SD), Structural Similarity Index Metric (SSIM), Quality Assessment(QAB/F).







