Manuscript accepted on :29-04-2025
Published online on: 20-05-2025
Plagiarism Check: Yes
Reviewed by: Dr. Abidin Çalişkan
Second Review by: Dr. Heamn Noori Abduljabbar
Final Approval by: Dr. Anton R Keslav
Kamal Upreti1* , Jossy George1
, Jossy George1 and Khushboo Malik2*
and Khushboo Malik2*
1Department of Computer Science and Engineering, Christ University, Delhi NCR, Ghaziabad, Uttar Pradesh, India.
2Department of Law, Christ University, Delhi NCR, Ghaziabad, Uttar Pradesh, India.
Corresponding Author E-mail: khushboo.usar@gmail.com
DOI : https://dx.doi.org/10.13005/bpj/3167
Abstract
Early and accurate classification of brain tumors plays a pivotal role in clinical decision-making and treatment planning. Manual methods are time-intensive and prone to variability, creating a need for robust automated solutions. This study aims to classify brain tumors from MRI scans using artificial intelligence techniques, specifically Logistic Regression (LR) and Support Vector Machines (SVM) with Radial Basis Function (RBF) kernels. The dataset, sourced from The Cancer Imaging Archive (TCIA), includes four classes: Meningioma, Glioma, Hypothalamic tumor, and No tumor. Preprocessing involved dimensionality reduction using Principal Component Analysis (PCA) to retain dominant features. Models were trained on an 80:20 train-test split, with LR achieving 99.83% training and 78.91% testing accuracy, while SVM performed better with 93.85% training and 81.88% testing accuracy. Error analysis revealed 104 misclassified samples, primarily due to structural similarity among tumor types. The findings suggest that SVM offers superior classification performance, and the study recommends further enhancement through deep learning models like Convolutional Neural Networks (CNNs) for improved diagnostic accuracy.
Keywords
Classification; Diagnosis; Early Detection; Machine Learning; MRI; Neuro-images
Download this article as:| Copy the following to cite this article: Upreti K, George J, Malik K. Automated Brain Tumor Segmentation in MRI Using AI for Improved Neurodiagnostics. Biomed Pharmacol J 2025;18(2). |
| Copy the following to cite this URL: Upreti K, George J, Malik K. Automated Brain Tumor Segmentation in MRI Using AI for Improved Neurodiagnostics. Biomed Pharmacol J 2025;18(2). Available from: https://bit.ly/3Zp1KqY |
Introduction
Any aberrant cell development in the brain, whether benign (non-malignant) or malignant (cancerous), is called a brain tumor. There are two types in it: primary, which starts in the brain, and secondary, which comes from other parts of the body. Headaches, seizures, loss of memory, vision loss, and loss of coordination are some of the symptoms.1 The cause is unknown, but inherited genetic mutations, radiation exposure, and family history can raise the risk. Diagnosis is done through MRI, CT scans, and occasionally a biopsy.2 Treatment can be surgery, radiation, chemotherapy, or targeted therapy. Early detection enhances outcomes, but prognosis varies with tumor type and stage.3 Medical research is raising the probability of survival and treatment. Brain tumors are categorized according to their origin, rate of growth, and malignancy also shown in Figure 1. Tumors of this kind can be categorized into primary, meaning they originate in the brain, or secondary, or metastatic, meaning they spread from other parts of the body. Primary tumors fall into one of two categories: malign (cancerous) or benign (non-cancerous).4 Common benign tumors are meningiomas, which are from the meninges, and pituitary adenomas, which are affecting hormone secretion. Malignant tumors are gliomas, which arise from glial cells, with subtypes like astrocytomas, oligodendrogliomas, and glioblastomas—the most lethal type.5 Medulloblastomas, frequent in children, and ependymomas, which are acting on cerebrospinal fluid pathways, are other types. Secondary tumors typically arise from cancer such as lung, breast, or melanoma that metastasizes to the brain.6 Classification is crucial for the decision of treatment and prognosis. Classical techniques for identifying brain tumors are predicated on imaging procedures and clinical assessments.7 The most common procedure is MRI, or magnetic resonance imaging that gives precise pictures of brain structures and determines abnormalities.
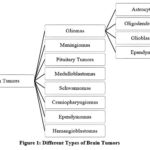 |
Figure 1: Different Types of Brain Tumors |
Computed Tomography (CT) scans employ the use of X-rays to create cross-sectional images that are commonly employed in emergency situations to get rapid assessment.8 Electroencephalography (EEG) is helpful in detecting abnormal brain activity that is causing seizures from tumors. Imaging of blood vessels within the brain, particularly those tumors influencing blood flow, is done through angiography. Positron Emission Tomography (PET) scan can trace the metabolic rate in an effort to identify benign or malignant tumors.9 The biopsy, in which a piece of the tumor tissue is removed and studied microscopically, is the only test for diagnosing malignancy. These classic techniques are vital in early diagnosis and in directing treatment.10
Early diagnosis of tumors for brain tumors is important to enhance the treatment outcome, as well as survival, in patients. Early diagnosis provides an opportunity for early medical therapy, avoiding progression and reducing complications.11 Less invasive procedures become possible with early diagnosis, enhancing the quality of life of the patient and avoiding neurological damage.12 Delayed detection results in tumor growth, which results in severe symptoms like seizures, memory loss, and loss of motor functions. Advanced imaging and AI-based diagnosis enhance the accuracy and speed of tumor detection in the initial stages.13 Early detection also assists in choosing the best treatment protocol, for example, surgery, radiation, or chemotherapy, before the metastasis of the tumor.14 Public awareness, frequent screening, and technological progress are significant in early detection and improved outcome.15 Machine learning is important in brain tumor detection by accuracy, speed, and autonomy in diagnosis in the healthcare sector. Progressive algorithms like Convolution Neural Networks (CNNs) and Deep Learning algorithms utilize MRI and CT scan images for the detection of tumors with great accuracy. Random Forest, SVMs (support vector machines) and artificial neural networks (ANNs) are generally employed to segment and classify tumors.16 These models help differentiate between benign and malignant tumors, minimizing the use of invasive tests like biopsies. Machine learning also supports the automated extraction of features, which enhances early detection and minimizes human error. Furthermore, predictive models help clinicians in evaluating patterns of tumor growth and response to therapy.17 The integration of AI with medical imaging not only improves the accuracy of diagnosis but also facilitates personalized planning for treatment for better patient outcomes. Machine learning improves the detection of brain tumors by enhanced accuracy, speed, and automation of diagnosis. Methods such as Deep Learning and CNNs interpret medical images with high precision, minimizing human error.18 Artificial intelligence-based models allow for automatic segmentation and classification of tumors, separating benign from malignant tumors with high efficiency.19 Predictive analytics allows tumor growth and response to therapy to be assessed, allowing for personalized treatment. Machine learning also reduces the necessity for invasive biopsies, ensuring that diagnosis is safer. Through ongoing learning from large data sets, the models improve over time, yielding better detection, prognosis, and medical decision-making.20
Research Objectives
The primary goal of the current study is to create a standalone and high-speed classification model that identifies brain tumors from machine learning and MRI scans. It should, per machine learning models, improve accuracy in diagnosis, minimize human involvement, and contribute a computationally effective way towards tumor classification, as indicated by machine learning models. Another of the primary goals is to ascertain the performance rate of machine learning models in successful brain tumor classification. The comparison offers the precise information concerning the strength and weaknesses of the models and facilitates identification of the best classifier to use for medical images. In addition to this, PCA is applied for reducing dimensionality as well as extracting features. The approach guarantees maximum computational effectiveness by removing redundant features without causing loss of important information such that it increases the accuracy of the model as well as minimizes processing time. Error analysis, which is another major part of this research, examines misclassification error and tries to identify reasons behind the same. Identification of false positives, false negatives, and inter-class misclassification has the advantage of fine-tuning the classification technique and revealing scope for change. Finally, the research aims to suggest future research avenues for improving the accuracy of classification using the examination of deep architectures of learning, advanced feature extraction algorithms, and ensemble learning approaches for better detection of tumors.
Paper Structure
There are five main sections to this study article. Part One (Introduction & Background) discusses the significance of brain tumor classification, drawbacks of manual diagnosis, and the promise of machine learning in increasing accuracy. Section 2 (Methodology) explains data preprocessing, which involves dataset acquisition, image normalization, PCA-based dimensionality reduction, model training, and evaluation methods. Section 3 (Performance Evaluation) discusses the comparative analysis between Logistic Regression and SVM with RBF Kernel in terms of training and testing accuracy results. Section 4 (Error Analysis) discusses misclassified samples, noting typical errors like erroneous positive and negative results, and discusses reasons for misclassification. Section 5 (Conclusion & Future Work) outlines major findings, addresses model limitations, and proposes future enhancements, such as deep learning methodologies such as CNNs and ensemble methods in order to improve the accuracy of classification.
Contribution of Study
This study suggests developing a categorization system based on machine learning in order to distinguish between brain tumor types based on MRI scans. The study contrasts the performance of machine learning models to determine the best method. It also assesses the effect of Principal Component Analysis (PCA) on feature extraction and performs a thorough error analysis to determine typical misclassifications and areas for improvement. This research advances the science of brain tumor classification with machine learning by developing a systematic automatic system for classification from MRI scans. The research applies machine learning methods, including Support Vector Machines (SVM) and Logistic Regression (LR) with RBF Kernel, to distinguish between various types of brain tumors. Through the use of machine learning, this research proposes to assist radiologists and medical professionals to provide accurate and quick diagnoses. A crucial part of this study entails comparing LR and SVM, reflecting data on reporting the effectiveness of traditional machine learning methods to identify tumors works. We also used Principal Component Analysis (PCA) to compactly trim superfluous MRI image dimensions preserving dominant features, to guarantee feature selection and computation efficiency with optimal features. The second significant contribution is rigorous error analysis, which identifies the dominant patterns of misclassification and their root causes. By studying the misclassified samples, the study provides further insight into false positives, false negatives, and inter-class confusion, and the door to the potential development of future models. Besides, based on the results, future studies’ recommendations from the study suggest using deep models such as Convolution Neural Networks (CNNs) in the aim of improving classification performance and robustness.
The three primary approaches to segmenting and categorizing brain tumors—region growth, shallow machine learning, and deep learning—are comprehensively explored in.21 It mentions the hassle and laborious task of manually segmenting brain tumors from MRI scans and the possible advantages of automation, including safer biopsies and non-invasive diagnosis. An automatic approach for precise brain tumor segmentation from MRI scans is outlined by.22 Pre-processing, image thresholding, and segmentation are all included in the system. Based on the application of Harvard datasets, this study compares Chan-vese and level set (LSM) approaches. The findings indicate that Chan-Vase is more effective in detecting tumors compared to LSM, while KIFCM is more effective compared to other clustering algorithms. This model outperforms current methods in brain tumor detection with less information loss and distance error. By introducing a deep learning architecture combining self-attention mechanisms and U-Net architecture,.23 try to increase the precision of brain tumor detection in neuro-oncology. The research compares and contrasts existing deep learning techniques, identifying the efficacy of the U-Net structure in detecting some features. The proposed approach sets a new benchmark in medical image segmentation by outperforming current breakthroughs in brain tumor segmentation performance.24 analyses detection of brain cancers with the assistance of Magnetic Resonance Imaging (MR). It proposes several methods depending on statistical image processing and artificial intelligence. Brain tumor morphology, available data sets, augmentation methods, component separation, and classification between Deep Learning, Transfer Learning, and Machine Learning models are also discussed in the study. Besides, it collects necessary information on tumor detection and advancement. Image segmentation, which splits digital images into small pieces to analyze, is an important part of image processing.25 reviews existing segmentation techniques and proposes new ones to facilitate improved information analysis and retrieval but eliminate their limitations.26 offer a lightweight U-Net solution for brain tumor segmentation from magnetic resonance imaging (MRI). Segmentation in real time is feasible with the lightweight U-Net without needing extensive data. It illustrates the use of three view planes for easier segmentation and is superior to standard benchmark approaches, providing promising results on the BITE dataset.27 present a model based on deep learning to handle challenges in brain tumor segmentation techniques. The model consists of three channels: five-layered segmentation capsule, six-layered transfer-learning capsule, and patch extraction. Two dropout layers and five segmentation channel levels with dropout layers are blended in the proposed 11-layered deep capsule network. High dice score coefficients (0.80, 0.76, 0.76) for whole tumor, active tumor, and core tumor were achieved by the model once it was able to learn on over 150 images successfully. In order to appropriately segment brain tumors without modalities, .28 suggests an Edge-aware Discriminative Feature Fusion Based Transformer U-Net (EA-DFFTU-Net). The pre-processing of the MRI input data comes before a ResNet-50 encoder for feature extraction. For extracting multiscale features, the Discriminative Feature Fusion based Transformer U-Net (DFFTU-Net) uses edge attention representations from the Edge Feature Module (EFM).Performance evaluation and comparison of the proposed EA-DFFTU-Net with existing methods were performed. Applying BraTS 2018 data,.29 compares various deep learning architectures such as U-Net, PSPNet, DeepLabV3+, and ResNet50. The results show that the 3D U-Net model performed best, with DSC of 0.90, HD of 10.69mm, and AVD of 11.15%; the PSPNet model also performed similarly, with DSC of 0.89, HD of 11.37mm, and AVD of 12.24%; the DeepLabV3+ and ResNet50 models performed less well, with DSCs of 0.85 and 0.83, respectively. The most lethal form of brain cancer, gliomas are most common. In order to increase brain tumor segmentation accuracy, scientists have created semi-automatic and completely automatic algorithms that use precise handmade feature-based or data-driven methodologies together with machine learning (ML) pipelines.30 In brain tumor detection, training of mr images created by heat map technique with cnn models-extraction of type- based activation sets and selection of best features by mrmr method .31 to precisely segment brain tumors in MRI images. The model emphasizes interdependencies by distributing and aggregating long-range information over a wider geographical area. Comparing with the existing methods, experimental results, the performance of segmentation is better. Healthcare workers and systems may be able to save time through the use of this model’s potential as an effective tool for medical image processing and performance improvement. An optimization-based classifier for MRI-based brain tumor classification is introduced by.32 The approach involves segmenting the output through a U-Net model, tumor classification through the Poor Bird Swarm Optimization algorithm (PRBSA), and pre-processing MRI through filtering and ROI extraction methods. Both the Bird Swarm Algorithm and Poor and Rich Optimization are employed to train the U-Net. The PRBSA-based U-Net achieved high sensitivity, specificity, and accuracy with a 94% accuracy rate.
Brain tumors present one of the most complex diagnostic challenges in neuro-oncology due to their morphological diversity, overlapping visual features, and the criticality of early detection. While recent advancements in deep learning have significantly enhanced segmentation and classification accuracy in medical imaging, such methods often depend on vast annotated datasets, high-end computational resources, and black-box decision-making processes that limit transparency and clinical adoption. This study addresses this critical gap by investigating whether traditional machine learning methods, particularly Support Vector Machines (SVM) with RBF kernels, can offer a reliable, interpretable, and resource-efficient solution for MRI-based brain tumor classification. The main objective is to evaluate the performance of classical models in a multi-class classification setting using real-world medical data from TCIA. By comparing SVM against Logistic Regression (LR), the study further aims to assess how nonlinear classifiers perform in the presence of structurally similar tumor types. The motivation is to establish a computationally lightweight benchmark model that maintains diagnostic reliability while remaining suitable for practical deployment in diverse healthcare environments.
Materials and Methods
This study proposes an MRI-based machine learning approach for brain tumor categorization, as illustrated in Figure 2. MRI data collection is the first step throughout this process, followed by data preprocessing that involves scaling, standardization, or grayscale conversion. Dimensionality reduction utilizing Principal Component Analysis (PCA) will be used to extract features, and the data will be split into 20% test and 80% training.
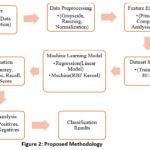 |
Figure 2: Proposed Methodology |
For classification, Support Vector Machines (SVM) and Logistic Regression (LR) using RBF Kernel, both optimized with Gradient Descent and Sequential Minimal Optimization (SMO), respectively, will be used. Accuracy, precision, and recall will be the metrics to evaluate the models, followed by error analysis to determine misclassifications. This approach is to build a strong and computerized system for detecting brain tumors, assisting radiologists in accurate diagnosis.
Data Collection and Preprocessing
Brain MRI scans that have had FLAIR abnormality masks manually segmented are among the data used in this study. The images, which were obtained from The Cancer Imaging Archive (TCIA), cover 110 individuals through the lower-grade tumor database The Cancer Genome Atlas (TCGA) on Kaggle. Each MRI scan includes FLAIR (fluid-attenuated inversion recovery) sequences and genomic cluster information. This data.csv document contains tumor genomic clusters and related patient data. In order to segment brain tumors automatically, the dataset was categorized into four distinct tumor types as shown in Figure 3:
No Tumor (Healthy brain scans)
Meningioma Tumor
Glioma Tumor
Pituitary Tumor
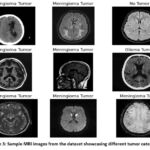 |
Figure 3: Sample MRI images from the dataset showcasing different tumor categories |
The images were obtained from a ZIP archive, which was extracted into separate folders for Training and Testing subsets. The dataset was structured systematically by mapping class labels to their respective MRI scans, ensuring proper labeling of each image for model training and evaluation. To ensure consistency and optimal model performance, several preprocessing steps were applied to the MRI images as shown in Figure 4: All MRI scans were loaded using OpenCV in grayscale mode, as color information is not essential for brain tumor segmentation. Loading the images in grayscale reduces computational complexity and allows the model to focus on the intensity variations that define tumor regions. Since the original MRI scans varied in resolution, Every image was scaled to a fixed 200×200 pixel size.. This step ensured uniformity across the dataset, making it easier for the model to process images without variations in input size. The fixed resolution also optimized memory usage and improved computational efficiency during training.
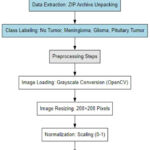 |
Figure 4: Approach for collecting and preprocessing data |
In order to improve machine learning algorithms’ performance, pixel values were standardized to fall between 0 and 1. This was accomplished by dividing the luminance of each pixel by 255, which standardizes the intensity values across different MRI scans. Normalization helps in:
Reducing the impact of varying intensity levels across different MRI scans.
Improving model convergence during training by maintaining a consistent input range.
Preventing numerical instability when performing calculations in deep learning models.
Each MRI image was flattened into a 1D vector before being used for classification. The original 200×200 pixel images were then mapped to feature vectors of size 40,000 (200×200). This mapping allowed machine learning algorithms to process the images as structured input data. Although this mapping discards spatial information, it simplifies the input format for conventional classification algorithms, such as Support Vector Machines (SVM ) or logistic regression.
Splitting and Dimensionality Reduction
The dataset was methodically divided into training and test subsets to provide a reliable training and assessment procedure. 80% of the data set was divided into training and 20% into testing using the train test splitting method from the sickest-learn package. This division technique allows the model to identify patterns in the vast majority of the data while preserving a distinct subset for objective performance evaluation. To provide stability across model run differences, a random seed value was employed during splitting, leaving train outcomes unaffected.
After splitting the dataset, Principal Component Analysis (PCA) was applied to solve the high-dimensional data problem. PCA was employed for dimensionality reduction since it preserves maximum variance and decreases computational complexity drastically. As MRI images are of high dimension, PCA can eliminate redundant and correlated features to improve model efficiency and prevent over fitting. Unlike t-SNE or UMAP, which are primarily visualization techniques, PCA gives a linear transformation that is still interpretable and scalable. It is more computationally efficient than Autoencoders and guarantees improved generalization for machine learning techniques include SVM (Support Vector Machine) and Logistic Regression (LR). This makes PCA the best option for precise and efficient brain tumor classification Because the MRI images were being resized into 200×200 pixels, the feature vector resulting from resizing had dimensions per image of 40,000. High-dimensional feature space resulting in this scenario can result in over fitting, higher computational overhead, and performance of the model being reduced. To minimize the issues, PCA was used for compressing data without losing 98% data variance. This process made it possible to retain the required information and delete redundant features and, in so doing, achieved improved computational complexity and generalization. PCA transform dramatically minimized feature count at maintaining the most notable features of the tumor differentiation. This benefited, in a good way, model kinds like Support Vector Machines (SVM) and Logistic Regression, whose classification has been aided by structured reduced-dimensional input. By applying dimensionality reduction, the model would be capable of improving performance, decreasing training time, and decreasing storage space, thus making the brain tumor classification task more efficient and scalable.
Model Selection and Training
LR was employed as the baseline while SVM with RBF Kernel to classify brain tumors based on MRI scans. It was divided between training and testing 80-20% in order to determine a fair evaluation of the model. Gradient Descent Optimization for LR and SMO for SVM were used. In LR, soft ax activation was turned on to facilitate multiclass classification, and SVM used an RBF kernel to provide better performance with non-linearly separable data. SVM was hyper parameterized using tuning of C (regularization) and gamma (kernel coefficient). Even though LR is computation efficient and interpretable, it suffers with non-linear patterns, whereas SVM performs extremely well with complex classifications but at the expense of computational effort. The results reveal SVM with RBF Kernel to be the top-performing model for classifying brain tumors in MRI images.
Results
The algorithms used for classifying brain tumors were evaluated for accuracy. Logical Regression (LR) and Support Vector Machine (SVM) using RBF Kernels were both of the models that were compared, as shown in Table 1. The model that used Logistic Regression recorded a training accuracy of 99.83%, indicating that it was capable of fitting the training dataset exceptionally well. However, its test accuracy was 78.91%, which proves that while it performed well on the training set, it was not able to generalize so much on unseen samples. The variation is a reflection of an overfitting issue because Logistic Regression needs linear separability and could not be able to detect the intricate patterns in MRI images. On the contrary, SVM classifier with RBF Kernel exhibited 93.85% accuracy on the training set and 81.88% accuracy on the test set. Although its accuracy in training was less than Logistic Regression, the better testing accuracy of SVM reflects better generalizability to out-of-sample data. This is to say, the SVM model is better able to handle MRI scan non-linearity and high dimensionality and therefore is the better choice in brain tumor classification. According to the findings, SVM with an RBF kernel performed better than logistic regression in testing accuracy and hence is a better model for this classification task. However, SVM is computationally more expensive and requires hyperparameter tuning, while Logistic Regression is simpler and faster but not as good on complex, high-dimensional data.
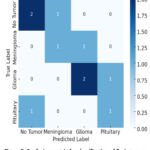 |
Figure 5: Confusion matrix for classification of Brain tumor |
The matrix of uncertainty heatmap in Figure 5 graphically displays the model’s classification performance by juxtaposing true tumor labels and predicted labels. Correctly classified instances are shown along the diagonal values, and off-diagonal values represent misclassifications. Misclassification between Meningioma and Glioma is seen in the model, probably due to the latter’s MRI similarity with the former, and also false positives under the “No Tumor” category, potentially resulting in unjustified medical treatments. False negatives are also dangerous as they don’t detect real tumor cases. From the Figure 6, Accuracy for SVM (81.88%) is greater than that of Logistic Regression (78.91%), suggesting SVM performs better overall. Precision, or the accuracy of positive classifications, is also improved in SVM (80.5%) over Logistic Regression (76.3%), and so SVM suppresses false positives better. The F1-Score is a precise or memory harmonic mean, is once greater for SVM (79.7%) compared to Logistic Regression (75.4%), indicating that SVM has both better precision-recall trade-offs and is a more consistent classifier here. This enhanced performance is primarily due to SVM’s ability to handle nonlinear decision boundaries using the RBF kernel, making it more effective for complex, high-dimensional MRI data. In contrast, LR assumes linear separability and is less capable of capturing intricate patterns, especially when tumor classes overlap in feature space. These results highlight the suitability of SVM for multi-class medical image classification tasks where subtle variations between categories must be captured accurately.
Table 1: Performance Evaluation of Models
| Models | Accuracy | Precision | Recall | F1-Score |
| LR | 78.91 | 76.3 | 74.5 | 75.4 |
| SVM | 81.88 | 80.5 | 78.9 | 79.7 |
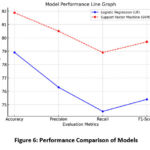 |
Figure 6: Performance Comparison of Models |
Error Analysis
Error analysis serves as crucial stage in measuring a machine learning model’s performance, especially in medical image tasks where accuracy is paramount. Misclassified samples in this research were determined through NumPy, which enabled us to investigate instances where the predictions of the model were not consistent with the true tumor labels. This discussion presents information about the type of errors, why they are occurring, and the ways in which model performance can be improved. For the detection of the misclassified samples, the actual labels and predicted labels of the test dataset were contrasted. Misclassified samples were retrieved with the help of the function of NumPy, which identified the indices where wrong predictions were made also shown in Figure 7. It illustrates MRI samples of pituitary and glioma tumors that display significant visual similarity, particularly in structure and location. This resemblance often causes confusion for the classification model, leading to mislabeling between the two classes. Such overlap highlights the limitation of traditional machine learning models in distinguishing closely related tumor types and supports the need for advanced feature extraction methods in future research. 104 misclassified samples were discovered, indicating a few inconsistencies in the classification ability of the model. These mistakes highlight possible issues like over fitting, imbalance of data, or the inability to differentiate between tumor types that appear visually similar on MRI scans.
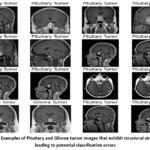 |
Figure 7: Examples of Pituitary and Glioma tumor images that exhibit structural similarities, leading to potential classification errors |
Discussion
On close inspection, some misclassifications revealed specific patterns. There was considerable confusion between Glioma and Meningioma tumors, presumably because both had similar structures on MRI scans. Likewise, some No Tumor images were mistakenly classified as images containing tumor, resulting in false positives—a disquieting occurrence in clinical diagnostics, since it might lead to unwarranted medical interventions. Table 2 provides a breakdown of the classification errors observed during the testing phase of the proposed SVM-based model. A total of 104 misclassified samples were identified, highlighting several underlying challenges. The most common misclassification occurred between meningioma and glioma classes, attributed to their overlapping morphological characteristics in MRI scans. False positives, where healthy (non-tumor) images were classified as tumor cases, raise concerns over unnecessary diagnostic escalation. Conversely, false negatives—tumor cases misclassified as “No Tumor”—pose a significant clinical risk due to potential delays in treatment. The table also indicates that tumor types with fewer examples in the training data exhibited disproportionately high misclassification rates, suggesting the presence of class imbalance, a known limitation in classical ML models. Potential causes include model overfitting, insufficient feature extraction, and similarity in pixel-level features among different tumor classes. To mitigate these issues, the table suggests adopting data augmentation, CNN-based architectures, hyperparameter tuning, and ensemble techniques as future enhancements for improving robustness and diagnostic safety.
Table 2: Error Analysis in Brain Tumor Classification
| Error Category | Details |
| Total Misclassified Samples | 104 misclassified samples identified using NumPy. |
| Common Misclassification | Confusion between Meningioma and Glioma due to structural similarities in MRI scans. |
| False Positives | Some non-tumor cases were incorrectly classified as tumor cases, leading to unnecessary alerts. |
| False Negatives | Some tumor cases were misclassified as “No Tumor,” increasing the risk of missed diagnoses. |
| Impact of Data Imbalance | Tumor types with fewer training samples showed a higher rate of misclassification. |
| Potential Causes | Overfitting, data imbalance, similar MRI features across tumor types, and limitations in feature extraction. |
| Possible Solutions | Implement deep learning models (CNNs), use data augmentation techniques, apply hyperparameter tuning, experiment with ensemble learning, and enhance feature extraction methods. |
The presence of these misclassification errors has numerous real-world implications. False negatives may cause undue delay in medical treatment, deteriorating the condition of a patient because of an undetected tumor. False positives may cause undue stress, further diagnostic tests, or even inappropriate treatments. These issues highlight the importance of accuracy in medical AI models so that errors must be minimized for model improvement.
In order to counter these mistakes and make the classification accuracy better, many approaches can be employed. The incorporation of Models for deep learning such as Convolution Neural Networks (CNNs) could enhance feature learning and the precision of classification. Data augmentation techniques like rotation, scaling, and changing contrast could be applied in order to raise dataset diversity and enable the model to generalize. The second method involves hyper parameter tuning of SVM and Logistic Regression models specifically to optimize their classification boundaries. Ensemble learning techniques, in which various classifiers like SVM, Neural Networks, and Decision Trees are employed with aggregation, can potentially enhance authenticity of prediction. In addition, more sophisticated feature extraction methods such as edge enhancement and texture analysis could enable the model to distinguish between tumor types more accurately by capturing slight differences. The 104 missing samples also serve to reflect the weaknesses of the existing model, especially in the differentiation of evidently similar tumors. While SVM with RBF Kernel was better than Logistic Regression, it still had mistaken which need to be fulfilled for improved performance. By means of implementing deep learning, adjusting model parameters, and revising dataset preprocessing techniques, future work can make such computerized brain tumor classification models more accurate, ultimately leading to better medical diagnoses. As shown in Table 3, while several recent studies report high accuracy using complex deep learning models, the present study offers a unique balance of performance, computational efficiency, and interpretability. Our model, based on Support Vector Machine (SVM) with RBF kernel and Principal Component Analysis (PCA), achieves a competitive accuracy of 81.88% on the real-world TCIA dataset. Unlike deep learning approaches that require extensive annotated data and high-end hardware, the proposed method is lightweight and deployable in low-resource clinical environments. Moreover, it successfully handles multi-class classification and provides transparent error analysis, making it not only efficient but also practical. This justifies its value as a strong baseline in brain tumor classification research, with potential for future enhancement using deep learning techniques.
Table 3: Comparative Analysis of MRI-Based Brain Tumor Classification and Segmentation Methods
| Study (Author & Year) | Methodology | Dataset | Model Type | Accuracy / Dice Score (%) | Remarks |
| 24 | Binary classification using SVM | Private MRI | Classical ML (SVM) | 80.2 | SVM applied on limited binary dataset |
| 22 | Chan-Vese vs. LSM segmentation | Harvard Dataset | Image Segmentation | ~90.0 | Pre-processing based segmentation study |
| 30 | CSPCNN + MRIPCNN dual CNN model | BraTS | Deep Learning (CNN) | 87.0 (DSC) | Specialized CNN pathways; strong performance |
| 31 | 3D UNet + Transformer | BraTS | Deep Learning (Hybrid) | 91.0 (DSC) | Exploits long-range spatial relationships |
| 32 | PRBSA-optimized U-Net | Private MRI | Deep Learning + Metaopt | 94.0 | Uses Poor Bird Swarm Optimization for training |
| Present Study | SVM (RBF Kernel) with PCA | TCIA | Classical ML (SVM) | 81.88 | Multi-class tumor classification; strong baseline performance |
The findings of this study contribute to the growing body of research focused on resource-aware AI solutions in medical diagnostics. The proposed SVM model, despite being a classical machine learning technique, demonstrates competitive accuracy and balanced performance metrics across four tumor classes, including glioma, meningioma, hypothalamic tumors, and non-tumor cases. This underscores its applicability in settings where deep learning solutions may be impractical due to infrastructure constraints or the unavailability of large, labeled datasets. Furthermore, the study highlights critical challenges in tumor classification, such as structural overlap and data imbalance, which remain relevant across both classical and modern AI paradigms. By identifying these limitations and proposing data-centric and architectural improvements, the work sets a strong foundation for future hybrid or deep learning extensions aimed at enhancing clinical decision support systems.
Conclusion
This study focused on four tumor classifications and assessed how well Support Vector Machines (SVM) and Logistic Regression (LR) classified brain tumor MRI datasets from The Cancer Imaging Archive (TCIA). According to the results, SVM performs better than LR when employing RBF kernels, with testing accuracy of 81.88% compared to 78.91%, respectively. Notwithstanding these encouraging results, the study points out many important drawbacks, most notably the difficulty of correctly categorizing tumors with overlapping structural characteristics and managing high-dimensional imaging data using conventional machine learning methods. Shallow models’ poor ability to derive abstract, hierarchical representations from complex medical images is the gap. Future studies should investigate deep learning architectures to overcome this, especially Convolutional Neural Networks (CNNs), which are better able to understand the textural and spatial information included in MRI data. Furthermore, multi-modal imaging inputs (e.g., integrating MRI with PET or CT), data augmentation, and ensemble learning techniques may enhance classification performance and clinical usefulness. In the end, integrating such sophisticated AI models into real-time diagnostic pipelines may improve precision, lessen the strain for radiologists, and facilitate neuro-oncology early intervention.
Acknowledgment
The authors wish to express their gratitude to the Centre for Research Projects (CRP) at CHRIST (Deemed to be University), Bangalore Central Campus, Bengaluru, India, for generously supporting this research with Seed Money for the academic year 2024-25 (Project Number CU: ORS: SM-24/95).
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
We have used images of tumor from public availably website kaggle.com – https://www.kaggle.com/code/abdallahwagih/brain-tumor-segmentation-unet-dice-coef-89-6/input
Author Contributions
- Kamal Upreti: Conceptualization, methodology, software development, data curation, model implementation, writing—original draft, and result analysis.
- Jossy George: Supervision, validation, funding acquisition, resources, and critical review of the manuscript.
- Khushboo Malik: Data preprocessing, experimental setup, performance evaluation, manuscript editing, and visualization.
Refrences
- Chen X, Zhang X, Xie H, et al. A bibliometric and visual analysis of artificial intelligence technologies-enhanced brain MRI research. Multimedia Tools and Applications. 2020;80(11):17335–63. https://doi.org/10.1007/s11042-020-09062-7
CrossRef - Gull S, Akbar S. Artificial Intelligence in Brain Tumor Detection through MRI Scans. In: CRC Press eBooks. 2021. p. 241–76. https://doi.org/10.1201/9781003097204-10
CrossRef - Gutman B, Shmilovitch AH, Aran D, Shelly S. Twenty-Five Years of AI in Neurology: The Journey of Predictive Medicine and Biological Breakthroughs (Preprint). JMIR Neurotechnology. 2024;3: e59556. https://doi.org/10.2196/59556
CrossRef - Al Bataineh AF, Nahar KM, Khafajeh H, Samara G, Alazaidah R, et al. Enhanced Magnetic Resonance Imaging-Based Brain Tumor Classification with a Hybrid Swin Transformer and ResNet50V2 Model. Applied Sciences. 2024;14(22):10154.
CrossRef - Muhammad SD, Kobti Z. An ensemble deep learning approach for enhanced classification of pituitary tumors. 2021 IEEE Symposium Series on Computational Intelligence (SSCI). 2023;427–32. https://doi.org/10.1109/ssci52147.2023.10371824
CrossRef - Góralczyk-Bińkowska A, Szmajda-Krygier D, Kozłowska E. The microbiota–gut–brain axis in psychiatric disorders. International journal of molecular sciences. 2022;23(19):11245.
CrossRef - Saeedi S, Rezayi S, Keshavarz H, R. Niakan Kalhori S. MRI-based brain tumor detection using convolutional deep learning methods and chosen machine learning techniques. BMC Medical Informatics and Decision Making. 2023;23(1):16.
CrossRef - Sultana OF, Bandaru M, Islam MA, Reddy PH. Unraveling the complexity of human brain: Structure, function in healthy and disease states. Ageing Research Reviews. 2024;100:102414.
CrossRef - Abidi N, Abidi Z, Hanrahan B, et al. Perspective Chapter: Future Impact of Artificial Intelligence on Medical subspecialties – Dermatology and Neurology. In: Artificial intelligence. 2024. https://doi.org/10.5772/intechopen.115279
CrossRef - Kazemivash, B., Suresh, P., Ye, D.H., Iraji, A., Liu, J., Plis, S., Kochunov, P. and Calhoun, V.D. st-DenseViT: A Weakly Supervised Spatiotemporal Vision Transformer for Dense Prediction of Dynamic Brain Networks. bioRxiv, 2024:2024-11.
CrossRef - Orouskhani M, Zhu C, Rostamian S, Zadeh FS, Shafiei M, Orouskhani Y. Alzheimer’s disease detection from structural MRI using conditional deep triplet network. Neuroscience Informatics. 2022;2(4):100066.
CrossRef - Bonnekoh LM, Seidenbecher S, Knigge K, et al. Long-term cortisol stress response in depression and comorbid anxiety is linked with reduced N-acetylaspartate in the anterior cingulate cortex. The World Journal of Biological Psychiatry. 2022;24(1):34–45. https://doi.org/10.1080/15622975.2022.2058084
CrossRef - Gull S, Akbar S. Artificial Intelligence in Brain Tumor Detection through MRI Scans. In: CRC Press eBooks. 2021. p. 241–76. https://doi.org/10.1201/9781003097204-10
CrossRef - Islam MK, Ali MS, Miah MS, Rahman MM, Alam MS, Hossain MA. Brain tumor detection in MR image using superpixels, principal component analysis and template based K-means clustering algorithm. Machine Learning with Applications. 2021;5:100044.
CrossRef - Al-Kadi OS, Al-Emaryeen RA, Al-Nahhas S, Almallahi IA, Braik R, Mahafza W. Empowering brain cancer diagnosis: harnessing artificial intelligence for advanced imaging insights. Reviews in the Neurosciences. 2024 Jun 25;35(4):399-419.
CrossRef - Zhu Z, He X, Qi G, Li Y, Cong B, Liu Y. Brain tumor segmentation based on the fusion of deep semantics and edge information in multimodal MRI. Information Fusion. 2023;91:376-87.
CrossRef - Ali S, Li J, Pei Y, Khurram R, Rehman KU, Mahmood T. A comprehensive survey on brain tumor diagnosis using deep learning and emerging hybrid techniques with multi-modal MR image. Archives of computational methods in engineering. 2022 Nov;29(7):4871-96.
CrossRef - Chattopadhyay A, Maitra M. MRI-based brain tumour image detection using CNN based deep learning method. Neuroscience informatics. 2022;2(4):100060.
CrossRef - Soomro TA, Zheng L, Afifi AJ, Ali A, Soomro S, Yin M, Gao J. Image segmentation for MR brain tumor detection using machine learning: a review. IEEE Reviews in Biomedical Engineering. 2022 Jun 23;16:70-90.
CrossRef - Zebari DA, Ibrahim DA, Zeebaree DQ et al. Systematic review of computing approaches for breast cancer detection based computer aided diagnosis using mammogram images. Applied Artificial Intelligence. 2021;35(15):2157-203.
CrossRef - Biratu ES, Schwenker F, Ayano YM, Debelee TG. A survey of brain tumor segmentation and classification algorithms. Journal of Imaging 2021;7(9):179. https://doi.org/10.3390/jimaging7090179
CrossRef - Babu Kr, Ksony N, Indira Nd, Prasad KV, Shameem S. An Effective Brain Tumor Detection from T1w MR Images Using Active Contour Segmentation Techniques. Journal of Physics Conference Series. 2021;1804(1):012174. https://doi.org/10.1088/1742-6596/1804/1/012174
CrossRef - Umarani CM, Gollagi SG, Allagi S, Sambrekar K, Ankali SB. Advancements in deep learning techniques for brain tumor segmentation: A survey. Informatics in Medicine Unlocked. 2024; 50:101576. https://doi.org/10.1016/j.imu.2024.101576
CrossRef - Solanki S, Singh UP, Chouhan SS, Jain S. Brain Tumor Detection and Classification Using Intelligence Techniques: An Overview. IEEE Access. 2023; 11:12870–86. https://doi.org/10.1109/access.2023.3242666
CrossRef - Sarma R and Gupta YK. A comparative study of new and existing segmentation techniques. IOP Conference Series Materials Science and Engineering. 2021;1022(1):012027. https://doi.org/10.1088/1757-899x/1022/1/012027
CrossRef - Walsh J, Othmani A, Jain M, Dev S. Using U-Net network for efficient brain tumor segmentation in MRI images. Healthcare Analytics. 2022; 2:100098. https://doi.org/10.1016/j.health.2022.100098
CrossRef - Sille R, Choudhury T, Chauhan P, Sharma D. A systematic approach for deep learning based brain tumor segmentation. Ingénierie Des Systèmes D Information. 2021;26(3):245–54. https://doi.org/10.18280/isi.260301
CrossRef - Jagadeesh B, Kumar GA. Brain tumor segmentation with missing MRI modalities using edge aware discriminative feature fusion based transformer U-net. Applied Soft Computing. 2024; 161:111709. https://doi.org/10.1016/j.asoc.2024.111709
CrossRef - Chauhan AS, Singh J, Kumar S, Saxena N, Gupta M, Verma P. Design and assessment of improved Convolutional Neural Network based brain tumor segmentation and classification system. Journal of Integrated Science and Technology. 2024;12(4). https://doi.org/10.62110/sciencein.jist.2024.v12.793
CrossRef - Ullah F, Nadeem M, Abrar M. Revolutionizing Brain Tumor Segmentation in MRI with Dynamic Fusion of Handcrafted Features and Global Pathway-based Deep Learning. KSII Transactions on Internet and Information Systems. 2024;18(1). https://doi.org/10.3837/tiis.2024.01.007
CrossRef - Çalışkan A. Diagnosis of malaria disease by integrating chi-square feature selection algorithm with convolutional neural networks and autoencoder network. Transactions of the Institute of Measurement and Control. 2023 ;45(5):975-85.
CrossRef - Nguyen-Tat TB, Nguyen TQT, Nguyen HN, Ngo VM. Enhancing brain tumor segmentation in MRI images: A hybrid approach using UNet, attention mechanisms, and transformers. Egyptian Informatics Journal. 2024; 27:100528. https://doi.org/10.1016/j.eij.2024.100528
CrossRef







