Manuscript accepted on :30-05-2025
Published online on: 13-06-2025
Plagiarism Check: Yes
Reviewed by: Dr. Daya Shankar Gautam
Second Review by: Dr. Alaa Saadi Abbood
Final Approval by: Dr. Patorn Piromchai
Department of Chemistry and Biochemistry, School of Medicine and Health Sciences, Atma Jaya Catholic University of Indonesia, Jakarta, Indonesia.
Corresponding Author E-mail: ferbian.siswanto@atmajaya.ac.id
DOI : https://dx.doi.org/10.13005/bpj/3173
Abstract
The loss of skeletal muscle mass and function during aging is associated with physical weakness and a higher risk of morbidity. MicroRNAs (miRNAs) have been proposed as promising biomarkers and therapeutic targets for aging, and several miRNAs are involved in the pathogenesis of various age-related diseases, underlining the significance of integrating miRNA with mRNA targets. The objectives of this study are to screen out the potential miRNA–mRNA pair(s) as biomarkers and therapeutic targets for skeletal muscle aging. In this study, two miRNA and four mRNA datasets were downloaded from the Gene Expression Omnibus (GEO) database. Comprehensive in silico analyses were performed to identify novel miRNA–mRNA pair(s) critically involved in skeletal muscle aging. A total of three miRNAs were found to be downregulated in aged muscle (miR-664b-3p, miR-208a-3p, and miR-365a-3p). In the four mRNA datasets, three common differentially expressed mRNAs were identified, one of which was consistently upregulated CDKN1A. The identified three miRNAs are potential biomarkers of skeletal muscle aging. The miR-208a-3p targets the 3' UTR of CDKN1A transcript. The regulatory network of miR-208a-3p expression during aging involves single nucleotide polymorphisms (SNPs) in the mature miRNA and its promoter/enhancers. This study established, for the first time, that miR-664b-3p, miR-208a-3p, and miR-365a-3p are potential biomarkers for skeletal muscle aging, but only miR-208a-3p can target CDKN1A. Therefore, miR-208a-3p–CDKN1A pair has the potential as a therapeutic agent for skeletal muscle aging.
Keywords
Aging; biomarkers; CDKN1A; Cellular senescence; Epigenetics; miR-208a; Skeletal muscle
Download this article as:| Copy the following to cite this article: Siswanto S. M. In Silico Analysis of MicroRNA-mRNA Interaction as Potential Biomarker in Skeletal Muscle Aging. Biomed Pharmacol J 2025;18(2). |
| Copy the following to cite this URL: Siswanto S. M. In Silico Analysis of MicroRNA-mRNA Interaction as Potential Biomarker in Skeletal Muscle Aging. Biomed Pharmacol J 2025;18(2). Available from: https://bit.ly/45TAntf |
Introduction
The World Health Organization (WHO) reports that the global population is aging rapidly, particularly in Asia, as life expectancy increases.1 By 2050, the number of individuals aged 60 years and older is projected to reach 2 billion—twice the number recorded in 2019.2 As the world’s fourth most populous country, Indonesia is experiencing a similar trend. In 2010, only 5% of the Indonesian population was elderly, a figure expected to rise to 11% by 2035.3 This demographic shift is primarily driven by a declining fertility rate. The elderly dependency ratio (the proportion of individuals aged 65+ relative to those aged 15–64) is projected to increase from under 10% in 2020 to over 46% by 2100.1 Life expectancy in Indonesia is also rising, from 72.5 years in 2015 to an estimated 75.5 years by 2045.4
As people live longer, limitations in physical activity become more common, negatively affecting quality of life. A key feature of aging associated with decreased physical activity is sarcopenia—a progressive decline in skeletal muscle mass and function.5 Beginning at age 30, skeletal muscle mass decreases by approximately 1% annually, accelerating after age 70.6 Sarcopenia is among the most prominent and debilitating age-related changes, contributing to physical disability and chronic disease. In Western countries, up to 42% of adults over 60 experience functional impairments.7 In Indonesia, an estimated 25–45% of older adults report physical limitations, which increase risks of falls, hospitalization, comorbidities, and premature death—thereby placing greater demands on healthcare systems.8
Preventing and managing physical decline is therefore essential for promoting healthy aging. Physical training remains the most widely recommended strategy to counteract muscle aging, tailored to individual needs and preferences.9 However, despite significant advances, the molecular mechanisms underlying skeletal muscle aging remain incompletely understood, and no definitive biomarker reliably predicts age-related functional decline.
MicroRNAs (miRNAs) are emerging as key regulators of aging and longevity. These small (19–22 nucleotide) noncoding RNAs typically bind to the 3′ untranslated region (UTR) of target mRNAs to repress translation or induce degradation.10 miRNAs influence various aging processes, including cellular proliferation and senescence.11 In skeletal muscle, a subset of miRNAs—known as myomiRs (e.g., miR-1, miR-133, miR-206, miR-208, miR-486, and miR-499)—are muscle-enriched and regulate key cellular processes such as differentiation, plasticity, and apoptosis.12 Altered expression of these miRNAs has been observed in aging muscle, potentially contributing to anabolic resistance and muscle loss.13 Despite increasing evidence, a comprehensive analysis of miRNA–mRNA interactions in skeletal muscle aging remains limited.
With advances in molecular biology, bioinformatics has become an essential tool in aging research. In silico analyses facilitate the identification of regulatory pathways, therapeutic targets, and disease mechanisms. While approximately 60% of protein-coding genes may be regulated by miRNAs, only a small fraction of these interactions has been experimentally validated.14 Several algorithms now enable the prediction of miRNA targets, expression profiles, signaling pathways, and transcription factor interactions.
In this study, miRNA expression profiles from aged muscle were analyzed using publicly available datasets from the Gene Expression Omnibus (GEO). Differentially expressed miRNAs and mRNAs were identified, followed by GO and KEGG pathway enrichment analyses. The predicted miRNA–mRNA interaction pairs were further refined. This work contributes to the understanding of the regulatory network involved in skeletal muscle aging and provides a foundation for future functional studies.
Materials and Methods
Microarray data
First, the publicly available miRNA dan mRNA datasets were screened by using the Gene expression omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo) database with the keywords “skeletal muscle”, “aging”, “miRNA” and “mRNA”. Public miRNA expression data sets (GSE23527 and GSE165632) and gene expression datasets (GSE1428, GSE167186, GSE111006, and GSE165630) were selected and downloaded. The GSE23527 contains miRNA array data derived from muscle biopsy samples of 36 men (19 young samples: 31±2 years old; 17 older samples: 73±3 years old). The GSE165632 miRNA array data of the vastus lateralis muscle contains data from 5 fully sedentary aged men (65 to 80 years) and 9 age-matched athletes. For gene/mRNA expression profiling in aging muscle, 10 young (19-25 years old) and 12 older (70-80 years old) males in the GSE1428 dataset, 19 young healthy (18-27 years old) and 24 old sarcopenic (67-92 years old) males in the GSE167186 dataset, 28 healthy old (68-77 years old) and 12 age-matched sarcopenic males in the GSE111006 dataset, and 5 fully sedentary aged men (65 to 80 years) and 9 age-matched athletes in the GSE165630 dataset were assessed. The datasets that compare sedentary and age-matched athletes were included due to a limited datasets that compare young and old samples for miRNA expression.
Differentially expressed miRNAs and mRNAs
The raw data of GSE23527, GSE165632, GSE1428, GSE167186, GSE111006, and GSE165630 were processed using a built-in interactive online tool used to identify differentially expressed miRNA and mRNA from GEO series called GEO2R (https://www.ncbi.nlm.nih.gov/geo/geo2r/). A false discovery rate (FDR) corrected p-value of < 0.05 was used to screen for differentially expressed miRNAs and mRNAs, and a screening threshold of |log2 fold change (FC)| >1.0 was chosen for each group. SRplot (http://www.bioinformatics.com.cn/srplot), an online platform for data analysis and visualization, was used to present and integrate the differentially expressed mRNAs and miRNAs in each dataset as volcano plots into Venn diagrams.
MiRNA targets prediction and miRNA–mRNA Target Network Analysis
Encyclopedia of RNA Interactomes (ENCORI, https://rnasysu.com/encori/) and miRTargetLink 2.0 (https://ccb-compute.cs.uni-saarland.de/mirtargetlink2) are two interactive bioinformatics platforms to facilitate the integrative and comprehensive annotation and discovery of RNA interactome. In the present study, the targets of the differentially expressed miRNAs were predicted using both ENCORI and miRTargetLink (version 2.0). To obtain a clear visualization of the miRNAs-mRNAs interaction network, the differentially expressed miRNAs and strongly validated target genes network were constructed using miRTargetLink (version 2.0). For this interactive network, only experimentally validated and manually curated miRNA–target interactions were used from miRTarBase.
Functional enrichment analysis of predicted targets of miRNAs
A popular web-based tool for functional annotation of genomic sequences, Database for Annotation, Visualization, and Integrated Discovery (DAVID, https://david.ncifcrf.gov/), was used to perform the Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathway and Gene Ontology (GO) biological process enrichment analysis of the predicted targets of differentially expressed miRNAs. A cut-off criterion of less than 0.05 for the FDR was applied. Using SRplot, Venn diagrams incorporating the significantly enriched KEGG pathway and GO biological process were created from both ENCORI and miRTargetLink 2.0.
Construction of miRNA–mRNA regulatory pair
MiRNA–mRNA pairs associated with aging of the skeletal muscle were chosen and assembled based on the target prediction outcomes of mRNAs and miRNAs with differential expression. The putative regulatory target of selected miRNA in the region of the selected gene was predicted using TargetScan7 (http://www.targetscan.org/). TargetScan7 is a web server that predicts miRNA targets using an algorithm based on conserved 8-mer, 7-mer, and 6mer sites that correspond to each miRNA’s seed region. To further validate the miRNA–mRNA pair involved in the aging process of skeletal muscle, miRTarBase (https://mirtarbase.cuhk.edu.cn/) was used, which is the most up-to-date and extensive collection of experimentally verified miRNA target interactions with enhanced annotation data. TargetScan7 was used to predict the conservation of target sequence across multiple genomes. The phyloP scores from the alignment of 100 genomes were used to calculate the conservation scores between seed binding positions and flanking positions.
Tissue and sub-cellular distribution of miRNA-mRNA pair
A visual display of selected miRNA and mRNA pair expression in a sub-cellular context and across human normal tissues was retrieved from an integrative online database, GeneCards (http://www.genecards.org/). RNAseq dataset from the Genotype-Tissue Expression (GTEx) project in the context of tissue distribution was selected for this study. The COMPARTMENTS resource, which combines manual literature curation, high-throughput microscopy-based screens, predictions from primary sequence, and automatic text mining mapped onto Gene Ontology terms, was used to determine the sub-cellular localizations of a specific miRNA and mRNA pair. Color codes are used to indicate the localization evidence’s confidence levels; light green (1) denotes low confidence and dark green (5) denotes high confidence. The color white (0) denotes the lack of evidence for localization.
Protein-protein interaction (PPI) networks of CDKN1A
The PPI network of the CDKN1A was constructed and visualized using the Search Tool for the Retrieval of Interacting Genes (STRING, https://string-db.org/) database to predict the key signalling pathways and cellular activities in skeletal muscle aging [21]. An interaction score > 0.9 as a threshold was used to identify the significant PPIs.
SNPs identification in miRNA genes and regulatory regions
An online miRNASNP-v3 database (http://bioinfo.life.hust.edu.cn/miRNASNP/#!/) that includes a catalogue of SNPs in 1,897 human pre-miRNAs (2,624 mature miRNAs) and SNP-induced gain and loss of miRNA targets was used to predict SNPs in pre- or mature miRNA and their predicted effects on mature miRNA expression. Based on the difference between the minimal free energy (MFE) between wild-type and mutant alleles (ΔG) calculated by RNAfold-v2, the predicted effects of SNPs on mature miRNA expression were calculated. The computational tool SNP2TFBS (https://epd.expasy.org/snp2tfbs/) was used to analyze the impact of SNPs in enhancer and promoter regions on transcription factor binding affinity. First, the GeneHancer framework of GeneCards was used to predict the promoter and enhancer regions of miRNAs. The UCSC website (https://genome.ucsc.edu/) was used to verify the obtained areas, and all SNP data within these areas was obtained. The motif of TFBS from the carefully selected JASPAR CORE 2014 vertebrate motif database was then analyzed using SNP2TFBS by computing the Position Weight Matrix (PWM). In order to predict transcription factors that control the expression of particular miRNAs, the TransmiR v2.0 database, which includes information derived from ChIP-seq evidence was used. Fig. 1 provides an overview of the research methodology employed in this study.
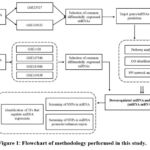 |
Figure 1: Flowchart of methodology performed in this study.Click here to view Figure |
Results
Differentially expressed miRNAs in aging muscle
The miRNA array results from the GEO database with the accession numbers GSE23527 and GSE165632 were chosen. The number of differentially expressed miRNAs obtained was 23 and 91 in GSE23527 and GSE165632, respectively. In GSE23527, eight upregulated and 15 downregulated miRNAs were observed (Fig. 2A), and in GSE165632, there were 53 upregulated and 38 downregulated miRNAs (Fig. 2B). The Venn diagram was used to identify overlapping miRNAs in both sets. Three miRNAs were identified as commonly downregulated miRNAs, including miR-664b-3p, miR-208a-3p, and miR-365a-3p (Fig. 2C).
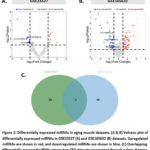 |
Figure 2: Differentially expressed miRNAs in aging muscle datasets. (A & B) Volcano plot of differentially expressed miRNAs in GSE23527 (A) and GSE165632 (B) datasets. Click here to view Figure |
Because changes in the levels of multiple miRNAs can produce biological outcomes that work in concert through redundant or cooperative mechanisms to regulate gene expression, pathway and biological processes enriched in the sets of predicted target genes of selected miRNAs were investigated. The validated and predicted gene targets of miR-664b-3p, miR-208a-3p, and miR-365a-3p were first retrieved using ENCORI and miRTargetLink 2.0 databases, and Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathway and Gene Ontology (GO) biological process enrichment analysis were constructed using DAVID. Venn diagram showed a total of 47 KEGG pathways were significantly enriched (FDR < 0.05) in ENCORI-predicted gene targets of the three miRNAs (Fig. 3A, B). Two pathways were significantly enriched in miRTargetLink-predicted sets of target genes of the three miRNAs, including apoptosis and pathway in cancer (Fig. 3C, D). The GO annotation demonstrated 214 significantly enriched biological processes in ENCORI-predicted gene targets of the three miRNAs (Fig. 3E, F). Two pathways, regulation of macromolecule metabolic process and regulation of metabolic process, were significantly enriched in miRTargetLink-predicted sets of target genes of the three miRNAs (Fig. 3G, H). To visualize the target gene network of miR-664b-3p, miR-208a-3p, and miR-365a-3p, the interactive network was built using miRTargetLink 2.0 databases. Of the strongly validated miRNA targets, nine were identified as the targets of miR-208a-3p, and the other nine were targets of miR-365a-3p. MiR-664b-3p has no targeted genes (Fig. 4).
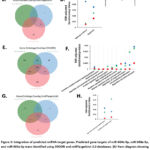 |
Figure 3: Integration of predicted miRNA target genes. Predicted gene targets of miR-664b-3p, miR-208a-3p, and miR-365a-3p were identified using ENCORI and miRTargetLink 2.0 databases. Click here to view Figure |
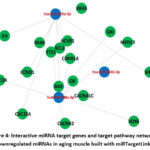 |
Figure 4: Interactive miRNA target genes and target pathway networks of downregulated miRNAs in aging muscle built with miRTargetLink 2.0.Click here to view Figure |
Differentially expressed mRNAs in aging muscle
The microarray results from the GEO database with the accession number GSE1428, GSE167186, GSE111006, and GSE165630 were chosen in the present study. A total of 899, 506, 5471 and 354 differentially expressed mRNAs were identified in GSE1428 (Fig. 5A), GSE167186 (Fig. 5B), GSE111006 (Fig. 5C), and GSE165630 (Fig. 5D), respectively. The Venn diagram identified three overlapping mRNAs in all datasets, including follistatin (FST), adrenoceptor alpha 2A (ADRA2A), and cyclin-dependent kinase inhibitor 1A (CDKN1A) (Fig. 5E). FST was upregulated in GSE167186, GSE111006, and GSE165630, but not in GSE1428 datasets. ADRA2A was upregulated in the GSE111006 and GSE165630 datasets. CDKN1A was consistently upregulated in all four datasets.
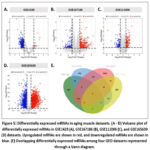 |
Figure 5: Differentially expressed mRNAs in aging muscle datasets. (A – D) Volcano plot of differentially expressed mRNAs in GSE1428 (A), GSE167186 (B), GSE111006 (C), and GSE165630 (D) datasets. Click here to view Figure |
Construction of miRNA-mRNA pair critical for skeletal muscle aging
In the present study, three miRNAs (miR-664b-3p, miR-208a-3p, miR-365a-3p) and three mRNAs (FST, ADRA2A, and CDKN1A) with altered expression in aging muscle were found. All three miRNAs were consistently downregulated in all datasets, whereas the expression of mRNAs differed. Since in most cases, miRNAs function as a guide by base-pairing with target mRNA to suppress its expression, this study focused on CDKN1A as a consistently upregulated gene in aging muscle. Analysis of the 3′ UTR region of human CDKN1A with TargetScan7 revealed several potential miRNAs, one of which was miR-208-3p with an exact match to positions 2-7 of the mature miRNA (the seed) and includes the A at position 1 (Fig. 6A, upper panel). Further analysis of miR-208a-3p–CDKN1A pair with ENCORI suggested the target-directed miRNA degradation (TDMD) score of 0.9878 (high confidence ≥ 0.8) and the evolutionary conservation (phyloP) scores of 3.239 (positive score= conserved) (Fig. 6A, lower panel).15 The predicted miR-208a-3p target site in the 3′ UTR of human CDKN1A is highly conserved among mammals (Fig. 6B). Of note, this study found a predicted miR-365a-3p target site within the poorly conserved region of CDKN1A 3′ UTR, while no miR-664b-3p target site was detected (data not shown).
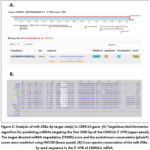 |
Figure 6: Analysis of miR-208a-3p target site(s) in CDKN1A gene. (A) TargetScan bioinformatics algorithm for predicting miRNAs targeting the first 1500 bp of the CDKN1A 3′ UTR (upper panel). Click here to view Figure |
Based on these results, miR-208a-3p–CDKN1A pair is potentially critical for skeletal muscle aging (Fig. 7A). The expression of the CDKN1A was high, and the expression of the miR-208a-3p was low in the aging muscle cohort (Fig. 7B-E). To further validate this proposed working model of the miR-208a-3p–CDKN1A pair, the information from miRTarBase was retrieved. This pair has been experimentally proven using reporter assay, Western blot, qPCR, and other methods (Fig. 7F). Next, the tissue and sub-cellular distribution of miR-208a-3p and CDKN1A were investigated. MiR-208a-3p was expressed only in a small number of tissues, while CDKN1A was expressed ubiquitously. These findings suggested that both were expressed in skeletal muscle (Fig. 7G). In the context of sub-cellular localization, both factors were expressed in almost all compartments of the cell, except lysosome, endosome, peroxisome (for miR-208a-3p), and Golgi apparatus (for both) (Fig. 7H). Thus, the interaction between miR-208a-3p and CDKN1A can occur in almost all parts of the cell, particularly in the cytosol and nucleus, where CDKN1A is dominantly expressed.
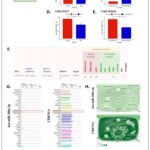 |
Figure 7: The relationship between miR-208a-3p and CDKN1A. (A) Hub gene CDKN1A mRNA was negatively regulated by miR-208a-3p. (B-C) The expression of CDKN1A mRNA in GSE1428 (B) and GSE165630 (C) datasets. Click here to view Figure |
CDKN1A controls the cell cycle and the G2 arrest caused by DNA damage. It is involved in the DNA damage-induced suppression of cellular growth via p53-dependent or p53-independent pathways. Additionally, CDKN1A suppresses the activity of cyclin-dependent kinases by preventing the phosphorylation of vital cyclin-dependent kinase substrates and halting the progression of the cell cycle.16 To visualize the network of CDKN1A, the protein-protein interaction (PPI) network of CDKN1A was downloaded from STRING (Fig. 8).
 |
Figure 8: The STRING protein-protein interaction network queried with CDKN1A. Coloured lines between the proteins indicate the various types of interaction evidence.Click here to view Figure |
Roles of SNPs in miRNA genes and regulatory regions
Studies have indicated that single nucleotide polymorphisms (SNPs) in various genes are important factors in determining human aging and longevity. In fact, previous study found that deletion SNP in the angiotensin-converting enzyme gene is responsible for skeletal muscle aging.17 To understand the mechanism by which the expression of miR-208a-3p in skeletal muscle is controlled by aging, SNPs in miRNA genes was analyzed computationally. The miRNASNPV3 was employed to search and predict the effects of SNPs in miR-208a-3p. Several SNPs in pre-miRNA and mature miRNA were identified in this study (Table 1). All SNPs, with the exception of rs1208723665 in pre-miRNA, had positive ΔG. It is anticipated that the pre-miRNA structure would be thermodynamically more stable with less MFE because the ΔG represents the difference in MFE between the mutant and the wild type alleles.18 The mature miRNA deletion variant rs755889109 exhibited the highest positive ΔG (7.1), indicating that it is the cause of the lowest mature miRNA expression rate.
Table 1: List of known SNPs in the miR-208a-3p and their effects on mature miRNA expression according to miRNASNPv3.
| SNP ID | Position | Ref/Alt | Region | ΔG | Predicted effect on mature miRNA expression |
| rs755889109 | chr14:23388606 | GC/G | in_mature | 7.1 | down |
| rs1346967645 | chr14:23388606 | G/A | in_mature | 2.4 | down |
| rs372515832 | chr14:23388616 | C/T | pre-miRNA | 3.6 | down |
| rs372515832 | chr14:23388616 | C/G | pre-miRNA | 2.7 | down |
| rs756125941 | chr14:23388617 | G/A | pre-miRNA | 2.3 | down |
| rs1208723665 | chr14:23388619 | C/T | pre-miRNA | -0.5 | mild |
Next, the impacts of SNPs on the regulatory regions of miR-208a-3p was further investigated. SNPs on the regulatory regions may modify miRNA expressions by altering TFBS for the transcription factor to bind—increase, decrease, or no effect. Both promoters and gene-distal cis-regulatory regions (or enhancers) of the MIR208A gene were predicted using the GeneHancer framework of GeneCards. One immediate upstream promoter (-1 to -2000 bp from TSS), one distal upstream enhancer (TSS distance= -56.4 kb), and one distal downstream enhancer (TSS distance= +60.4 kb) were selected in this study. Obtained regions were checked at the UCSC site (https://genome.ucsc.edu/), and all SNP information in these areas was retrieved. Based on SNP2TFBS, the rs73604575, rs28730769, rs111403367, and rs911390 polymorphisms most likely create the new transcription factor recognition. On the contrary, rs555976716, rs57660219, and rs200618133 in the promoter region and rs115881412 and rs60487758 in the enhancer region were likely to reduce the expression of miR-208a-3p due to a negative Scorediff (Table 2). A negative score means a more significant PWM score in the wild-type allele (reference genome, Ref) than the alternating allele (Alt), which could give rise to the loss of the original transcription factor recognition site.
Table 2: List of SNPs located in the promoter and enhancer region of MIR208A coding gene and their effects on transcription factor binding activity performed by SNP2TFBS.
| Region | Type | More PWM score on Alt (Scorediff +) missing in ref | More PWM score on Ref (Scorediff -) missing in alt | Neutral |
| chr14:23,859,360-23,861,360 | Proximal Upstream Promoter (TSS distance= 0 kb) |
rs73604575 rs28730769 |
rs555976716 rs57660219 rs200618133 |
rs73604573 rs28730770 rs112405990 rs28730771 |
| chr14:23,910,811-23,917,840 | Distal Upstream Enhancer (TSS distance= -56.4 kb) |
rs111403367 rs911390 |
rs115881412 rs60487758 |
rs11621360 rs59824205 rs73587650 rs17092504 rs73587646 |
| chr14:23,797,010-23,797,862 | Distal Downstream Enhancer (TSS distance= +60.4 kb) |
– | – | rs7161120 rs8003299 rs4981469 rs576209370 rs4981468 |
The rs555976716, rs57660219, and rs200618133 are polymorphisms in the KLF5 binding site with a PMW score of -1, while rs115881412 and rs60487758 are variations in the ZNF263 binding site with a PMW score of -223. Based on these PMW scores, the low expression of miR-208a-3p in aging muscle is likely due to rs115881412 and rs60487758 polymorphisms that affect the binding of transcription factor ZNF263 to the enhancer region of MIR208A.
Transcription factor-miRNA regulatory network in aging muscle
To study the regulation of miR-208a-3p in skeletal muscle aging, transcription factor(s) responsible for the regulation of miR-208a-3p expression was predicted using TransmiR database. Several transcription factors were predicted to regulate the expression of miR-208a-3p, including BATF, E2F6, ELF1, GTF2I, JUND, MAX, RUNX, SFPQ, TET2, and ZNF263 (Table 3). To confirm which transcription factor(s) plays role in the regulation of miR-208a-3p expression in skeletal muscle during aging, the expression of these transcription factors in the GSE1428 dataset was analyzed. The results showed that only BATF mRNA levels were downregulated in aging muscle (Fig. 9). Based on these results, the basic leucine zipper ATF-like transcription factor (BATF) is likely responsible for the reduced expression of miR-208a-3p during skeletal muscle aging.
Table 3: List of predicted TFBS in the regulatory region of mir-208a retrieved from TransmiR v2.0 database.
| TSS | TF name | Binding site |
| chr14: 23408277 | BATF | chr14: 23409573-23409727(score=1000) |
| chr14: 23388667 | E2F6 | chr14: 23390142-23390312(score=900) |
| chr14: 23388668 | ELF1 | chr14: 23392733-23392849(score=435) |
| chr14: 23388669 | GTF2I | chr14: 23388219-23389081(score=1000) |
| chr14: 23388670 | GTF2I | chr14: 23388292-23389006(score=1000) |
| chr14: 23388671 | GTF2I | chr14: 23388301-23388996(score=1000) |
| chr14: 23388672 | GTF2I | chr14: 23388368-23388932(score=806) |
| chr14: 23388673 | GTF2I | chr14: 23388387-23388890(score=896) |
| chr14: 23388674 | GTF2I | chr14: 23388424-23388868(score=562) |
| chr14: 23388675 | GTF2I | chr14: 23388439-23388843(score=476) |
| chr14: 23388676 | JUND | chr14: 23390002-23390177(score=815) |
| chr14: 23388677 | JUND | chr14: 23390014-23390187(score=997) |
| chr14: 23388678 | JUND | chr14: 23389975-23390197(score=1000) |
| chr14: 23388679 | MAX | chr14: 23390106-23390280(score=608) |
| chr14: 23388680 | RUNX1 | chr14: 23390196-23390411(score=286) |
| chr14: 23388681 | SFPQ | chr14: 23389881-23390355(score=515) |
| chr14: 23388682 | TET2 | chr14: 23390176-23390398(score=295) |
| chr14: 23408277 | ZNF263 | chr14: 23410222-23410586(score=891) |
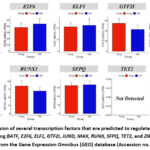 |
Figure 9: The expression of several transcription factors that are predicted to regulate miR-208a-3p during muscle aging, including BATF, E2F6, ELF1, GTF2I, JUND, MAX, RUNX, SFPQ, TET2, and ZNF263. Click here to view Figure |
Discussion
Age-related skeletal muscle loss, or sarcopenia, progresses at an estimated rate of 1% per year beginning in midlife.6 This involuntary loss of muscle mass and function compromises mobility, reduces quality of life, and increases the risk of fall-related injuries. Sarcopenia is also associated with systemic conditions such as metabolic syndrome, non-alcoholic fatty liver disease, and cardiovascular disease.19 Its pathophysiology is multifactorial, involving impaired protein turnover, mitochondrial dysfunction, chronic inflammation, satellite cell exhaustion, oxidative stress, and apoptosis.20 Despite growing knowledge, the molecular mechanisms driving skeletal muscle aging remain incompletely defined.
Recent research has turned attention toward the regulatory role of microRNAs (miRNAs) and their target mRNAs. Some skeletal muscle-specific miRNAs (myomiRs) have been identified as critical modulators of age-related muscle degeneration.21 In this study, microarray datasets from the GEO database were used to identify differentially expressed miRNAs and mRNAs in aged muscle tissues. Three miRNAs (miR-664b-3p, miR-208a-3p, and miR-365a-3p) were consistently dysregulated in aged muscle, with enrichment analyses linking their target genes to apoptosis and macromolecule metabolism—both key processes in muscle aging.22
Subsequent analysis identified three differentially expressed genes (FST, ADRA2A, and CDKN1A), with a notable miRNA–mRNA interaction between miR-208a-3p and CDKN1A. Although miR-208a has been studied extensively in cardiovascular diseases,23 its role in skeletal muscle aging remains poorly understood. In contrast, miR-208b is known to influence muscle fiber-type specification and proliferation, but its inconsistent expression limits its utility as a biomarker. The results of present study reveal a consistent downregulation of miR-208a in aging muscle, suggesting its potential diagnostic and therapeutic value. Notably, miR-208a has been shown to regulate mitochondrial biogenesis and function, processes integral to aging.24
CDKN1A (p21/Cip1/Waf1), a cyclin-dependent kinase inhibitor encoded on chromosome 6p21.2, is a key regulator of cell cycle arrest and senescence.25 It inhibits cyclin-CDK complexes and DNA replication by binding to proliferating cell nuclear antigen (PCNA).16 CDKN1A expression is elevated in aging tissues and is widely recognized as a senescence marker.26 In this study, CDKN1A emerged as a central target of miR-208a-3p, supporting a role for this regulatory pair in skeletal muscle aging. Previous research has confirmed the interaction of miR-208a-3p and CDKN1A in various cancer contexts, including lung27 and prostate cancers28, and the analogous miR-208b–CDKN1A pair has been shown to promote muscle cell proliferation.29
Genetic variants such as single nucleotide polymorphisms (SNPs) in miRNA genes or their regulatory regions can affect miRNA expression and function.30 Analysis in this study showed that all known SNPs in miR-208a-3p, except rs1208723665, had positive ΔG values—suggesting enhanced miRNA expression. Among these, rs755889109 (a GC/G deletion) had the highest ΔG and may promote maturation of miR-208a-3p. Although the clinical significance of this SNP remains unclear, it may influence aging by modulating miRNA and CDKN1A levels.
Around 90% of disease-associated SNPs lie in non-coding regions31, and many are located in miRNA promoters or enhancers where they affect transcription factor binding.32 Data in the present study suggest that low miR-208a-3p expression in aging muscle could be due to SNPs such as rs115881412 and rs60487758, which disrupt the binding of ZNF263 to the enhancer region of MIR208A. ZNF263 is a key transcription factor involved in gene regulation and cell proliferation and has been implicated in skeletal muscle development.33 Previous findings show ZNF263 levels remain unchanged with age34, suggesting the downregulation of miR-208a-3p is likely due to loss of enhancer function rather than reduced ZNF263 expression.
Using the TransmiR database, several potential regulators of miR-208a-3p were identified. Among them, BATF was the only transcription factor downregulated in aging muscle. BATF, a bZIP transcription factor, is known for its role in lymphocyte differentiation and immune function35, but its involvement in muscle aging is uncharacterized. These findings suggest that BATF may regulate miR-208a-3p and warrant further investigation.
While this in silico analysis offers valuable insights, several limitations must be acknowledged. The study relies on publicly available transcriptomic datasets, which may vary in quality and experimental conditions. Bioinformatics predictions lack biological context and do not account for post-transcriptional modifications or tissue-specific regulation. Functional validation using techniques such as qRT-PCR, luciferase reporter assays, and in vivo models is essential to confirm the proposed interactions and assess their relevance as diagnostic or therapeutic targets in skeletal muscle aging.
Conclusion
The present in silico study elucidated the mechanistic miRNA-mRNA regulatory network related to the aging process of skeletal muscle using several datasets from the GEO database. CDKN1A is famously known to be involved in cellular senescence, which is consistent with KEGG pathway enrichment and GO annotation of predicted miR-208a-3p targets. The miR-208a-3p–CDKN1A pair might serve as a potential biomarker and therapeutic target to ameliorate and block aging-related changes in skeletal muscle. Following a deep screening of miR-208a-3p, this study provides a comprehensive analysis of the SNPs and their effects on miR-208a-3p biogenesis and function, including pre-miRNA processing level, miRNA-target interaction, and transcript level, which may affect skeletal muscle aging. However, it is recommended that more clinical investigations be conducted in the laboratory with several cohorts and a larger number of individuals, stratified by age and training level, in order to provide further validation.
Acknowledgement
The author thanks the School of Medicine and Health Sciences, Atma Jaya Catholic University of Indonesia for providing facilities.
Funding Sources
This study was supported by Grant-in-Aid from Atma Jaya Catholic University of Indonesia (Grant no. 0878 /III.D.FKIK-KP.104.01/02/2024).
Conflict of Interest
The author(s) do not have any conflict of interest
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable.
Author Contributions
The sole author was responsible for the conceptualization, methodology, data collection, analysis, writing, and final approval of the manuscript.
References
- Kudrna G, Le T, Piggott J. Macro-Demographics and Ageing in Emerging Asia: the Case of Indonesia. J Popul Ageing. 2022;15(1):7-38. doi:10.1007/s12062-022-09358-6
CrossRef - World Health Organization. Ageing and health. Published 2022. Accessed August 28, 2023. https://www.who.int/news-room/fact-sheets/detail/ageing-and-health
- Basrowi RW, Rahayu EM, Khoe LC, Wasito E, Sundjaya T. The Road to Healthy Ageing: What Has Indonesia Achieved So Far? Nutrients. 2021;13(10):3441. doi:10.3390/nu13103441
CrossRef - BPS-StatisticsIndonesia. Indonesia Population Projection 2015–2045 Result of SUPAS 2015. Revised Ed.; 2018.
CrossRef - Larsson L, Degens H, Li M, et al. Sarcopenia: Aging-Related Loss of Muscle Mass and Function. Physiol Rev. 2019;99(1):427-511. doi:10.1152/physrev.00061.2017
CrossRef - Mercken EM, Majounie E, Ding J, et al. Age-associated miRNA Alterations in Skeletal Muscle from Rhesus Monkeys reversed by caloric restriction. Aging (Albany NY). 2013;5(9):692-703. doi:10.18632/aging.100598
CrossRef - Louie GH, Ward MM. Sex Disparities in Self-Reported Physical Functioning: True Differences, Reporting Bias, or Incomplete Adjustment for Confounding? J Am Geriatr Soc. 2010;58(6):1117-1122. doi:10.1111/j.1532-5415.2010.02858.x
CrossRef - Satria A, Yeni Y, Akbar H, et al. Relationship between depression and physical disability by gender among elderly in Indonesia. Universa Med. 2022;41(2):104-113. doi:10.18051/UnivMed.2022.v41.104-113
CrossRef - van Baak MA, Pramono A, Battista F, et al. Effect of different types of regular exercise on physical fitness in adults with overweight or obesity: Systematic review and meta‐analyses. Obes Rev. 2021;22(S4). doi:10.1111/obr.13239
CrossRef - de Lucia C, Komici K, Borghetti G, et al. microRNA in Cardiovascular Aging and Age-Related Cardiovascular Diseases. Front Med. 2017;4. doi:10.3389/fmed.2017.00074
CrossRef - Ekowati AL, Milas Siswanto F. Epigenetic Alterations in Aging: A Brief Review. J Urban Heal Res. 2024;2(3):25-41. doi:10.25170/juhr.v2i3.5759
CrossRef - Kovanda A, Režen T, Rogelj B. MicroRNA in skeletal muscle development, growth, atrophy, and disease. Wiley Interdiscip Rev RNA. 2014;5(4):509-525. doi:10.1002/wrna.1227
CrossRef - Jung HJ, Lee KP, Kwon KS, Suh Y. MicroRNAs in Skeletal Muscle Aging: Current Issues and Perspectives. Journals Gerontol Ser A. 2019;74(7):1008-1014. doi:10.1093/gerona/gly207
CrossRef - Muljo SA, Kanellopoulou C, Aravind L. MicroRNA targeting in mammalian genomes: genes and mechanisms. WIREs Syst Biol Med. 2010;2(2):148-161. doi:10.1002/wsbm.53
CrossRef - Simeone I, Rubolino C, Noviello TMR, et al. Prediction and pan-cancer analysis of mammalian transcripts involved in target directed miRNA degradation. Nucleic Acids Res. 2022;50(4):2019-2035. doi:10.1093/nar/gkac057
CrossRef - Kreis, Louwen, Yuan. The Multifaceted p21 (Cip1/Waf1/CDKN1A) in Cell Differentiation, Migration and Cancer Therapy. Cancers (Basel). 2019;11(9):1220. doi:10.3390/cancers11091220
CrossRef - Handayani MDN, Sadewa AH, Farmawati A, Rochmah W. Deletion Polymorphism of Angiotensin-Converting Enzyme Gene Is Associated with Low Muscle Mass in Elderly People in Jakarta, Indonesia. Kobe J Med Sci. 2018;64(3):E119-E125. http://www.ncbi.nlm.nih.gov/pubmed/30666042
- Moraghebi M, Negahi AA, Bazireh H, et al. The Analysis of SNPs’ Function in miR-21 and miR146a/b in Multiple Sclerosis and Active Lesions: An In Silico Study. Bioinform Biol Insights. 2022;16:117793222211163. doi:10.1177/11779322221116322
CrossRef - Kim SH, Jeong JB, Kang J, et al. Association between sarcopenia level and metabolic syndrome. Mogi M, ed. PLoS One. 2021;16(3):e0248856. doi:10.1371/journal.pone.0248856
CrossRef - Wiedmer P, Jung T, Castro JP, et al. Sarcopenia – Molecular mechanisms and open questions. Ageing Res Rev. 2021;65:101200. doi:10.1016/j.arr.2020.101200
CrossRef - McGregor RA, Poppitt SD, Cameron-Smith D. Role of microRNAs in the age-related changes in skeletal muscle and diet or exercise interventions to promote healthy aging in humans. Ageing Res Rev. 2014;17:25-33. doi:10.1016/j.arr.2014.05.001
CrossRef - Wohlgemuth SE, Seo AY, Marzetti E, Lees HA, Leeuwenburgh C. Skeletal muscle autophagy and apoptosis during aging: Effects of calorie restriction and life-long exercise. Exp Gerontol. 2010;45(2):138-148. doi:10.1016/j.exger.2009.11.002
CrossRef - Zhao X, Wang Y, Sun X. The functions of microRNA-208 in the heart. Diabetes Res Clin Pract. 2020;160:108004. doi:10.1016/j.diabres.2020.108004
CrossRef - Mekala N, Kurdys J, Vicenzi AP, et al. MiR 208a Regulates Mitochondrial Biogenesis in Metabolically Challenged Cardiomyocytes. Cells. 2021;10(11):3152. doi:10.3390/cells10113152
CrossRef - Dutto I, Tillhon M, Cazzalini O, Stivala LA, Prosperi E. Biology of the cell cycle inhibitor p21CDKN1A: molecular mechanisms and relevance in chemical toxicology. Arch Toxicol. 2015;89(2):155-178. doi:10.1007/s00204-014-1430-4
CrossRef - Perez K, Ciotlos S, McGirr J, et al. Single nuclei profiling identifies cell specific markers of skeletal muscle aging, frailty, and senescence. Aging (Albany NY). Published online December 13, 2022. doi:10.18632/aging.204435
CrossRef - Tang Y, Cui Y, Li Z, et al. Radiation-induced miR-208a increases the proliferation and radioresistance by targeting p21 in human lung cancer cells. J Exp Clin Cancer Res. 2016;35(1):7. doi:10.1186/s13046-016-0285-3
CrossRef - Guo H, Xu Y, Fu Q. Curcumin inhibits growth of prostate carcinoma via miR-208-mediated CDKN1A activation. Tumor Biol. 2015;36(11):8511-8517. doi:10.1007/s13277-015-3592-y
CrossRef - Wang J, Song C, Cao X, et al. MiR‐208b regulates cell cycle and promotes skeletal muscle cell proliferation by targeting CDKN1A. J Cell Physiol. 2019;234(4):3720-3729. doi:10.1002/jcp.27146
CrossRef - Sun G, Yan J, Noltner K, et al. SNPs in human miRNA genes affect biogenesis and function. RNA. 2009;15(9):1640-1651. doi:10.1261/rna.1560209
CrossRef - Hindorff LA, Sethupathy P, Junkins HA, et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci. 2009;106(23):9362-9367. doi:10.1073/pnas.0903103106
CrossRef - Moraghebi M, Maleki R, Ahmadi M, Negahi AA, Abbasi H, Mousavi P. In silico Analysis of Polymorphisms in microRNAs Deregulated in Alzheimer Disease. Front Neurosci. 2021;15. doi:10.3389/fnins.2021.631852
CrossRef - Yang Y, Fan X, Yan J, et al. A comprehensive epigenome atlas reveals DNA methylation regulating skeletal muscle development. Nucleic Acids Res. 2021;49(3):1313-1329. doi:10.1093/nar/gkaa1203
CrossRef - Cui J, Liu J, Fan L, et al. A zinc finger family protein, ZNF263, promotes hepatocellular carcinoma resistance to apoptosis via activation of ER stress-dependent autophagy. Transl Oncol. 2020;13(12):100851. doi:10.1016/j.tranon.2020.100851
CrossRef - Kurachi M, Barnitz RA, Yosef N, et al. The transcription factor BATF operates as an essential differentiation checkpoint in early effector CD8+ T cells. Nat Immunol. 2014;15(4):373-383. doi:10.1038/ni.2834
CrossRef
Abbreviations
ADRA2A: Adrenoceptor Alpha 2A,
BATF: Basic Leucine Zipper Transcription Factor,
CDKN1A: Cyclin-dependent kinase inhibitor 1A,
ChIP: Chromatin immunoprecipitation,
DAVID: Database for annotation, visualization, and integrated discovery,
ENCORI: encyclopaedia of RNA interactomes,
FC: Fold change,
FST: Follistatin,
GEO: Gene expression omnibus,
GO: Gene ontology,
GTEx: Genotype-tissue expression,
KEGG: Kyoto encyclopedia of genes and genomes,
MFE: Minimal free energy,
miRNA: Micro-RNA,
mRNA: Messenger RNA,
PMW: Position weight matrix,
PPI: Protein-protein interaction,
SNP: Single nucleotide polymorphism,
STRING: Search tool for the retrieval of interacting genes,
TDMD: Target-directed miRNA degradation,
TSS: Transcription start site,
UTR: Untranslated region,
ZNF263: Zinc finger protein 263








