Manuscript accepted on :23-01-2026
Published online on: 17-04-2026
Plagiarism Check: Yes
Reviewed by: Dr. Moumita Hazra
Second Review by: Dr. Nadhim M. H
Final Approval by: Dr. Eman Refaat Youness
Nargis Khanam1 , Ankur Singh2
, Ankur Singh2 , Mona Srivastava3
, Mona Srivastava3 , Abhay Kumar Yadav1
, Abhay Kumar Yadav1 , Shani Vishwakarma1
, Shani Vishwakarma1 , Anil Kumar Maurya1
, Anil Kumar Maurya1 , Suchitra Singh1
, Suchitra Singh1 , Shraddha Chaurasiya1
, Shraddha Chaurasiya1 , Surbhi Singh1
, Surbhi Singh1 , Vaibhav Jaiswal4
, Vaibhav Jaiswal4 , Khushbu Yadav1
, Khushbu Yadav1 , Pooja Dubey1
, Pooja Dubey1 , Nitish Kumar Singh5
, Nitish Kumar Singh5 , Ashish5
, Ashish5 and Royana Singh1,5*
and Royana Singh1,5*
1Department of Anatomy, Institute of Medical Science, Banaras Hindu University, Varanasi, UP, India.
2Department of Pediatrics, Institute of Medical Science, Banaras Hindu University, Varanasi, UP, India.
3Department of Psychiatry, Institute of Medical Science, Banaras Hindu University, Varanasi, UP, India.
4Department of Kaumar Bharitya/Balroga, Faculty of Ayurveda, Institute of Medical Sciences, Banaras Hindu University, Varanasi, UP, India.
5Multidisiplinary Research Unit, Institute of Medical Science, Banaras Hindu University, Varanasi, UP, India.
Corresponding Author E-mail: royanasingh@bhu.ac.in
Abstract
Autism is a Spectrum Disorder, which represents a multifaceted neurodevelopmental disorder distinguished by impairments in daily life behaviors such as communication, social engagement, and the manifestation of repetitive behaviors. The incidence of autism can be affected by variations in diagnostic criteria and demographic factors. Research indicates that trace elements such as magnesium, selenium, chromium, zinc, and copper are integral to optimal brain function, with deficiencies in these elements potentially exacerbating the characteristics of Autism spectrum disorder. Additionally, genetic variables, including copy number variations, significantly influence the etiology of Autism Spectrum Disorder. The present investigation was carried out on samples of 35 children diagnosed with Autism Spectrum Disorder alongside 35 age-matched healthy controls, ranging from 3 to 14 years of age. To investigate the possible link between genes and Autism Spectrum Disorder, genetic analysis was conducted using CGH microarray, while a trace element study was carried out using atomic absorption spectrophotometry. After that, the docking of proteins (CNTNAP3 and SPATA31A1) with metals was performed. The results of the trace element analysis show significant variations in zinc and copper levels (P-value is 0.0003 and P-value <0.0001, respectively), with the ASD group having lower zinc and higher copper levels than the controls, changing the zinc/copper ratio. The result of the CGH microarray identified CNVs in the CNTNAP3 and SPATA31A1 genes. The result of docking demonstrated that zinc bound to CNTNAP3 and SPATA31A1 protein more strongly, with binding scores of 6.280 and 9.805, respectively, than copper.
Keywords
Autism Spectrum Disorder (ASD); Copper; Copy Number Variations (CNV); Trace elements; Zinc
| Copy the following to cite this article: Khanam N, Singh A, Srivastava M, Yadav A. K, Vishwakarma S, Maurya A. K, Singh S, Chaurasiya S, Singh S, Jaiswal V, Yadav K, Dubey P, Singh N. K, Ashish A, Singh R. Investigating the Role of Trace Elements and Copy Number Variants in Autism Spectrum Disorder: A Focus on CNTNAP3 and SPATA31A1 genes. Biomed Pharmacol J 2026;19(2). |
| Copy the following to cite this URL: Khanam N, Singh A, Srivastava M, Yadav A. K, Vishwakarma S, Maurya A. K, Singh S, Chaurasiya S, Singh S, Jaiswal V, Yadav K, Dubey P, Singh N. K, Ashish A, Singh R. Investigating the Role of Trace Elements and Copy Number Variants in Autism Spectrum Disorder: A Focus on CNTNAP3 and SPATA31A1 genes. Biomed Pharmacol J 2026;19(2). Available from: https://bit.ly/4cLjBQ5 |
Introduction
Autism spectrum disorder (ASD) is a complex neuro-developmental condition, several difficulties, such as difficulties in communication, social interaction, and the presence of restricted and repetitive behaviors or interests can characterize it.1,2 Autism spectrum disorder can have a relatively common co-occurrence with Intellectual Disability.3,4 About one-third of children with ASD also have intellectual disability.5 Different populations and countries have different prevalence rates of autism spectrum disorder; about 0.76% of children worldwide may have autism spectrum disorder .6 However, the prevalence rate can be altered due to diagnostic criteria assessment methods and population characterization. The etiology of ASD is thought to be influenced by genetic causes and environmental influences, several causes collectively shaping the manifestation of the disorder.7 Within this complex framework, the significance of minerals and trace elements emerges as they assume fundamental roles in the operations of the central nervous system. Changes in their equilibrium, such as deficiencies or surpluses, have been assumed as feasible factors that might cause neurological disorders, such as ASD.8
Several metals play an essential role in maintaining the proper homeostatic conditions of the brain and body.9,10 Magnesium, a chelating agent, is essential for ≥300 biochemical reactions in the body.11,12 Magnesium has several roles in the body, such as maintaining the normal functions of nerve and muscle, supporting the maintenance of a normal and healthy immune system, ensuring a steady heartbeat, and preserving bone strength.13,14 Additionally, it contributes to the regulation of blood glucose levels and assists in the production of energy and protein.15 Selenium is a critical element in numerous enzymes and proteins, known as selenoproteins, which are integral to DNA synthesis and the protection against cellular damage and infections.16,17 Both selenium and selenoproteins are involved in various biological processes, with particular significance in brain development and function.18-20 Chromium (Cr) exists in two oxidation states: +3 and +6. Among these, only chromium (+3) is an essential trace element for the body, and it has a crucial role in regulating glucose and cholesterol metabolism.21,22 Most research on chromium levels in hair indicates that children with autism spectrum disorder (ASD) tend to have altered levels of this element compared to control groups.23 Zinc, an indispensable micronutrient, is naturally present in specific alimentary sources and is additionally obtainable in the form of dietary supplements.24,25 It serves a pivotal function in multiple dimensions of cellular metabolism, being requisite for the catalytic action of various enzymatic processes.26,27 Additionally, zinc contributes to increasing immunological function, synthesis of protein and DNA, wound healing, and cell signaling and division. Several studies have found that low zinc levels are associated with neurodevelopmental diseases.28,29 Copper is a mineral present throughout the body, playing a vital role in various physiological functions.30 It aids in the production of red blood cells and supports the health of nerve cells and the immune system.31 Additionally, copper is involved in the formation of collagen, a crucial component of bones and connective tissues. It may also function as an antioxidant, reducing free radicals that can cause cellular and DNA damage.32,33 Copper is essential for the absorption of iron and is necessary for energy production in the body.34
In some studies, it is mentioned that there is a potential link between zinc deficiency and autism spectrum disorder in newborns and younger children.35 The study also indicated that infants (newly born children) with ASD have various degrees of zinc deficiency in the lower hair scalp.36 They analyzed trace elements in hair scalp, including zinc, and found zinc deficiency potentially associated with autism.37 According to a specific study, it has been demonstrated that there exists a notable reduction in Zn levels (P<0.001) among children diagnosed with autism, as opposed to the corresponding control cohort.38 Moreover, the study has also established a considerable elevation in the Zn/Cu ratio (P<0.001) within the population of autistic children.39,40
Among genetic causes, both cytogenetic and molecular genetic causes are involved. Several genes and proteins are involved in the proper functioning of the brain, neurons, and nerve conduction. Any alterations in these genes and proteins can cause neurodevelopmental disorders.41,42 These changes in genes can be point mutations, Single nucleotide polymorphisms (SNPs), and copy number variations (CNVs).43,44 CNV can be defined as any difference in the copy number of a gene (part of DNA) among individuals.45,46 Copy number variations in particular genes can involve particular symptoms, and sometimes, they can cause a specific disease.47 Copy number variations can be detected by CGH (Comparative Genomic Hybridization) microarray.48,49 Copy number variations of the CNTNAP (Contactin-associated protein-like) gene family play an essential role in ASD.50
In this study, we have performed an analysis of trace elements (Magnesium (Mg), Selenium (Se), Chromium (Ch), Zinc (Zn), and Copper (Cu)) in the serum samples of children with autism spectrum disorder and healthy control children of the same age group for a case-control study. Besides the trace element analysis, a microarray for copy number variation has been done to know the genes involved. The genetic heterogeneity of ASD is reflected in its clinical variability, with individuals exhibiting a wide range of symptoms and severity levels. This heterogeneity underscores the complexity of ASD genetics and highlights the importance of personalized approaches to diagnosis and treatment. Alongside uncommon genetic variations with significant impact sizes, prevalent genetic variants with minor effect sizes also have a role in ASD risk. Docking of proteins (whose genes are showing copy number variation) with trace elements Zn++, Cu+, and Cu++ have been performed by using the MIB2 (Metal Ion-Binding site prediction and modeling server) tool to know the interaction of these proteins with the particular trace element to determine if these copy number variations in genes alter the level of trace in the sample body.
Materials and Methods
Study populations
This study was done at the Department of Anatomy, Institute of Medical Sciences, Banaras Hindu University. Ethical approval has been taken from the institutional ethical committee. A total of 35 cases of autism spectrum disorder were enrolled in this study after proper use of the Indian Scale for Assessment of Autism (ISAA),51 and following the norm of the Diagnostic and Statistical Manual of Mental Disorders, Fifth Edition (DSM-5).52 This study includes children who have an ISAA score above 70; children having an ISAA score less than 70 are excluded. The age limit is between 3-14 years, which is the developmental age; cases with 18 years and above are excluded. The control group consists of 35 healthy asymptomatic children of the same age group. Informed consent from the parents of each child has been taken before sample collection.
Sample collection
Blood samples were collected from autistic children in clot activator vials as well as in EDTA vials for DNA extraction by venipuncture to facilitate serum separation and DNA Extraction respectively, from the Outpatient Department (OPD) under the supervision of the co-supervisors at the Department of Psychiatry and Genetic Clinic of Department of Pediatrics, Institute of Medical Sciences (IMS), Banaras Hindu University (BHU). Parental consent has been obtained as the research population (autistic children) is incapable of providing consent. Ethical clearance has been taken from the ethical committee’s Institute of Medical Science Banaras Hindu University (IMS BHU) ethical letter number (Dean/2022/EC/3825)
Trace element analysis
Serum preparation
The serum was separated from all 35 cases and 35 control blood samples, which were taken in a clot activator vial. After the separation of the serum, the serum was precisely preserved at a temperature of -80ºC to ensure its preservation for subsequent utilization. In preparation for experimentation, the serum samples were methodically transitioned to a temperature of -20ºC, a necessary step to be completed two days before the experimental phase. Subsequently, the evening before the experiment, the serum samples were gently transitioned to a temperature of 4ºC.
Preparation of Diluent
The method involves preparing the diluent solution by combining precise amounts of 1% Nitric acid and 0.4% Triton X with distilled water. In a 50 mL Falcon tube, accurately measure 500 µL of 1% Nitric acid and 200 µL of 0.4% Triton X. Add 49.3 mL of distilled water to the Falcon tube containing the Nitric acid and Triton X. Thoroughly mix the contents to ensure uniformity. The resulting solution constitutes the prepared diluent.
Preparation of Standard
The standard preparation procedure involves using two different volumes to achieve specific concentrations. For a 50 mL standard solution with a concentration of 1 ppm, dissolve 50 µL of the stock solution in 49.95 mL of 1% Nitric acid. Conversely, for a 10 mL standard solution with a concentration of 20 ppb, dissolve 0.2 mL of the stock solution in 9.8 mL of 1% Nitric acid.
Preparation of Blank
To prepare a blank, combine 300 µL of 1% Nitric acid with 800 µL of the diluent in a sampler tube. These preparations are crucial for ensuring precise calibration and accurate baseline measurements, which are essential for AAS analysis.
Analysis of trace elements in the sample serum
Serum levels of trace elements (Mg, Se, Cr, Zn, Cu) were quantified using the Perkin Elmer Atomic Absorption Spectrophotometer, Pinnacle 900T Graphite Furnace method,53 in the Department of Anatomy, Institute of Medical Sciences, Banaras Hindu University. The Atomic Absorption Spectroscopy (AAS) technique involves the absorption of electromagnetic radiation by gasified atoms of an element. This occurs as the temperature rises, causing the element to gasify. The method relies on calculating the absorbed beam by the Lambert-Beer law. According to this law, atoms or substances absorb specific wavelengths from the transmitted beam, causing them to transition from a ground state to an excited state. After absorption, the atom stays in the excited state for a short duration, typically between 10^-6 and 10^-9 seconds, before returning to its original state and releasing the absorbed beam. The absorption wavelengths are distinct for each element, enabling their detection and quantification in a sample using AAS analysis. The evaluation of trace elements, notably magnesium, selenium, Chromium, zinc, and copper, was conducted using the analytical technique of Atomic Absorption Spectroscopy (AAS). This analytical process enabled the acquisition of elemental data. The obtained results were then meticulously compared against established normal ranges to facilitate further comprehensive analysis.
Statistical analysis
Statistical analysis was performed with GraphPad Prism 9 software.54 Normality and lognormality tests (Shapiro-Wilk normality test and Kolmogorov-Smirnov normality test) have been performed to check the normal distribution of data. After the normality check, we performed the Mann–Whitney test. Graphs were created with GraphPad Prism 9 software,54 and compiled using CorelDRAW.55 Statistical significance was set at P < 0.05 (*).
CGH microarray
DNA Extraction
DNA was extracted from blood using the QIAamp® DNA Blood Mini kit (Qiagen, Hilden, Germany), for this purpose, we followed the protocol provided by the manufacturer. The concentration and purity of the genomic DNA were measured using a spectrophotometer, following the manufacturer’s guidelines.
Microarray
DNA copy number detection and genome-wide SNP genotyping were performed on samples using the G3 Human Gene Expression 8x60K v3 Microarray Kit. The procedures for DNA digestion, ligation, PCR amplification, fragmentation, labeling, and hybridization were carried out at the MRU lab according to the protocol provided by Agilent. Data interpretation of microarray. CNVs detected through chromosomal microarrays were analyzed by using the CNVxplorer tool. We identified specific amplifications and deletions in certain genes.
Docking
Binding site prediction and docking of CNTNAP3 (Contactin-associated protein-like 3) with Zn++, Cu+, and Cu++, as well as docking of SPATA31A1 (spermatogenesis-associated protein 31 subfamily A member 1) with Zn++, Cu+, and Cu++, have been done. The binding site and evaluation of the binding ability of Zn++, Cu+, and Cu++ ions with CNTNAP3 protein (uniport ID Q9BZ76) as well as with SPATA31A1 Protein (uniport ID Q5TZJ5) have been done by using the MIB2 server.56 The MIB2 server primarily forecasts binding sites and facilitates the docking of metal ions by utilizing both protein sequences and structures. The 3D configurations of CNTNAP3 with Zn++, Cu+, and Cu++ ions, along with SPATA31A with Zn++, Cu+, and Cu++ ions, were submitted to the MIB2() service. The optimal docking site for CNTNAP3 with Zn++ and Cu++ was selected based on the highest binding score, as was SPATA31A1 with Zn++, Cu+, and Cu++. The resulting docked structure was subsequently visualized and examined using the PyMoL Visualizer.57 The MIB2 server primarily forecasts binding sites and docks metal ions by utilizing both protein sequences and structures.
Results
Result of trace elements
Quantities of various trace elements were compared and measured between the case and control groups, are illustrated in Figure 1. Statistical analysis (Mann-Whitney test) was conducted to evaluate the significance of the differences between the groups.
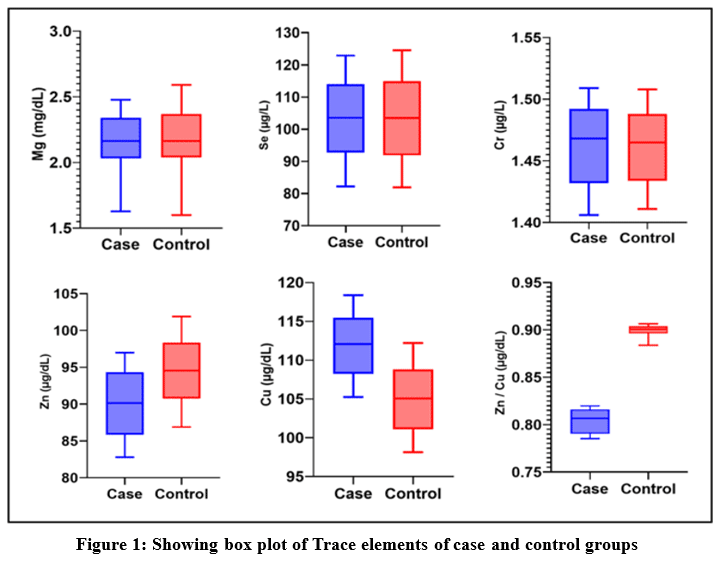 |
Figure 1: Showing box plot of Trace elements of case and control groups |
Table 1 provides details of the Mann-Whitney test observation. The values of Magnesium among the case and control have a median of 2.165 mg/dL and 2.163 mg/dL, respectively, indicating approximately the same value in both case and control. There is no statistically significant difference with a P-value of 0.9326. Selenium also has the same median value among case and control; both have a 103.6µg/L median value with a P-value of 0.9071. The level of Chromium is also approximately the same value with a median 1.468 µg/L and 1.465 µg/L in cases and control, respectively, with a P-value of 0.9326. The level of Zinc in the serum sample of cases is significantly low in comparison to controls. Showing significant differences between case and control, the median values were 90.13µg/dL and 94.56µg/dL, respectively. P – value is 0.0003**. The level of Copper is significantly higher among cases compared to the control, with medians of 112.1µg/dL and 105.1 µg/dL, respectively. There is a statistically significant difference with a P-value <0.0001**. Zinc levels are significantly decreased in cases in comparison to controls, whereas copper levels are significantly higher in cases compared to controls. So, the Zinc / Copper ratio is also altered. The zinc/copper ratio is significantly low among cases in comparison to the control with a median of 0.8065µg/dL and 0.9006 µg/dL in cases and control, respectively. This difference is significant with a p-value of <0.0001**.
Table 1: Showing the result of statical analysis by using the Mann-Whitney test.
| Trace element | Case Median (n=35) | Control Median (n = 35) | IQR | U Value | P Value | |
| Case | Control | |||||
| Magnesium
(mg/dL) |
2.165 | 2.163 | 2.031 – 2.342 | 2.039 – 2.368 | 605 | 0.9326 |
| Selenium
(µg/L) |
103.6 | 103.6 | 92.78 – 113.98 | 91.93 – 114.89 | 602 | 0.9071 |
| Chromium
(µg/L) |
1.468 | 1.465 | 1.432 – 1.492 | 1.434 – 1.488 | 605 | 0.9326 |
| Zinc
(µg/dL) |
90.13 | 94.56 | 85.86 – 94.32 | 90.78 – 98.36 | 309 | 0.0003 |
| Copper
(µg/dL) |
112.1 | 105.1 | 108.24 – 115.44 | 101.11 – 108.78 | 162.5 | <0.0001 |
| Zinc / Copper
(µg/dL) |
0.8065 | 0.9006 | 0.7902 – 0.816 | 0.8961 – 0.9039 | 0 | <0.0001 |
Note- IQR is interquartile range, U is Mann-Whitney U value, and P is the probability value
Result of CGH Microarray
In our CNV analysis by CGH microarray, we observed several CNV on different chromosomes, among which the CNTNAP3 gene family and SPATA31A1 gene family CNV is common in all samples. We analyzed all CNV on CNVxplorer and found the CNTNAP3 gene family and SPATA gene family likely pathogenic in autism. The result of the CGH microarray CNV analysis is shown in Table 2.
Table 2: Showing the result of CGH microarray CNV analysis
| Sn. | Chromosome number | Start – stop bp | Cytoband | Size | Probe | Amplification/
Gain/ Loss/ Deletion |
Gene names | CNVxplore analysis |
| Case -1 | Chr-9 | 38355321-46084155 | p13.2-p11.2 | 7728.835 | 72 | Gain | ALDH1B1, IGFBPL1, FAM96C, ANKRD18A, FAM201A, CNTNAP3, SPATA31A1, FAM74A1, ZNF658B, LOC105379450 | Likely pathogenic
CNTNAP3, SPATA31A1 |
| Chr-9 | 67363066-71627946 | q13- q21.11 | 4264.881 | 44 | Gain | LOC102723709, FAM27E3, FAM27B, ANKRD20A1, ANKRD20A3, MIR4477A, MIR4477B, FRG1JP, FRG1HP, MIR1299 | Not pathogenic for autism | |
| Chr-14 | 106405703-106513022 | Q32.33 | 107.32 | 3 | Gain | ADAM6 | Not pathogenic for autism | |
| Case -2 | Chr-9 | 38355321-46084155 | p13.2-p11.2 | 7728.835 | 12 | Gain | ALDH1B1, IGFBPL1, FAM96C, ANKRD18A, FAM201A, CNTNAP3, SPATA31A1,
FAM74A1, ZNF658B, LOC105379450 |
Likely pathogenic
CNTNAP3, SPATA31A1 |
| Chr-19 | 537965-15521582 | p13.3-p13.12 | 14983.618 | 557 | Gain | CDC34,
G2MM, BSG, HCN2, POLRMT, FGF22, RNF126, FSTL3, PRSS57, |
Not pathogenic for autism | |
| Case -3 | Chr-9 | 43505843-43841603 | p12-p11.2 | 335.761 | 5 | Gain | Fam74A7, SPATA31A6, CNTNAP3B, CNTNAP3P2 | Likely pathogenic
SPATA31A6 CNTNAP3B CNTNAP3P2 |
| Chr-14 | 106405703-106513022 | Q32.33- | 107.32 | 3 | Gain | ADAM6 | Not pathogenic for autism | |
| Case -4 | Chr-6 | 32605385-32631881 | P21.32 | 26.497 | 9 | Deletion | HLA-DQA1,
HLA-DQB1, HLA-DQB1-AS1 |
Not pathogenic for autism |
| Chr-9 | 43505843-43841603 | p12-p11.2 | 335.761 | 5 | Gain | Fam74A7, SPATA31A6, CNTNAP3B, CNTNAP3P2 | Likely pathogenic
SPATA31A6 CNTNAP3B CNTNAP3P2 |
|
| Chr-14 | 106405703-106513022 | Q32.33- | 107.32 | 3 | Gain | ADAM6 | Not pathogenic for autism |
Result of Docking
Table 3. Showing the binding site and binding score of proteins with particular metals.
Table 3: Showing the binding residue and binding score of CNTNAP3 and SPATA31A1 with Zn++, Cu+, and Cu++
| Sn. | Protein | Metal | Binding residue | Binding score |
| 1. | CNTNAP3 | Zn++ | 593E, 596K, 597H | 6.280 |
| Cu+ | 223H, 231H | 4.951 | ||
| Cu++ | 223H, 231H | 5.400 | ||
| 2. | SPATA31A1 | Zn++ | 239H, 241D | 9.805 |
| Cu+ | 238S, 239H, 240C | 7.430 | ||
| Cu++ | 1334S, 1386H | 9.600 |
As previously described in the method part, the docking of Zn++, Cu+, and Cu++ ions to CNTNAP3 protein (UniProt ID Q9BZ76) as well as SPATA31A1 protein (UniProt ID – Q5TZJ5) has been done. In our observation we find zinc has more binding affinity with both protein CNTNAP3 as well as SPATA31A1 than copper. Zn++ binds with CNTNAP3 protein at 593E, 596K, and 597H residues, having a binding score of 6.280. The protein structure and binding site of CNTNAP3 are shown in Figure 2. The binding score and residue of Zn++ with SPATA31A1 protein are 9.805 and 239H, 241D, respectively. The protein structure and binding site of SPATA31A1 are shown in Figure 3. Both ions of copper, Cu+ and Cu++, have less binding score in comparison to Zinc.
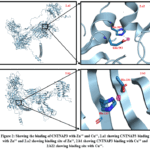 |
Figure 2: Showing the binding of CNTNAP3 with Zn++ and Cu++, 2.a1 showing CNTNAP3 binding with Zn++ and 2.a2 showing binding site of Zn++, 2.b1 showing CNTNAP3 binding with Cu++ and 2.b22 showing binding site with Cu++. |
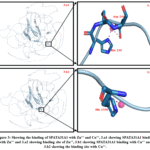 |
Figure 3: Showing the binding of SPATA31A1 with Zn++ and Cu++, 3.a1 showing SPATA31A1 binding with Zn++ and 3.a2 showing binding site of Zn++, 3.b1 showing SPATA31A1 binding with Cu++ and 3.b2 showing the binding site with Cu++. |
Discussion
This study was conducted to compare the levels of various trace elements between a case group and a control group to identify potential differences that might contribute to the pathophysiology of autism spectrum disorder. In our trace element analysis, we observed a significant decrease in zinc and an increase in copper levels in the case group compared to the control group. In contrast, the levels of magnesium, selenium, and chromium showed no significant difference between the two groups. The zinc / Copper ratio is also significantly lower in cases than in the control. Zinc plays a crucial role in our body and nervous system.58,59 A deficiency of zinc can lead to the development of autism spectrum disorder because zinc plays a role in neurogenesis 60,61, and low zinc decreases neurogenesis, as shown in Figure 4. Copper is also related to normal neuronal development; both deficiency and excess concentrations of copper can alter normal neural development.62,63 Copper is also associated with nerve conduction.64 High copper can have a negative effect on nerve conduction and can lead to damage to the myelin sheath.65 Copper toxicity is more common in Wilson’s disease, which is one of the causes behind autistic features.66 The effect of high copper concentration is shown in Figure 5. In this study, we found lower serum zinc levels in cases (90.13 µg/dL) than in control (94.56µg/dL) with a P-value of 0.0003. We also observed higher copper in the serum of cases (112.1µg/dL) than control (105.1 µg/dL) and P-value <0.0001. A low zinc/copper ratio among cases in comparison to control (0.8065 µg/dL among cases and 0.9006 µg/dL among control) is also noticed; this difference is also significant with a P-value <0.0001. Low zinc and high copper are a biomarker of autism, low zinc/copper ratio is also a cause of autism spectrum disorder.67 Besides zinc and copper, we also performed a trace element analysis of magnesium, selenium, and chromium, but we found significant differences only in zinc and copper.
To evaluate CNV, we also performed CGH microarray. In our study of CGH microarray for CNV analysis, it is observed that copy number variation has occurred in some genes and cytoband which is mentioned in table1. Copy number variations on the p arm of chromosome number 9 with cytoband p13.2-p11.2 and p12-p11.2 are common in all samples. This CNV has a common gene CNTNAP3 gene family and SPATA31A1 gene family. CNTNAP3 is a neuro-connexin protein.68 Connexin proteins are transmembrane proteins and play a role in cell divisions, cell recognitions, and neurogenesis and found gap junctions, neuro-connexin is involved in cell-cell recognition in the nervous system.69 CNTNAP3 interacts with synaptic adhesion protein.70 We have analyzed the result with the CNVxplorer tool we found that CNTNAP3 and SPATA31A1 have been likely pathogenic. The association of the CNTNAP3 gene with the autism gene family has already been reported in the literature.71-73
So, we have performed the docking of CNTNAP3 and SPATA31A1 proteins (product of CNTNAP3 and SPATA31A1 gene) with Zn++, Cu+, and Cu++. Shown in Table 3. Both of these proteins have binding affinity with Zn++, Cu+, and Cu++ but zinc shows a higher binding score than copper. The binding score of CNTNAP3 protein with ZN++ is 6.280, and the binding scores of Cu+ and Cu++ are 4.951 and 5.400, respectively. So, we can say the CNTNAP3 protein binds with zinc with more affinity than with copper. On the other hand, SPATA31A1 protein also shows a high binding affinity with Zn++, having a binding score of 9.805, a binding score of Cu+, and Cu++ having a binding score of 7.430 and 9.600, respectively. This protein also shows a higher binding score with zinc than with copper. Thus, both of these proteins bind more frequently with zinc than copper, and so the level of zinc in serum became low in comparison to the control.
The protein products of both genes CNTNAP3 and SPATA31A1 have a high binding affinity for Zn++. According to our results, both of these genes have an increased copy number (there is a duplication in these genes). So, the amount of protein products produced by these genes will be elevated. As mentioned earlier in the result, these proteins have a strong binding score for zinc. Figure 6 shows that the increase in the copy number can lead to high sequestration of Zn++ from the blood. This can result in a lower amount of Zn++ in the serum, leading to a zinc deficiency. Zn++ has many roles in the body, such as immune function, neurotransmission, and cellular growth and repair. Therefore, the copy number variation may lead to zinc imbalances, which could cause the symptoms of autism spectrum disorder (ASD) in children.
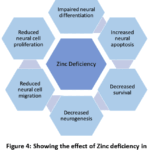 |
Figure 4: Showing the effect of Zinc deficiency in neurodevelopment and neurogenesis |
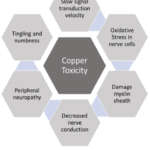 |
Figure 5: Showing the effect of copper toxicity in nerve conduction |
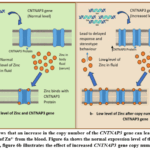 |
Figure 6: Shows that an increase in the copy number of the CNTNAP3 gene can lead to elevated sequestration of Zn²⁺ from the blood. Figure 6a shows the normal expression level of the CNTNAP3 gene, figure 6b illustrates the effect of increased CNTNAP3 gene copy number. |
Implications
This study holds important implications for understanding autism spectrum disorder (ASD), particularly about how copy number variations (CNVs) in the genes CNTNAP3 and SPATA31A1 contribute to the disorder’s underlying pathophysiology. These CNVs are linked to zinc (Zn++) deficiency in children with autism, which is a crucial aspect of the study. Even inside the same family where children share the same food and water sources, there may be significant differences in their trace elements levels. This variation can be due to the sequestration of trace elements by proteins (Zn++ is sequestrated by CNTNAP3 and SPATA31A1), which may hinder the availability of this essential trace element. The levels of Zinc, Copper, and other trace elements in the body have several roles in the body. Many physiological processes of the body, such as nerve conduction, immune response, and several metabolic processes. So, understanding the impact of CNVs on trace element levels is crucial to know how such genetic factors could influence the development and severity of autism spectrum disorder in affected children.
Limitations and future research
This research has contributed valuable understandings into the autism spectrum disorder. However, has many limitations that could hamper the results. A significant limitation is the relatively small sample size (N = 35), which affects the statistics of the study and the detection of delicate effects or the drawing of definitive conclusions. Furthermore, the study’s hospital-based based may introduce certain changes in disease ratio. Populations within hospitals may display higher occurrences of specific diseases or conditions compared to the general population due to healthcare-seeking behaviors and the prevalence of more severe or complex cases. To avoid these limitations, future research should consider using larger sample sizes to improve statistical validity. A population-based study would yield a more characteristic overview of the targeted demographic, provide more accurate assessments of the autistic children under investigation. Such practices may enhance the validity of the findings and provide a more comprehensive understanding of the phenomena being examined.
Conclusion
The concentrations of zinc and copper significantly impact human physiology and are associated with neurogenesis and nerve conduction. In children, low zinc levels, elevated copper levels, and a reduced zinc-to-copper ratio in the bloodstream can manifest as autistic symptoms. During developmental stages, insufficient zinc and excessive copper levels can result in delayed responses and stereotypic behaviors in children. Adequate zinc levels are crucial during developmental periods for proper neurogenesis. The zinc-to-copper ratio is particularly important during these developmental stages.
Acknowledgment
This study was conducted at the Multidisciplinary Research Unit, an ICMR-DHR-funded laboratory located at the Department of Anatomy, Institute of Medical Sciences (IMS), Banaras Hindu University (BHU). The first author of this work is a recipient of a Senior Research Fellowship from the Indian Council of Medical Research, contributing significantly to the study.
Funding sources
The first author is qualified for the ICMR-JRF 2020 in life sciences and received an ICMR JRF for 2 years, followed by an SRF. file no. 3/1/3/JRF-2020/HRD(LS)135018/39.
Conflict of interest
The author(s) do not have any conflict of interest
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This study was done at the Department of Anatomy, Institute of Medical Sciences, Banaras Hindu University. Ethical approval has been taken from the institutional ethical committee
Informed Consent Statement
Informed consent from the parents of each child has been taken prior to sample collection.
Clinical Trial Registration
This research does not involve any clinical trials
Permission to reproduce material from other sources.
Not Applicable
Author contributions
- Nargis Khanam: Conceptualizations, Methodology, Writing – Original Draft.
- Ankur Singh: Supervision, Review & Editing,
- Mona Srivastava: Supervision: Review & Editing .
- Abhay Yadav: Methodology,
- Shani Vishwakarma: Methodology,
- Anil Kumar Maurya: Analysis,
- Suchitra Singh: Docking,
- Shraddha Chaurasiya: Visualization & figure preparations,
- Surbhi Singh: Docking,
- Khushbu Yadav: sample collection,
- Vaibhav Jaiswal: sample collection,
- Pooja Dubey: Review
- Nitish Kumar Singh: Review
- Ashish: data analysis,
- Royana Singh: Resources, Supervision
References
- DeNucci BL, de Lima LS, Mota IF, et al. Current knowledge, challenges, new perspectives of the study, and treatments of autism spectrum disorder. Reprod Toxicol. 2021;106:82-93.
CrossRef - Kurzius-Spencer M, Pettygrove S, Christensen D, et al. Behavioral problems in children with autism spectrum disorder with and without co-occurring intellectual disability. Res Autism Spectr Disord. 2018;56:61-71.
CrossRef - Bertelli MO. ASD and intellectual disability. In: Bertelli MO, Deb S, eds. Psychopathology in Adolescents and Adults With Autism Spectrum Disorders. Cham, Switzerland: Springer International Publishing; 2019:111-130.
CrossRef - Schieve LA, Clayton HB, Durkin MS, Wingate MS, Drews-Botsch C. Comparison of perinatal risk factors associated with autism spectrum disorder (ASD), intellectual disability (ID), and co-occurring ASD and ID. J Autism Dev Disord. 2015;45(8):2361-2372.
CrossRef - Baio J. Prevalence of autism spectrum disorder among children aged 8 years—Autism and Developmental Disabilities Monitoring Network, 11 sites, United States, 2014. MMWR Surveill Summ. 2018;67(6):1-23.
CrossRef - Issac A, Halemani K, Shetty A, et al. The global prevalence of autism spectrum disorder in children: a systematic review and meta-analysis. Osong Public Health Res Perspect. 2025;16(1):3-15.
CrossRef - Sauer AK, Stanton J, Hans S, Grabrucker A. Autism spectrum disorders: etiology and pathology. In: Autism Spectrum Disorders. Brisbane (AU): Exon Publications; 2021:1-5.
CrossRef - Jomova K, Makova M, Alomar SY, et al. Essential metals in health and disease. Chem Biol Interact. 2022;367:110173.
CrossRef - Jomova K, Vondrakova D, Lawson M, Valko M. Metals, oxidative stress and neurodegenerative disorders. Mol Cell Biochem. 2010;345(1):91-104.
CrossRef - Jan AT, Azam M, Siddiqui K, et al. Heavy metals and human health: mechanistic insight into toxicity and counter defense system of antioxidants. Int J Mol Sci. 2015;16(12):29592-29630.
CrossRef - Fiorentini D, Cappadone C, Farruggia G, Prata C. Magnesium: biochemistry, nutrition, detection, and social impact of diseases linked to its deficiency. Nutrients. 2021;13(4):1136.
CrossRef - Smith SW. The role of chelation in the treatment of other metal poisonings. J Med Toxicol. 2013;9(4):355-369.
CrossRef - Fatima G, Dzupina A, Alhmadi HB, et al. Magnesium matters: a comprehensive review of its vital role in health and diseases. Cureus. 2024;16(10):e72917.
CrossRef - Mathew AA, Panonnummal R. “Magnesium”—the master cation—as a drug—possibilities and evidences. Biometals. 2021;34(5):955-986.
CrossRef - Stanojević M, Djuricic N, Parezanovic M, et al. The impact of chronic magnesium deficiency on excitable tissues—translational aspects. Biol Trace Elem Res. 2025;203(2):707-728.
CrossRef - Hariharan S, Dharmaraj S. Selenium and selenoproteins: its role in regulation of inflammation. Inflammopharmacology. 2020;28(3):667-695.
CrossRef - Zoidis E, Seremelis I, Kontopoulos N, Danezis GP. Selenium-dependent antioxidant enzymes: actions and properties of selenoproteins. Antioxidants (Basel). 2018;7(5):66.
CrossRef - Zhang F, Li X, Wei Y. Selenium and selenoproteins in health. Biomolecules. 2023;13(5):799.
CrossRef - Solovyev ND. Importance of selenium and selenoprotein for brain function: from antioxidant protection to neuronal signalling. J Inorg Biochem. 2015;153:1-2.
CrossRef - Pillai R, Uyehara-Lock JH, Bellinger FP. Selenium and selenoprotein function in brain disorders. IUBMB Life. 2014;66(4):229-239.
CrossRef - Genchi G, Lauria G, Catalano A, Carocci A, Sinicropi MS. The double face of metals: the intriguing case of chromium. Appl Sci. 2021;11(2):638.
CrossRef - Morvaridzadeh M, Estevao MD, Qorbani M, et al. The effect of chromium intake on oxidative stress parameters: a systematic review and meta-analysis. J Trace Elem Med Biol. 2022;69:126879.
CrossRef - Skalny AV, Simashkova NV, Klyushnik TP, et al. Hair toxic and essential trace elements in children with autism spectrum disorder. Metab Brain Dis. 2017;32(1):195-202.
CrossRef - Saghazadeh A, Ahangari N, Hendi K, Saleh F, Rezaei N. Status of essential elements in autism spectrum disorder: systematic review and meta-analysis. Rev Neurosci. 2017;28(7):783-809.
CrossRef - Rashaid AH, Nusair SD, Alqhazo MT, et al. Heavy metals and trace elements in scalp hair samples of children with severe autism spectrum disorder: a case-control study on Jordanian children. J Trace Elem Med Biol. 2021;67:126790.
CrossRef - Awuchi CG, Igwe VS, Amagwula IO, Echeta CK. Health benefits of micronutrients (vitamins and minerals) and their associated deficiency diseases: a systematic review. Int J Food Sci. 2020;3(1):1-32.
CrossRef - Silva CS, Moutinho C, Ferreira da Vinha A, Matos C. Trace minerals in human health: iron, zinc, copper, manganese and fluorine. Int J Sci Res Methodol. 2019;13(3):57-80.
- Li Z, Liu Y, Wei R, Yong VW, Xue M. The important role of zinc in neurological diseases. Biomolecules. 2022;13(1):28.
CrossRef - Bonaventura P, Benedetti G, Albarède F, Miossec P. Zinc and its role in immunity and inflammation. Autoimmun Rev. 2015;14(4):277-285.
CrossRef - Collins JF. Copper: basic physiological and nutritional aspects. In: Collins JF, ed. Molecular, Genetic, and Nutritional Aspects of Major and Trace Minerals. San Diego, CA: Academic Press; 2017:69-83.
CrossRef - Prohaska JR, Lukasewycz OA. Effects of copper deficiency on the immune system. In: Gershwin ME, German JB, Keen CL, eds. Antioxidant Nutrients and Immune Functions. Boston, MA: Springer; 1990:123-143.
CrossRef - Wang Y, Zhang W, Yao Q. Copper-based biomaterials for bone and cartilage tissue engineering. J Orthop Transl. 2021;29:60-71.
CrossRef - Linder MC. The relationship of copper to DNA damage and damage prevention in humans. Mutat Res. 2012;733(1-2):83-91.
CrossRef - Nishito Y, Kambe T. Absorption mechanisms of iron, copper, and zinc: an overview. J Nutr Sci Vitaminol (Tokyo). 2018;64(1):1-7.
CrossRef - Yasuda H, Yoshida K, Yasuda Y, Tsutsui T. Infantile zinc deficiency: association with autism spectrum disorders. Sci Rep. 2011;1:129.
CrossRef - Rashaid AH, Nusair SD, Alqhazo MT, et al. Heavy metals and trace elements in scalp hair samples of children with severe autism spectrum disorder: a case-control study on Jordanian children. J Trace Elem Med Biol. 2021;67:126790.
CrossRef - Fiore M, Barone R, Copat C, et al. Metal and essential element levels in hair and association with autism severity. J Trace Elem Med Biol. 2020;57:126409.
CrossRef - Goyal DK, Neil JR, Simmons SD, et al. Zinc deficiency in autism: a controlled study. Insights Biomed. 2019;4:12.
- Li SO, Wang JL, Bjørklund G, Zhao WN, Yin CH. Serum copper and zinc levels in individuals with autism spectrum disorders. Neuroreport. 2014;25(15):1216-1220.
CrossRef - El-Meshad GM, Abou El-Khair MS, Moharam NM, Abd El-Nabi SA. The plasma zinc/serum copper ratio as a biomarker in children with autism spectrum disorders. Menoufia Med J. 2017;30(3):727-733.
- Rylaarsdam L, Guemez-Gamboa A. Genetic causes and modifiers of autism spectrum disorder. Front Cell Neurosci. 2019;13:470832.
CrossRef - Parenti I, Rabaneda LG, Schoen H, Novarino G. Neurodevelopmental disorders: from genetics to functional pathways. Trends Neurosci. 2020;43(8):608-621.
CrossRef - Pös O, Radvanszky J, Buglyó G, et al. DNA copy number variation: main characteristics, evolutionary significance, and pathological aspects. Biomed J. 2021;44(5):548-559.
CrossRef - Fanciulli M, Petretto E, Aitman TJ. Gene copy number variation and common human disease. Clin Genet. 2010;77(3):201-213.
CrossRef - Almal SH, Padh H. Implications of gene copy-number variation in health and diseases. J Hum Genet. 2012;57(1):6-13.
CrossRef - Zarrei M, MacDonald JR, Merico D, Scherer SW. A copy number variation map of the human genome. Nat Rev Genet. 2015;16(3):172-183.
CrossRef - Morrow EM. Genomic copy number variation in disorders of cognitive development. J Am Acad Child Adolesc Psychiatry. 2010;49(11):1091-1104.
CrossRef - Carreira IM, Ferreira SI, Matoso E, et al. Copy number variants prioritization after array-CGH analysis: a cohort of 1000 patients. Mol Cytogenet. 2015;8:103.
CrossRef - Boone PM, Bacino CA, Shaw CA, et al. Detection of clinically relevant exonic copy-number changes by array CGH. Hum Mutat. 2010;31(12):1326-1342.
CrossRef - Murdoch JD, Gupta AR, Sanders SJ, et al. No evidence for association of autism with rare heterozygous point mutations in contactin-associated protein-like 2 (CNTNAP2), or in other contactin-associated proteins or contactins. PLoS Genet. 2015;11(1):e1004852.
CrossRef - Manohar H, Kishore T, Jacob P. Indian Scale for Assessment of Autism (ISAA): issues with the current assessment scale and recommendations for disability assessment. Indian J Psychol Med. 2025;47(1):76-79.
CrossRef - Lolk A. Neurokognitive lidelser. In: Diagnostic and Statistical Manual of Mental Disorders. Washington, DC: American Psychiatric Association; 2013.
- Mokgohloa CP, Thomas MS, Mokgalaka NS, Ambushe AA. Speciation of chromium in river sediments by graphite furnace-atomic absorption spectrometry after microwave-assisted extraction. Int J Environ Anal Chem. 2022;102(18):6454-6468.
CrossRef - GraphPad Software. GraphPad Prism. Version 9. San Diego, CA: GraphPad Software; 2020.https://www.graphpad.com/guides/prism/9/statistics/
- Corel Corporation. CorelDRAW Graphics Suite. Version 2020. Ottawa, ON: Corel Corporation; 2020.Lu CH, Chen CC, Yu CS, et al.
- Lu CH, Chen CC, Yu CS, Liu YY, Liu JJ, Wei ST, Lin YF. MIB2: metal ion–binding site prediction and modeling server. Bioinformatics. 2022;38(18):4428-4429.
CrossRef - Schrödinger, LLC, DeLano W. PyMOL. Published 2020. http://www.pymol.org/pymol.
- Wang B, Fang T, Chen H. Zinc and central nervous system disorders. Nutrients. 2023;15(9):2140.
CrossRef - Benarroch E. What are the functions of zinc in the nervous system? Neurology. 2023;101(16):714-720.
CrossRef - Ross MM, Hernandez-Espinosa DR, Aizenman E. Neurodevelopmental consequences of dietary zinc deficiency: a status report. Biol Trace Elem Res. 2023;201(12):5616-5639.
CrossRef - Alsufiani HM, Alkhanbashi AS, Laswad NA, et al. Zinc deficiency and supplementation in autism spectrum disorder and Phelan-McDermid syndrome. J Neurosci Res. 2022;100(4):970-978.
CrossRef - Manto M. Abnormal copper homeostasis: mechanisms and roles in neurodegeneration. Toxics. 2014;2(2):327-345.
CrossRef - Gromadzka G, Tarnacka B, Flaga A, Adamczyk A. Copper dyshomeostasis in neurodegenerative diseases—therapeutic implications. Int J Mol Sci. 2020;21(23):9259.
CrossRef - D’Ambrosi N, Rossi L. Copper at synapse: release, binding and modulation of neurotransmission. Neurochem Int. 2015;90:36-45.
CrossRef - Gaier ED, Eipper BA, Mains RE. Copper signaling in the mammalian nervous system: synaptic effects. J Neurosci Res. 2013;91(1):2-19.
CrossRef - Lorincz MT. Wilson disease and related copper disorders. In: Handb Clin Neurol. Vol 147. Amsterdam, Netherlands: Elsevier; 2018:279-292.
CrossRef - Scassellati C, Bonvicini C, Benussi L, Ghidoni R, Squitti R. Neurodevelopmental disorders: metallomics studies for the identification of potential biomarkers associated to diagnosis and treatment. J Trace Elem Med Biol. 2020;60:126499.
CrossRef - Tong DL, Chen RG, Lu YL, et al. The critical role of ASD-related gene CNTNAP3 in regulating synaptic development and social behavior in mice. Neurobiol Dis. 2019;130:104486.
CrossRef - Vicario N, Zappalà A, Calabrese G, et al. Connexins in the central nervous system: physiological traits and neuroprotective targets. Front Physiol. 2017;8:1060.
CrossRef - Tong DL, Chen RG, Lu YL, et al. The critical role of ASD-related gene CNTNAP3 in regulating synaptic development and social behavior in mice. Neurobiol Dis. 2019;130:104486.
CrossRef - Poot M. A candidate gene association study further corroborates involvement of contactin genes in autism. Mol Syndromol. 2014;5(5):229-235.
CrossRef - Zou Y, Zhang WF, Liu HY, et al. Structure and function of the contactin-associated protein family in myelinated axons and their relationship with nerve diseases. Neural Regen Res. 2017;12(9):1551-1558.
CrossRef - Bamford RA, Widagdo J, Takamura N, et al. The interaction between contactin and amyloid precursor protein and its role in Alzheimer’s disease. Neuroscience. 2020;424:184-202.
CrossRef
Abbreviations
ASD – Autism Spectrum Disorder
SNP – Single Nucleotide Polymorphism
CNV – Copy Number Variations
CGH – Comparative Genomic Hybridization
CNTNAP – Contactin-associated protein-like
ISAA – Indian Scale for Assessment of Autism
DSM-5 – Diagnostic and Statistical Manual of Mental Disorders, Fifth Edition
OPD – Outpatient Department
IMS – Institute of Medical Sciences
BHU – Banaras Hindu University
AAS – Atomic Absorption Spectroscopy
MIB2 – Metal Ion-Binding site prediction and modeling server
CNTNAP3 – Contactin-associated protein-like 3
SPATA31A1 – spermatogenesis-associated protein 31 subfamily A member 1
UniProt ID – Universal Protein resource identification (UniProt Knowledgebase – UniProtKB)







