Manuscript accepted on :28-10-2025
Published online on: 10-11-2025
Plagiarism Check: Yes
Reviewed by: Dr. Rashid Akhundov
Second Review by: Dr. Abdulrahman Rasheed Mahmood
Final Approval by: Dr. Prabhishek Singh
Sakthi Periyasamy and Priya Deivasigamani*
and Priya Deivasigamani*
Department of Pharmaceutical Chemistry, SRM College of Pharmacy, Faculty of Medicine and Health Sciences, SRM Institute of Science and Technology, Kattankulathur, Chengalpattu, Tamilnadu, India.
Corresponding Author E-mail: priyad@srmist.edu.in
DOI : https://dx.doi.org/10.13005/bpj/3298
Abstract
Pinocembrin, a naturally occurring flavonoid, possesses anti-inflammatory, antioxidant, antimicrobial, and neuroprotective properties. The multi-target potential of this study is assessed through molecular docking, ADMET (Absorption, Distribution, Metabolism, Excretion, and Toxicity) predictions, and Density Functional Theory (DFT) analysis. Ten disease-relevant protein targets were selected based on literature and SuperPred predictions. The docking with AutoDock Vina demonstrated a strong binding to nitric oxide synthase (–10.3 kcal/mol), cytochrome P450 1A2 (–10.0 kcal/mol), and the delta opioid receptor (–9.5 kcal/mol), indicating potential roles in inflammation control, metabolic modulation, and neuroprotection. The docking was performed in blind mode, wherein the grid parameters were configured to encompass the entire receptor surface and ligand binding regions, with the exhaustiveness value set to 8 to ensure adequate sampling of the conformational space. DFT results indicated a stable and moderately reactive nature, aligning with its pharmacological potential. ADMET profiling predicted high intestinal absorption, blood–brain barrier permeability, and compliance with Lipinski’s rule, though possible hERG (Human Ether-à-go-go Related Gene (potassium channel)) inhibition and CYP (Cytochrome P450 enzymes) interactions highlight the need for further safety evaluation. Overall, the findings position pinocembrin as a promising lead compound for the development of therapeutics targeting neurological, inflammatory, and metabolic disorders.
Keywords
ADMET Prediction; Auto Dock Vina; Flavonoid; Molecular Docking; Multi-target Drug; Pinocembrin
Download this article as:| Copy the following to cite this article: Periyasamy S, Deivasigamani P. Computational Investigation of Pinocembrin as a Multi-Target Ligand: A Molecular Docking Study. Biomed Pharmacol J 2025;18(4). |
| Copy the following to cite this URL: Periyasamy S, Deivasigamani P. Computational Investigation of Pinocembrin as a Multi-Target Ligand: A Molecular Docking Study. Biomed Pharmacol J 2025;18(4). Available from: https://bit.ly/3LT9e1u |
Introduction
Pinocembrin (5,7-dihydroxyflavanone) is also found in many other plants, such as in several genera of the Piperaceae family, including five hundred species and fourteen genera considered to be rich in pinocembrin. However, it has been found that the most diverse and international species are found in the two genera of Peperomia and Piper, which contain 600 and 700 species, respectively.1,2,3 In addition, this family of plants has been isolated with Pinocembrin (Fig 1) from plants belonging to the Lauraceae and Asteraceae families; these families are huge. Africa has 600 species of Helichrysum and roughly 244 of these are in South Africa; approximately 250 species of Cryptocarya are essentially found in tropical and subtropical regions.1,4 Additionally, pinocembrin was found in honey and in the aerial parts of Flourensia oolepis S.F. Blake (Asteraceae).5 Flavonoid pinocembrin is a naturally existing component, and it is found in tea, red wine, fruits, vegetables, nuts and seeds, herbs, spices, propolis, honey, and some plants, such as wild marjoram and ginger roots.6,7 Pharmacological character of the Pinocembrin makes it possible to be used to treat various illnesses, include cancer and problems with blood vessels, as well as inflammation by preventing bacterial colonization.8,9 Researchers have seen that pinocembrin can also be biosynthesised. In this project, artificial phenyl propanoid pathway was designed to produce the flavanone pinocembrin from microorganisms. On a single pET plasmid in E. coli, CHS from the licorice plant Glycyrrhiza echinata, PAL from the yeast Rhodotorula rubra, 4CL from the actinomycete S. coelicolor and CHI from the plant Pueraria lobata were combined to achieve this. In pharmacological research, it has shown some interesting qualities which may be applicable in treatment of diseases, e.g. cancer, cardiovascular disorders as well as endotoxin shock. Indeed, as has been shown by previous research, pinocembrin is a neuroprotective and anti-inflammatory compound and reduces reactive oxygen species (ROS), stabilizes the blood- brain barrier and changes mitochondrial activity, and controls apoptosis.10-17
The pharmacological activity of pinocembrin can be further identified throug in silico studies, where a Computer- aided drug design tools can deploy different types of powerful and highly efficient models to afford quantitative explanation of the structure–activity relationship (SAR) observance.18 The previous docking studies indicated that pinocembrin exhibits strong CYP1A2 inhibition and possesses potential drug metabolism and interactions through strong contact with the enzyme’s active site.19 Molecular docking results indicated a strong favorable binding to Plasmodium falciparum DHFR-TS having an affinity greater than the widely prescribed drug chloroquine. Previous research on the antioxidant pinocembrin has demonstrated strong antiplasmodial activity, in agreement with this observation. This result serves as a lead chemical for the development of antimalarial drug.20 The studies of binding with alpha synuclein, a critical target of Parkinson’s disease, have shown that, like curcumin, pinocembrin has a high binding affinity, suggesting a potential function of the neuroprotection.21 Literature proved that pinocembrin has high LibDock scores and stable interactions with the critical ovarian cancer targets such as AKT1, TNF and BCL2 and reinforces the role of this compound in the modulation of apoptosis pathways.22 In extension, (2S)-Pinocembrin also showed a fairly high binding affinity for MAPKs, especially p38α and ERK2, suggesting a possible mechanism of action whereby it inhibits PGE2 production by antagonizing this (and perhaps other) signalling back to base.23 Pinocembrin was also found to show good binding affinity to BACE1 and might therefore be important in the management of Alzheimer’s disease.24 Based on previous research, AutoDock Vina was used in our study to dock pinocembrin with ten different proteins. The Comparative Assessment of Scoring Functions (CASF) is a standard for docking and compares scoring functions.25-28 AdMET (Absorption, Distribution, Metabolism, Excretion and Toxicity) profiling in the form of pharmacokinetic and toxicological properties of pharmacokinetic and toxicological properties of pinocembrin was performed using a common web-based prediction tool – PreADMET. Only a couple of dozen of the comprehensive predictions that PreADMET provides involve human intestinal absorption (HIA), Caco-2 permeability, MDCK cell permeability, blood- brain barrier (BBB) penetration, skin permeability, and carcinogenicity. However, these computational methods provide essential clues to the drug ability and safety profile of – Pinocembrin – supporting it as a potential therapeutic. Potential protein target and PDB structure retrieval for molecular docking studies were accomplished using SuperPred to predict potential protein targets.29 These in silico methods bring forward important information regarding Pinocembrin’s therapeutic efficacy and safety profile. Fig 1 represents the structure of Pinocembrin.
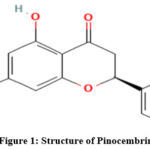 |
Figure 1: Structure of Pinocembrin |
Materials and Methods
Protein Preperation
Thus, molecular docking studies were carried out on ten disease relevant protein targets to evaluate the multi- target pharmacological potential for pinocembrin. These targets were selected from predictions by the SuperPred server as well as supporting data from previously conducted research studies. For inflammation, Cyclooxygenase-2 (COX-2; PDB ID: 6NOD) was selected because it functions as an inhibitor, while acetylcholinesterase (AChE; PDB ID: 6OL9) was used as the target for Alzheimer’s disease. For Parkinson’s disease, Alpha-synuclein (PDB ID: 1XQ8) was chosen as the target. To investigate the anti-photoaging activity, Staphylococcus proteinase (PDB ID: 1G4K) was selected. Included for pain modulation was the delta-opioid receptor (PDB ID: 4N6H). The nitric oxide synthase (NOS; PDB ID: 3E7G) was used, as it plays an important role in inflammatory and neurodegenerative conditions. There is a selection of a relevant target, plasmepsin II (PDB ID: 1J3I), for its antiplasmodial activity. To study inhibition of drug metabolism, the structure of CYP1A2 (PDB ID: 2HI4) was selected, while its relevance to the cardiovascular system is reflected by plasminogen activator inhibitor-1 (PAI-1; PDB ID: 3CVM). The central nervous system modulation was also studied using the GABA-A receptor (PDB ID: 6X3T). The docking simulations to predict the binding affinity and interaction profile of pinocembrin were carried out using AutoDock Vina for the retrieved protein structures from the RCSB Protein Data Bank.30,31
Ligand Preperation
Chem 3D 16.0 32 was used to generate a three-dimensional structure of Pinocembrin, after which it is refined using molecular mechanics force fields to give a low energy conformation to make sure that the ligand starts off energetically favourable.
Molecular Docking
The resulting molecular interactions between pinocembrin and 6NOD, 2HI4, 1J3I, 6OL9, 1XQ8, 1G4K, 4N6H, 6X3T, 3E7G and 3CVM, one of the most widely used programs as a means to predict small molecule binding affinities and conformations to macromolecular targets were examined using computational docking study using AutoDock Vina.33 Finally, an appropriate format of the optimized ligand and macromolecules to use with AutoDock Vina, generally the PDBQT format, including atomic coordinates, atomic charges, and types of atoms, was created. Potential pinocembrin was assumed to bind the active site of each protein based on available literature and structural data; when no such information existed, blind docking was used to explore the entire protein surface.
Geometry Optimization and Dft Calculations
The geometry of Pinocembrin was optimized using Avogadro. Then the optimized Structure was further subjected to DFT Calculations using ORCA Software. The energy minimization, HOMO–LUMO energy
gap, dipole moment, and molecular electrostatic potential were calculated and analyzed.
Drug Likeness and Admet Prediction
PreADMET and SWISS ADME web server was used for computation of drug similarity and pharmacokinetic characteristics of pinocembrin absorption, distribution, metabolism, excretion, and toxicity (ADMET). Human intestine absorption (% HIA), Caco-2 permeability, MDCK cell permeability, skin permeability, blood–brain barrier penetration and carcinogenicity were predicted as different attributes.
Results
Molecular Docking Studies
Molecular docking for Pinocembrin binding to ten specific protein targets involved with different therapeutic pathways was carried out using AutoDock Vina (Table 1). Then, binding energies, critical amino acid interactions, and structure-activity connections for the compound were evaluated to assess its potential multi-target properties. The lowest binding energies for pinocembrin were found for 3E7G (nitric oxide synthase), 4N6H (delta opioid receptor), and 2HI4 (CYP1A2 Inhibition) in molecular docking.
The 2D interaction plots indicate hydrogen bonding amino acid residues, whereas the 3D views demonstrate pinocembrin’s active site occupancy. These interactions suggest pinocembrin can strongly inhibit these targets (Fig. 2– Fig. 4). The 2D and 3D interactions of Pinocembrin with other targets were also represented in Fig. 5 – Fig. 11.
Table 1: Interactions of Pinocembrin Against Ten Different Proteins
|
Activity |
PDB ID |
Binding Energy (kcal/mol) |
Detailed Interactions |
|
Anti-inflammatory |
6NOD |
-7.4 |
Conventional hydrogen bonds: LYS A:596, ALA A:597, TYR A:454; Pi-alkyl interactions: ARG A:586, VAL A:444; |
|
Anti-Alzheimer |
6OL9 |
-7.9 |
Conventional hydrogen bond: TYR A:481; Pi–pi stacking: TRP A:477; |
|
Anti-Parkinson |
1XQ8 |
-5.0 |
Conventional hydrogen bonds: LYS A:10, GLU A:13, SER A:9; Pi interaction: LYS A:6; |
|
Anti-photoaging |
1G4K |
-4.3 |
Conventional hydrogen bonds: GLN A:236, ASN A:240; Pi–cation interaction: PHE A:83; |
|
Delta Opioid Receptor |
4N6H |
-9.5 |
Conventional hydrogen bond: TYR A:129; Pi–sigma interactions: VAL A:281, MET A:132; Pi–alkyl interactions: ILE A:277, ILE A:304, VAL A:217; |
|
Nitric Oxide Synthase |
3E7G |
-10.3 |
Conventional hydrogen bond: TYR D:489; Pi–pi stacked interactions: PHE D:369, TRP D:194; Pi–sigma interaction: ALA D:197 |
|
Anti Plasmodial |
1J3I |
-8.5 |
Conventional hydrogen bond: SER A:111; Pi–pi sigma interactions: ILE A:112, LEU A:46; |
|
CYP1A2 Inhibition |
2HI4 |
-10.0 |
Conventional hydrogen bond: ASN A:312; Pi–alkyl interaction: ALA A:317; Pi–pi stacked interactions: GLY A:316, PHE A:226; Pi–sigma interaction: LEU A:497; |
|
Plasminogen activator Inhibitor-1 |
3CVM |
-6.6 |
Conventional hydrogen bonds: ASN A:365, ASP A:231; Carbon–hydrogen bonds: ARG A:268, GLU A:225; Pi–alkyl interaction: PRO A:227; Pi–carbon interaction: LYS A:263 |
|
GABA Receptor |
6X3T |
-5.1 |
Conventional hydrogen bonds: ARG A:114, ASP A:89; Pi–sigma interaction: VAL A:111; Pi–cation interaction: ARG A:86; Pi–alkyl interaction: LYS A:112; |
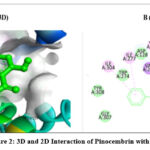 |
Figure 2: 3D and 2D Interaction of Pinocembrin with 4N6H |
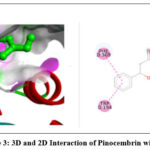 |
Figure 3: 3D and 2D Interaction of Pinocembrin with 3E7G |
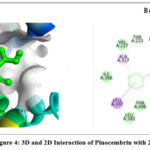 |
Figure 4: 3D and 2D Interaction of Pinocembrin with 2HI4 |
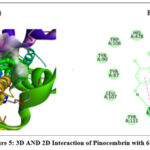 |
Figure 5: 3D AND 2D Interaction of Pinocembrin with 6OL9 |
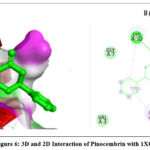 |
Figure 6: 3D and 2D Interaction of Pinocembrin with 1XQ8 |
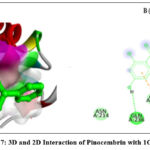 |
Figure 7: 3D and 2D Interaction of Pinocembrin with 1G4K |
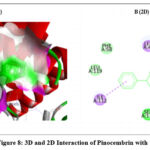 |
Figure 8: 3D and 2D Interaction of Pinocembrin with 1J3I |
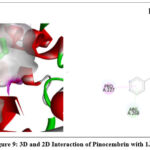 |
Figure 9: 3D and 2D Interaction of Pinocembrin with 1J3I |
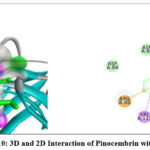 |
Figure 10: 3D and 2D Interaction of Pinocembrin with 6X3T |
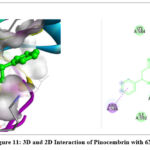 |
Figure 11: 3D and 2D Interaction of Pinocembrin with 6X3T |
DFT Calculations
Pinocembrin’s electrical and reactivity properties were studied by Density Functional Theory (DFT) simulations. The molecule’s chemical reactivity and stability were evaluated using the HOMO-LUMO energy gap. The HOMO and LUMO values were used to generate a number of global descriptors, including ionization energy, electron affinity, global hardness and softness, chemical potential, electronegativity, and electrophilicity index (Table 2). The ability of the molecule to give or receive electrons is revealed by these properties, which is crucial for forecasting its chemical activity and possible biological interactions (Fig. 12).
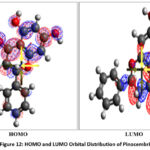 |
Figure 12: HOMO and LUMO Orbital Distribution of Pinocembrin |
Table 2: DFT-Derived Quantum Chemical Descriptors for Pinocembrin
|
Descriptor |
Value (eV) |
|
EHOMO |
–5.125 |
|
ELUMO |
–2.023 |
|
Band Gap (ΔE) |
3.102 |
|
Ionization Energy (IE) |
5.125 |
|
Electron Affinity (EA) |
2.023 |
|
Chemical Hardness (η) |
1.551 |
|
Chemical Softness (S) |
0.645 |
|
Chemical Potential (μ) |
–3.574 |
|
Electronegativity (χ) |
3.574 |
|
Electrophilicity Index (ω) |
4.118 |
ADME and Drug-Likeness Profile
Using PreADMET, we assessed the pharmacokinetic properties of Pinocembrin and it was determined that it exhibits favourable drug-like properties. It was not in violation of Lipinski’s rule of five and therefore showed as drug like with CMC-like, WDI-like and lead-like filters. Good bioavailability of the compound resulted in it being an ideal oral candidate.
Human intestinal absorption was shown to be greater than 92.35%, Caco-2 cell permeability was moderate (0.8×10−6 cm/sec), and MDCK cell permeability was high (8.6×10−6 cm/sec), suggesting efficient absorption and cellular transport. Skin permeability was low however, CNS activity via the blood-brain barrier (BBB) penetration of 0.91 was observed, indicating CNS activity.
Since Pinocembrin was predicted to inhibit cytochrome P450 (CYP) enzymes CYP2C9, CYP2C19 and CYP3A4, it has the potential to cause drug-drug interactions. The compound had a high degree of plasma protein binding (98.45%), and hence affected its pharmacokinetics and distribution.
SWISS ADME results showed that it has high gastrointestinal absorption, BBB permeability, no PAINS alarm, and no Lipinski’s Rule infractions were all demonstrated by pinocembrin. CYP1A2 is expected to be inhibited, but not other important CYP enzymes. Additionally, the molecule demonstrated a synthetic accessibility score of 3.21, which indicates ease of synthesis, and moderate water solubility (Table 4).
Both the white (HIA+) and yellow yolk (BBB+) zones of the BOILED-Egg diagram (Fig.13) contain the red dot that represents pinocembrin (Table 3). This supports the compound’s use in CNS illnesses by showing that it may be absorbed orally and cross the blood–brain barrier.
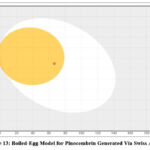 |
Figure 13: Boiled-Egg Model for Pinocembrin Generated Via Swiss ADME |
Table 3: Boiled Egg Regions and Meaning
|
Region |
Meaning |
|
White Region |
High probability of passive GI absorption (HIA+) |
|
Yellow Region (Yolk) |
High probability of BBB permeability (CNS-active) |
Table 4: PreADMET and Swiss ADME Data
|
Parameter |
Tool Used |
Result |
Interpretation |
|
Human Intestinal Absorption (HIA) |
PreADMET |
92.35% |
Excellent absorption capacity |
|
Caco-2 Permeability |
PreADMET |
0.8 × 10⁻⁶ cm/sec |
Moderate absorption |
|
MDCK Cell Permeability |
PreADMET |
8.6 × 10⁻⁶ cm/sec |
High membrane permeability |
|
Blood–Brain Barrier (BBB) Penetration |
PreADMET |
0.91 |
Likely to cross the BBB (CNS active) |
|
Skin Permeability |
PreADMET |
Low |
Limited transdermal absorption |
|
CYP Inhibition (CYP1A2) |
PreADMET |
Inhibitor |
May affect drug metabolism; potential for drug-drug interaction |
|
Plasma Protein Binding (PPB) |
PreADMET |
98.45% |
High binding, may influence distribution |
|
hERG Inhibition |
PreADMET |
Medium risk |
Needs further cardiotoxicity evaluation |
|
Mutagenicity (Ames Test) |
PreADMET |
TA100 (+), mutagenic |
Caution advised for genotoxic potential |
|
Carcinogenicity (Rodents) |
PreADMET |
Non-carcinogenic |
Safe in long-term rodent models |
|
Lipinski Rule of Five |
SWISS ADME |
0 violations |
Drug-like properties confirmed |
|
Veber/Ghose/Muegge/Egan Rules |
SWISS ADME |
All passed |
Consistent with oral bioavailability |
|
GI Absorption |
SWISS ADME |
High |
Good candidate for oral delivery |
|
BBB Permeation |
SWISS ADME |
Yes |
CNS activity supported |
|
CYP Inhibition (CYP1A2) |
SWISS ADME |
Inhibitor |
Confirms PreADMET result |
|
PAINS Alerts |
SWISS ADME |
0 alerts |
No false positives in assays |
|
Synthetic Accessibility |
SWISS ADME |
3.21 |
Moderately easy to synthesize |
|
Water Solubility (ESOL) |
SWISS ADME |
Soluble |
Suitable for oral formulation |
|
Bioavailability Score |
SWISS ADME |
0.55 |
Acceptable oral bioavailability |
Toxicity Profile
Pinocembrin was predicted to be mutagenic in the Ames test (TA100 positive) and non-carcinogenic in rats and mice, something reassuring from its long-term safety profile. Medium risk was shown for hERG inhibition, and the compound therefore should be investigated for cardiotoxicity further, particularly if used in CNS-related applications. However, its low aquatic toxicity suggests that it will have minimal environmental impact and will be developed further.
Discussion
Investigations into docking have shown that pinocembrin has multi-target activity, especially through strong interactions with enzymes of the nitric oxide synthase, CYP1A2 and delta opioid receptor. These are in line with potential anti-inflammatory, neuroprotective, and analgesic effects. Reduced binding scores (i.e. with elastase and GABA receptors) indicated minimal anti-photoaging and GABAergic modulatory activities. The DFT showed that the 3.102 eV energy difference indicated chemical stability and enough reactivity for biological processes. High electrophilicity indicates that the molecule can interact with protein active site nucleophilic amino acid residues, supporting molecular docking data. Pharmacokinetic predictions confirmed pinocembrin to be attractive as an orally active candidate with the possibility of crossing the blood-brain barrier. These are especially important in CNS-related signals like pain management and Alzheimer. However, CYP enzyme blockage and the medium-risk hERG blockage highlight the importance of considering any potential side effects and drug-drug interactions during the development of a drug under clinical conditions. There was no difference between prediction of toxicity and long-term safety, indicating that the risk of carcinogenicity was not present, but the risks of mutagenicity and cardiotoxicity should be investigated. The results suggest dose optimization and subsequent in vitro / in vivo validation. Overall, pinocembrin is a promising in vivo multi-target lead molecule with a pharmacological repertoire of anti-inflammatory, neuroprotective, and cardiovascular activities. Its positive ADME-drug likeness profile supports its potential. However, its estimated mutagenicity and hERG inhibitory capabilities point to the need for experimental validation and precautions in CNS-directed therapy.
Conclusion
In the present study, the in silico potential therapeutic multi-target efficacy of the natural flavonoid pinocembrin was investigated by molecular docking and ADMET predictions. Docking studies showed strong binding affinities with important target proteins (-9.5 for Delta Opioid Receptor; -10.0 for Cytochrome P450 1A2; and -10.3 for Nitric Oxide Synthase) . Potential uses in pain management, anti-inflammatory as well as neuroprotection are demonstrated with these interactions. Pharmacokinetic study also confirmed good oral bioavailability, strong intestinal absorption, blood–brain barrier permeability, and appropriate drug-likeness. However, cautious further testing is necessary due to the hERG risk and CYP450 enzyme inhibition. On the whole, they suggest pinocembrin to be a good lead compound for multi-target therapeutics, especially in the areas of inflammatory diseases, neurological diseases, and metabolic dysfunctions. More research with molecular dynamics simulations, toxicity profiling, and in vitro validation is required to verify its effectiveness and safety.
Acknowledgement
The authors acknowledge Department of Pharmaceutical Chemistry, SRM College of Pharmacy, SRMIST, for providing the necessary infra structure facilities and support during the research
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest-
The author(s) do not have any conflict of interest.
Data Availability Statement-
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials
Permission to reproduce material from other sources
Not Applicable
Author’s Contribution
- Sakthi Periyasamy: Conceptualization, Methodology, Data Collection, Molecular Docking Studies, DFT Calculations, ADMET Analysis, Writing Original Draft.
- Priya Devasigamani: Supervision, Validation, Writing – Review & Editing.
References
- Lan X, Wang W, Li Q, Wang J. The natural flavonoid Pinocembrin: molecular targets and potential therapeutic applications. Molecular Neurobiology. 2015;53(3):1794-1801. doi:10.1007/s12035-015-9125-2.
CrossRef - Danelutte AP, Lago JHG, Young MCM, Kato MJ. Antifungal flavanones and prenylated hydroquinones from Piper crassinervium Kunth. Phytochemistry. 2003;64(2):555-559. doi:10.1016/s0031-9422(03)00299-1
CrossRef - López A, Ming DS, Towers GHN. Antifungal Activity of Benzoic Acid Derivatives from Piper lanceaefolium. Journal of Natural Products. 2001;65(1):62-64. doi:10.1021/np010410g
CrossRef - Feng R, Guo ZK, Yan CM, Li EG, Tan RX, Ge HM. Anti-inflammatory flavonoids from Cryptocarya chingii. Phytochemistry. 2012;76:98-105. doi:10.1016/j.phytochem.2012.01.007
CrossRef - Jaganathan SK, Mandal M. Antiproliferative effects of honey and of its polyphenols: a review. BioMed Research International. 2009;2009(1). doi:10.1155/2009/830616
CrossRef - Jiang H, Morgan JA. Optimization of an in vivo plant P450 monooxygenase system in Saccharomyces cerevisiae. Biotechnology and Bioengineering. 2003;85(2):130-137. doi:10.1002/bit.10867.
CrossRef - Miyahisa I, Funa N, Ohnishi Y, Martens S, Moriguchi T, Horinouchi S. Combinatorial biosynthesis of flavones and flavonols in Escherichia coli. Applied Microbiology and Biotechnology. 2005;71(1):53-58. doi:10.1007/s00253-005-0116-5
CrossRef - Touil YS, Fellous A, Scherman D, Chabot GG. Flavonoid-Induced morphological modifications of endothelial cells through microtubule stabilization. Nutrition and Cancer. 2009;61(3):310-321. doi:10.1080/01635580802521346
CrossRef - Manthey J, Guthrie N, Grohmann K. Biological properties of citrus flavonoids pertaining to cancer and inflammation. Current Medicinal Chemistry. 2001;8(2):135-153. doi:10.2174/0929867013373723
CrossRef - Bertelli D, Papotti G, Bortolotti L, Marcazzan GL, Plessi M. 1H‐NMR Simultaneous identification of Health‐Relevant compounds in propolis extracts. Phytochemical Analysis. 2011;23(3):260-266. doi:10.1002/pca.1352
CrossRef - Danert FC, Zampini C, Ordoñez R, Maldonado L, Bedascarrasbure E, Isla MI. Nutritional and Functional Properties of Aqueous and Hydroalcoholic Extracts from Argentinean Propolis. Natural Product Communications. 2014;9(2). doi:10.1177/1934578×1400900209.
CrossRef - Gao M, Zhu SY, Tan CB, Xu B, Zhang WC, Du GH. Pinocembrin protects the neurovascular unit by reducing inflammation and extracellular proteolysis in MCAO rats. Journal of Asian Natural Products Research. 2010;12(5):407-418. doi:10.1080/10286020.2010.485129
CrossRef - Horinouchi S. Combinatorial biosynthesis of non-bacterial and unnatural flavonoids, stilbenoids and curcuminoids by microorganisms. The Journal of Antibiotics. 2008;61(12):709-728. doi:10.1038/ja.2008.85
CrossRef - Massaro CF, Katouli M, Grkovic T, et al. Anti-staphylococcal activity of C-methyl flavanones from propolis of Australian stingless bees (Tetragonula carbonaria) and fruit resins of Corymbia torelliana (Myrtaceae). Fitoterapia. 2014;95:247-257. doi:10.1016/j.fitote.2014.03.024
CrossRef - Miyahisa I, Kaneko M, Funa N, et al. Efficient production of (2S)-flavanones by Escherichia coli containing an artificial biosynthetic gene cluster. Applied Microbiology and Biotechnology. 2005;68(4):498-504. doi:10.1007/s00253-005-1916-3
CrossRef - Stashenko EE, Martínez JR, Ruíz CA, et al. Lippia origanoides chemotype differentiation based on essential oil GC‐MS and principal component analysis. Journal of Separation Science. 2009;33(1):93-103. doi:10.1002/jssc.200900452
CrossRef - Tuchinda P, Reutrakul V, Claeson P, et al. Anti-inflammatory cyclohexenyl chalcone derivatives in Boesenbergia pandurata. Phytochemistry. 2002;59(2):169-173. doi:10.1016/s0031-9422(01)00451-4
CrossRef - Lewis RA. A general method for exploiting QSAR models in lead optimization. Journal of Medicinal Chemistry. 2005;48(5):1638-1648. doi:10.1021/jm049228d
CrossRef - Bhatt S, Dhiman S, Kumar V, et al. Assessment of the CYP1A2 Inhibition-Mediated Drug Interaction Potential for Pinocembrin using in silico, in vitro, and in vivo approaches. ACS Omega. 2022;7(23):20321-20331. doi:10.1021/acsomega.2c02315
CrossRef - Melaku Y, Solomon M, Eswaramoorthy R, Beifuss U, Ondrus V, Mekonnen Y. Synthesis, antiplasmodial activity and in silico molecular docking study of pinocembrin and its analogs. BMC Chemistry. 2022;16(1). doi:10.1186/s13065-022-00831-z
CrossRef - Priya M, Zochedh A, Mohan YS, Chandran K, Arumugam K, Sultan AB. Network analysis and molecular modeling studies of pinocembrin a bioactive phytochemical of Dodonaea viscosa against Parkinson’s disease. In Silico Pharmacology. 2024;12(2). doi:10.1007/s40203-024-00268-3
CrossRef - Wang G, Cheng J, Yao M, Li J, Chen T, Zhang J, Du W, Chen Y. Network pharmacology-based strategy to reveal the mechanism of pinocembrin against ovarian cancer. Naunyn-Schmiedeberg’s Archives of Pharmacology. 2025 Apr;398(4):3803-15.
CrossRef - Teruna HY, Rullah K, Hendra R, et al. Inhibitory Effect of (2S)‐Pinocembrin From Goniothalamus macrophyllus on the Prostaglandin E2 Production in Macrophage Cell Lines. Advances in Pharmacological and Pharmaceutical Sciences. 2024;2024(1). doi:10.1155/2024/8811022.
CrossRef - Youn K, Jun M. Biological Evaluation and Docking Analysis of Potent BACE1 Inhibitors from Boesenbergia rotunda. Nutrients. 2019;11(3):662. doi:10.3390/nu11030662
CrossRef - Trott O, Olson AJ. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. Journal of Computational Chemistry. 2009;31(2):455-461. doi:10.1002/jcc.21334
CrossRef - Cheng T, Li X, Li Y, Liu Z, Wang R. Comparative assessment of scoring functions on a diverse test set. Journal of Chemical Information and Modeling. 2009;49(4):1079-1093. doi:10.1021/ci9000053
CrossRef - Li Y, Liu Z, Li J, et al. Comparative assessment of scoring functions on an updated benchmark: 1. Compilation of the test set. Journal of Chemical Information and Modeling. 2014;54(6):1700-1716. doi:10.1021/ci500080q
CrossRef - Wang R, Fang X, Lu Y, Wang S. The PDBbind Database: Collection of Binding Affinities for Protein−Ligand Complexes with Known Three-Dimensional Structures. Journal of Medicinal Chemistry. 2004;47(12):2977-2980. doi:10.1021/jm030580l
CrossRef - Dunkel M, Günther S, Ahmed J, Wittig B, Preissner R. SuperPred: drug classification and target prediction. Nucleic acids research. 2008 May 22;36(suppl_2):W55-9.
CrossRef - Das S, Chakrabarti S. Classification and prediction of protein–protein interaction interface using machine learning algorithm. Scientific Reports. 2021;11(1). doi:10.1038/s41598-020-80900-2
CrossRef - Song L, Ning S, Hou J, Zhao Y. Performance of protein-ligand docking with CDK4/6 inhibitors: a case study. Mathematical Biosciences & Engineering. 2020;18(1):456-470. doi:10.3934/mbe.2021025
CrossRef - ChemOffice. (2016). PerkinElmer informatics. http://www.cambridgesoft.com.
- Martins MO, Da Silva IZ, Fagan SB, Santos AFD. Docking fundamentals for simulation in nanoscience. Disciplinarum Scientia Série Ciências Naturais E Tecnológicas. 2021;22(3):67-76. doi:10.37779/nt.v22i3.4106.
CrossRef
Abbrevations List
ADMET – Absorption, Distribution, Metabolism, Excretion, and Toxicity
DFT – Density Functional Theory
hERG – Human Ether-à-go-go Related Gene (potassium channel)
CYP – Cytochrome P450 enzymes
pET – Plasmid Expression Vector (commonly used in recombinant protein expression)
CYP1A2 – Cytochrome P450 Family 1 Subfamily A Member 2
DHFR-TS – Dihydrofolate Reductase–Thymidylate Synthase
AKT1 – Protein Kinase B (AKT serine/threonine kinase 1)
TNF – Tumor Necrosis Factor
BCL2 – B-cell Lymphoma 2 (anti-apoptotic protein)
MAPK – Mitogen-Activated Protein Kinase
ERK2 – Extracellular Signal-Regulated Kinase 2
PGE2 – Prostaglandin E2
BACE1 – Beta-Site Amyloid Precursor Protein Cleaving Enzyme 1
HIA – Human Intestinal Absorption
BBB – Blood–Brain Barrier
MDCK – Madin–Darby Canine Kidney (cell line used in permeability studies)
PDB – Protein Data Bank
COX2 – Cyclooxygenase-2
ACHE – Acetylcholinesterase
NOS – Nitric Oxide Synthase
GABA – Gamma-Aminobutyric Acid
RCSB – Research Collaboratory for Structural Bioinformatics (maintains PDB)
PDBQT – Protein Data Bank, Qt (file format for AutoDock)
ORCA – ORCA Quantum Chemistry Program Package
HOMO – Highest Occupied Molecular Orbital
LUMO – Lowest Unoccupied Molecular Orbital
CMC – Critical Micelle Concentration
WDI – World Drug Index







